Regulating Tumor Suppressor Genes: Post-Translational Modifications
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Supplemental Materials ZNF281 Enhances Cardiac Reprogramming
Supplemental Materials ZNF281 enhances cardiac reprogramming by modulating cardiac and inflammatory gene expression Huanyu Zhou, Maria Gabriela Morales, Hisayuki Hashimoto, Matthew E. Dickson, Kunhua Song, Wenduo Ye, Min S. Kim, Hanspeter Niederstrasser, Zhaoning Wang, Beibei Chen, Bruce A. Posner, Rhonda Bassel-Duby and Eric N. Olson Supplemental Table 1; related to Figure 1. Supplemental Table 2; related to Figure 1. Supplemental Table 3; related to the “quantitative mRNA measurement” in Materials and Methods section. Supplemental Table 4; related to the “ChIP-seq, gene ontology and pathway analysis” and “RNA-seq” and gene ontology analysis” in Materials and Methods section. Supplemental Figure S1; related to Figure 1. Supplemental Figure S2; related to Figure 2. Supplemental Figure S3; related to Figure 3. Supplemental Figure S4; related to Figure 4. Supplemental Figure S5; related to Figure 6. Supplemental Table S1. Genes included in human retroviral ORF cDNA library. Gene Gene Gene Gene Gene Gene Gene Gene Symbol Symbol Symbol Symbol Symbol Symbol Symbol Symbol AATF BMP8A CEBPE CTNNB1 ESR2 GDF3 HOXA5 IL17D ADIPOQ BRPF1 CEBPG CUX1 ESRRA GDF6 HOXA6 IL17F ADNP BRPF3 CERS1 CX3CL1 ETS1 GIN1 HOXA7 IL18 AEBP1 BUD31 CERS2 CXCL10 ETS2 GLIS3 HOXB1 IL19 AFF4 C17ORF77 CERS4 CXCL11 ETV3 GMEB1 HOXB13 IL1A AHR C1QTNF4 CFL2 CXCL12 ETV7 GPBP1 HOXB5 IL1B AIMP1 C21ORF66 CHIA CXCL13 FAM3B GPER HOXB6 IL1F3 ALS2CR8 CBFA2T2 CIR1 CXCL14 FAM3D GPI HOXB7 IL1F5 ALX1 CBFA2T3 CITED1 CXCL16 FASLG GREM1 HOXB9 IL1F6 ARGFX CBFB CITED2 CXCL3 FBLN1 GREM2 HOXC4 IL1F7 -

Cellular and Molecular Signatures in the Disease Tissue of Early
Cellular and Molecular Signatures in the Disease Tissue of Early Rheumatoid Arthritis Stratify Clinical Response to csDMARD-Therapy and Predict Radiographic Progression Frances Humby1,* Myles Lewis1,* Nandhini Ramamoorthi2, Jason Hackney3, Michael Barnes1, Michele Bombardieri1, Francesca Setiadi2, Stephen Kelly1, Fabiola Bene1, Maria di Cicco1, Sudeh Riahi1, Vidalba Rocher-Ros1, Nora Ng1, Ilias Lazorou1, Rebecca E. Hands1, Desiree van der Heijde4, Robert Landewé5, Annette van der Helm-van Mil4, Alberto Cauli6, Iain B. McInnes7, Christopher D. Buckley8, Ernest Choy9, Peter Taylor10, Michael J. Townsend2 & Costantino Pitzalis1 1Centre for Experimental Medicine and Rheumatology, William Harvey Research Institute, Barts and The London School of Medicine and Dentistry, Queen Mary University of London, Charterhouse Square, London EC1M 6BQ, UK. Departments of 2Biomarker Discovery OMNI, 3Bioinformatics and Computational Biology, Genentech Research and Early Development, South San Francisco, California 94080 USA 4Department of Rheumatology, Leiden University Medical Center, The Netherlands 5Department of Clinical Immunology & Rheumatology, Amsterdam Rheumatology & Immunology Center, Amsterdam, The Netherlands 6Rheumatology Unit, Department of Medical Sciences, Policlinico of the University of Cagliari, Cagliari, Italy 7Institute of Infection, Immunity and Inflammation, University of Glasgow, Glasgow G12 8TA, UK 8Rheumatology Research Group, Institute of Inflammation and Ageing (IIA), University of Birmingham, Birmingham B15 2WB, UK 9Institute of -

WO 2019/079361 Al 25 April 2019 (25.04.2019) W 1P O PCT
