Next-Generation Sequencing Approach For
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Y Chromosomal Noncoding Rnas Regulate Autosomal Gene Expression Via Pirnas in Mouse Testis Hemakumar M
bioRxiv preprint doi: https://doi.org/10.1101/285429; this version posted January 15, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Y chromosomal noncoding RNAs regulate autosomal gene expression via piRNAs in mouse testis Hemakumar M. Reddy1,2,16, Rupa Bhattacharya1,3,16, Zeenath Jehan1,4, Kankadeb Mishra1,5, Pranatharthi Annapurna1,6, Shrish Tiwari1, Nissankararao Mary Praveena1, Jomini Liza Alex1, Vishnu M Dhople1,7, Lalji Singh8, Mahadevan Sivaramakrishnan1,9, Anurag Chaturvedi1,10, Nandini Rangaraj1, Thomas Michael Shiju1,11, Badanapuram Sridevi1, Sachin Kumar1, Ram Reddy Dereddi1,12, Sunayana M Rayabandla1,13, Rachel A. Jesudasan1,14*, 15* 1Centre for Cellular and Molecular Biology (CCMB), Uppal Road, Hyderabad, Telengana – 500007, India. Present address: 2Brown University BioMed Division, Department of Molecular Biology, Cell Biology and Biochemistry, 185 Meeting Street room 260, Sidney Frank Life Sciences Building, Providence, RI 02912, USA. 3 Pennington NJ 08534, USA. 4Department of Genetics and Molecular Medicines, Vasavi Medical and Research Centre, 6- 1-91 Khairatabad, Hyderabad 500 004 India. 5Department of Cell Biology, Memorial Sloan Kettering Cancer Centre, Rockefeller Research Laboratory, 430 East 67th Street, RRL 445, New York, NY 10065, USA. 6Departments of Orthopaedic Surgery & Bioengineering, University of Pennsylvania, 376A Stemmler Hall, 36th Street & Hamilton Walk, Philadelphia, PA 19104.USA. 7Ernst-Moritz-Arndt-University of Greifswald Interfaculty Institute for Genetics and Functional Genomics, Department of Functional Genomics, Friedrich-Ludwig-Jahn-Straße 15 a, 17487 Greifswald, Germany. 8 Deceased. 9Jubilant Biosystems Ltd., #96, Industrial Suburb, 2nd Stage, Yeshwantpur, Bangalore- 560022, Karnataka, India. -

Involvement of DPP9 in Gene Fusions in Serous Ovarian Carcinoma
Smebye et