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization I International Bureau (10) International Publication Number (43) International Publication Date WO 2019/079361 Al 25 April 2019 (25.04.2019) W 1P O PCT (51) International Patent Classification: CA, CH, CL, CN, CO, CR, CU, CZ, DE, DJ, DK, DM, DO, C12Q 1/68 (2018.01) A61P 31/18 (2006.01) DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, HN, C12Q 1/70 (2006.01) HR, HU, ID, IL, IN, IR, IS, JO, JP, KE, KG, KH, KN, KP, KR, KW, KZ, LA, LC, LK, LR, LS, LU, LY, MA, MD, ME, (21) International Application Number: MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, PCT/US2018/056167 OM, PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SA, (22) International Filing Date: SC, SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, 16 October 2018 (16. 10.2018) TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. (25) Filing Language: English (84) Designated States (unless otherwise indicated, for every kind of regional protection available): ARIPO (BW, GH, (26) Publication Language: English GM, KE, LR, LS, MW, MZ, NA, RW, SD, SL, ST, SZ, TZ, (30) Priority Data: UG, ZM, ZW), Eurasian (AM, AZ, BY, KG, KZ, RU, TJ, 62/573,025 16 October 2017 (16. 10.2017) US TM), European (AL, AT, BE, BG, CH, CY, CZ, DE, DK, EE, ES, FI, FR, GB, GR, HR, HU, ΓΕ , IS, IT, LT, LU, LV, (71) Applicant: MASSACHUSETTS INSTITUTE OF MC, MK, MT, NL, NO, PL, PT, RO, RS, SE, SI, SK, SM, TECHNOLOGY [US/US]; 77 Massachusetts Avenue, TR), OAPI (BF, BJ, CF, CG, CI, CM, GA, GN, GQ, GW, Cambridge, Massachusetts 02139 (US). -
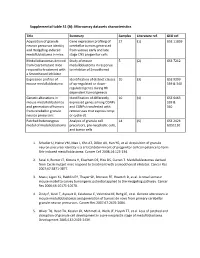
Supplemental Table S1 (A): Microarray Datasets Characteristics
Supplemental table S1 (A): Microarray datasets characteristics Title Summary Samples Literature ref. GEO ref. Acquisition of granule Gene expression profiling of 27 (1) GSE 11859 neuron precursor identity cerebellar tumors generated and Hedgehog‐induced from various early and late medulloblastoma in mice. stage CNS progenitor cells Medulloblastomas derived Study of mouse 5 (2) GSE 7212 from Cxcr6 mutant mice medulloblastoma in response respond to treatment with to inhibitor of Smoothened a Smoothened inhibitor Expression profiles of Identification of distinct classes 10 (3) GSE 9299 mouse medulloblastoma of up‐regulated or down‐ 339 & 340 regulated genes during Hh dependent tumorigenesis Genetic alterations in Identification of differently 10 (4) GSE 6463 mouse medulloblastomas expressed genes among CGNPs 339 & and generation of tumors and CGNPs transfected with 340 from cerebellar granule retroviruses that express nmyc neuron precursors or cyclin‐d1 Patched heterozygous Analysis of granule cell 14 (5) GSE 2426 model of medulloblastoma precursors, pre‐neoplastic cells, GDS1110 and tumor cells 1. Schuller U, Heine VM, Mao J, Kho AT, Dillon AK, Han YG, et al. Acquisition of granule neuron precursor identity is a critical determinant of progenitor cell competence to form Shh‐induced medulloblastoma. Cancer Cell 2008;14:123‐134. 2. Sasai K, Romer JT, Kimura H, Eberhart DE, Rice DS, Curran T. Medulloblastomas derived from Cxcr6 mutant mice respond to treatment with a smoothened inhibitor. Cancer Res 2007;67:3871‐3877. 3. Mao J, Ligon KL, Rakhlin EY, Thayer SP, Bronson RT, Rowitch D, et al. A novel somatic mouse model to survey tumorigenic potential applied to the Hedgehog pathway. Cancer Res 2006;66:10171‐10178. -

Trim24 Targets Endogenous P53 for Degradation
Trim24 targets endogenous p53 for degradation Kendra Alltona,b,1, Abhinav K. Jaina,b,1, Hans-Martin Herza, Wen-Wei Tsaia,b, Sung Yun Jungc, Jun Qinc, Andreas Bergmanna, Randy L. Johnsona,b, and Michelle Craig Bartona,b,2 aDepartment of Biochemistry and Molecular Biology, Program in Genes and Development, Graduate School of Biomedical Sciences and bCenter for Stem Cell and Developmental Biology, University of Texas M.D. Anderson Cancer Center, 1515 Holcombe Boulevard, Houston, TX 77030; and cDepartment of Molecular and Cellular Biology, Baylor College of Medicine, 1 Baylor Plaza, Houston, TX 77030 Edited by Carol L. Prives, Columbia University, New York, NY, and approved May 15, 2009 (received for review December 23, 2008) Numerous studies focus on the tumor suppressor p53 as a protector regulator of p53 suggests that potential therapeutic targets to of genomic stability, mediator of cell cycle arrest and apoptosis, restore p53 functions remain to be identified. and target of mutation in 50% of all human cancers. The vast majority of information on p53, its protein-interaction partners and Results and Discussion regulation, comes from studies of tumor-derived, cultured cells Creation of a Mouse and Stem Cell Model for p53 Analysis. To where p53 and its regulatory controls may be mutated or dysfunc- facilitate analysis of endogenous p53 in normal cells, we per- tional. To address regulation of endogenous p53 in normal cells, formed gene targeting to create mouse embryonic stem (ES) we created a mouse and stem cell model by knock-in (KI) of a cells and mice (12), which express endogenously regulated p53 tandem-affinity-purification (TAP) epitope at the endogenous protein fused with a C-terminal TAP tag (p53-TAPKI, Fig. -

Transcriptional and Post-Transcriptional Regulation of ATP-Binding Cassette Transporter Expression
Transcriptional and Post-transcriptional Regulation of ATP-binding Cassette Transporter Expression by Aparna Chhibber DISSERTATION Submitted in partial satisfaction of the requirements for the degree of DOCTOR OF PHILOSOPHY in Pharmaceutical Sciences and Pbarmacogenomies in the Copyright 2014 by Aparna Chhibber ii Acknowledgements First and foremost, I would like to thank my advisor, Dr. Deanna Kroetz. More than just a research advisor, Deanna has clearly made it a priority to guide her students to become better scientists, and I am grateful for the countless hours she has spent editing papers, developing presentations, discussing research, and so much more. I would not have made it this far without her support and guidance. My thesis committee has provided valuable advice through the years. Dr. Nadav Ahituv in particular has been a source of support from my first year in the graduate program as my academic advisor, qualifying exam committee chair, and finally thesis committee member. Dr. Kathy Giacomini graciously stepped in as a member of my thesis committee in my 3rd year, and Dr. Steven Brenner provided valuable input as thesis committee member in my 2nd year. My labmates over the past five years have been incredible colleagues and friends. Dr. Svetlana Markova first welcomed me into the lab and taught me numerous laboratory techniques, and has always been willing to act as a sounding board. Michael Martin has been my partner-in-crime in the lab from the beginning, and has made my days in lab fly by. Dr. Yingmei Lui has made the lab run smoothly, and has always been willing to jump in to help me at a moment’s notice. -
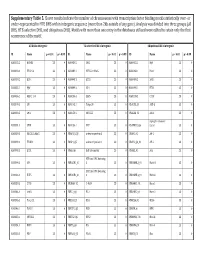
Supplementary Table 5. Clover Results Indicate the Number Of
Supplementary Table 5. Clover results indicate the number of chromosomes with transcription factor binding motifs statistically over‐ or under‐represented in HTE DHS within intergenic sequence (more than 2kb outside of any gene). Analysis was divided into three groups (all DHS, HTE‐selective DHS, and ubiquitous DHS). Motifs with more than one entry in the databases utilized were edited to retain only the first occurrence of the motif. All DHS x Intergenic TEselective DHS x Intergenic Ubiquitous DHS x Intergenic ID Name p < 0.01 p > 0.99 ID Name p < 0.01 p > 0.99 ID Name p < 0.01 p > 0.99 MA0002.2 RUNX1 23 0 MA0080.2 SPI1 23 0 MA0055.1 Myf 23 0 MA0003.1 TFAP2A 23 0 MA0089.1 NFE2L1::MafG 23 0 MA0068.1 Pax4 23 0 MA0039.2 Klf4 23 0 MA0098.1 ETS1 23 0 MA0080.2 SPI1 23 0 MA0055.1 Myf 23 0 MA0099.2 AP1 23 0 MA0098.1 ETS1 23 0 MA0056.1 MZF1_1‐4 23 0 MA0136.1 ELF5 23 0 MA0139.1 CTCF 23 0 MA0079.2 SP1 23 0 MA0145.1 Tcfcp2l1 23 0 V$ALX3_01 ALX‐3 23 0 MA0080.2 SPI1 23 0 MA0150.1 NFE2L2 23 0 V$ALX4_02 Alx‐4 23 0 myocyte enhancer MA0081.1 SPIB 23 0 MA0156.1 FEV 23 0 V$AMEF2_Q6 factor 23 0 MA0089.1 NFE2L1::MafG 23 0 V$AP1FJ_Q2 activator protein 1 23 0 V$AP1_01 AP‐1 23 0 MA0090.1 TEAD1 23 0 V$AP4_Q5 activator protein 4 23 0 V$AP2_Q6_01 AP‐2 23 0 MA0098.1 ETS1 23 0 V$AR_Q6 half‐site matrix 23 0 V$ARX_01 Arx 23 0 BTB and CNC homolog MA0099.2 AP1 23 0 V$BACH1_01 1 23 0 V$BARHL1_01 Barhl‐1 23 0 BTB and CNC homolog MA0136.1 ELF5 23 0 V$BACH2_01 2 23 0 V$BARHL2_01 Barhl2 23 0 MA0139.1 CTCF 23 0 V$CMAF_02 C‐MAF 23 0 V$BARX1_01 Barx1 23 0 MA0144.1 Stat3 23 0 -

Cardiac SARS‐Cov‐2 Infection Is Associated with Distinct Tran‐ Scriptomic Changes Within the Heart
Cardiac SARS‐CoV‐2 infection is associated with distinct tran‐ scriptomic changes within the heart Diana Lindner, PhD*1,2, Hanna Bräuninger, MS*1,2, Bastian Stoffers, MS1,2, Antonia Fitzek, MD3, Kira Meißner3, Ganna Aleshcheva, PhD4, Michaela Schweizer, PhD5, Jessica Weimann, MS1, Björn Rotter, PhD9, Svenja Warnke, BSc1, Carolin Edler, MD3, Fabian Braun, MD8, Kevin Roedl, MD10, Katharina Scher‐ schel, PhD1,12,13, Felicitas Escher, MD4,6,7, Stefan Kluge, MD10, Tobias B. Huber, MD8, Benjamin Ondruschka, MD3, Heinz‐Peter‐Schultheiss, MD4, Paulus Kirchhof, MD1,2,11, Stefan Blankenberg, MD1,2, Klaus Püschel, MD3, Dirk Westermann, MD1,2 1 Department of Cardiology, University Heart and Vascular Center Hamburg, Germany. 2 DZHK (German Center for Cardiovascular Research), partner site Hamburg/Kiel/Lübeck. 3 Institute of Legal Medicine, University Medical Center Hamburg‐Eppendorf, Germany. 4 Institute for Cardiac Diagnostics and Therapy, Berlin, Germany. 5 Department of Electron Microscopy, Center for Molecular Neurobiology, University Medical Center Hamburg‐Eppendorf, Germany. 6 Department of Cardiology, Charité‐Universitaetsmedizin, Berlin, Germany. 7 DZHK (German Centre for Cardiovascular Research), partner site Berlin, Germany. 8 III. Department of Medicine, University Medical Center Hamburg‐Eppendorf, Germany. 9 GenXPro GmbH, Frankfurter Innovationszentrum, Biotechnologie (FIZ), Frankfurt am Main, Germany. 10 Department of Intensive Care Medicine, University Medical Center Hamburg‐Eppendorf, Germany. 11 Institute of Cardiovascular Sciences, -

Trim24-Regulated Estrogen Response Is Dependent on Specific Histone Modifications in Breast Cancer Cells
The Texas Medical Center Library DigitalCommons@TMC The University of Texas MD Anderson Cancer Center UTHealth Graduate School of The University of Texas MD Anderson Cancer Biomedical Sciences Dissertations and Theses Center UTHealth Graduate School of (Open Access) Biomedical Sciences 12-2012 TRIM24-REGULATED ESTROGEN RESPONSE IS DEPENDENT ON SPECIFIC HISTONE MODIFICATIONS IN BREAST CANCER CELLS Teresa T. Yiu Follow this and additional works at: https://digitalcommons.library.tmc.edu/utgsbs_dissertations Part of the Biochemistry Commons, Cancer Biology Commons, Medicine and Health Sciences Commons, and the Molecular Biology Commons Recommended Citation Yiu, Teresa T., "TRIM24-REGULATED ESTROGEN RESPONSE IS DEPENDENT ON SPECIFIC HISTONE MODIFICATIONS IN BREAST CANCER CELLS" (2012). The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences Dissertations and Theses (Open Access). 313. https://digitalcommons.library.tmc.edu/utgsbs_dissertations/313 This Dissertation (PhD) is brought to you for free and open access by the The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences at DigitalCommons@TMC. It has been accepted for inclusion in The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences Dissertations and Theses (Open Access) by an authorized administrator of DigitalCommons@TMC. For more information, please contact [email protected]. TRIM24-REGULATED ESTROGEN RESPONSE IS DEPENDENT ON SPECIFIC HISTONE -

Reorganization of the Host Epigenome by a Viral Oncogene
Downloaded from genome.cshlp.org on September 29, 2021 - Published by Cold Spring Harbor Laboratory Press Research Reorganization of the host epigenome by a viral oncogene Roberto Ferrari,1,2 Trent Su,1,3 Bing Li,1 Giancarlo Bonora,1,4 Amit Oberai,1 Yvonne Chan,1 Rajkumar Sasidharan,5 Arnold J. Berk,6,7 Matteo Pellegrini,2,5 and Siavash K. Kurdistani1,2,7,8,9 1Department of Biological Chemistry, David Geffen School of Medicine, University of California, Los Angeles, California 90095, USA; 2Eli and Edythe Broad Center of Regenerative Medicine and Stem Cell Research, University of California, Los Angeles, California 90095, USA; 3Division of Oral Biology and Medicine, School of Dentistry, David Geffen School of Medicine, University of California, Los Angeles, California 90095, USA; 4UCLA Bioinformatics Interdepartmental Degree Program, David Geffen School of Medicine, University of California, Los Angeles, California 90095, USA; 5Department of Molecular, Cellular, and Developmental Biology, University of California, Los Angeles, California 90095, USA; 6Department of Microbiology, Immunology and Molecular Genetics, University of California, Los Angeles, California 90095, USA; 7Molecular Biology Institute, University of California, Los Angeles, California 90095, USA; 8Department of Pathology and Laboratory of Medicine, David Geffen School of Medicine, University of California, Los Angeles, California 90095, USA Adenovirus small e1a oncoprotein causes ~70% reduction in cellular levels of histone H3 lysine 18 acetylation (H3K18ac). It is unclear, however, where this dramatic reduction occurs genome-wide. ChIP-sequencing revealed that by 24 h after expression, e1a erases 95% of H3K18ac peaks in normal, contact-inhibited fibroblasts and replaces them with one-third as many at new genomic locations. -
Figure S1. Reverse Transcription‑Quantitative PCR Analysis of ETV5 Mrna Expression Levels in Parental and ETV5 Stable Transfectants
Figure S1. Reverse transcription‑quantitative PCR analysis of ETV5 mRNA expression levels in parental and ETV5 stable transfectants. (A) Hec1a and Hec1a‑ETV5 EC cell lines; (B) Ishikawa and Ishikawa‑ETV5 EC cell lines. **P<0.005, unpaired Student's t‑test. EC, endometrial cancer; ETV5, ETS variant transcription factor 5. Figure S2. Survival analysis of sample clusters 1‑4. Kaplan Meier graphs for (A) recurrence‑free and (B) overall survival. Survival curves were constructed using the Kaplan‑Meier method, and differences between sample cluster curves were analyzed by log‑rank test. Figure S3. ROC analysis of hub genes. For each gene, ROC curve (left) and mRNA expression levels (right) in control (n=35) and tumor (n=545) samples from The Cancer Genome Atlas Uterine Corpus Endometrioid Cancer cohort are shown. mRNA levels are expressed as Log2(x+1), where ‘x’ is the RSEM normalized expression value. ROC, receiver operating characteristic. Table SI. Clinicopathological characteristics of the GSE17025 dataset. Characteristic n % Atrophic endometrium 12 (postmenopausal) (Control group) Tumor stage I 91 100 Histology Endometrioid adenocarcinoma 79 86.81 Papillary serous 12 13.19 Histological grade Grade 1 30 32.97 Grade 2 36 39.56 Grade 3 25 27.47 Myometrial invasiona Superficial (<50%) 67 74.44 Deep (>50%) 23 25.56 aMyometrial invasion information was available for 90 of 91 tumor samples. Table SII. Clinicopathological characteristics of The Cancer Genome Atlas Uterine Corpus Endometrioid Cancer dataset. Characteristic n % Solid tissue normal 16 Tumor samples Stagea I 226 68.278 II 19 5.740 III 70 21.148 IV 16 4.834 Histology Endometrioid 271 81.381 Mixed 10 3.003 Serous 52 15.616 Histological grade Grade 1 78 23.423 Grade 2 91 27.327 Grade 3 164 49.249 Molecular subtypeb POLE 17 7.328 MSI 65 28.017 CN Low 90 38.793 CN High 60 25.862 CN, copy number; MSI, microsatellite instability; POLE, DNA polymerase ε.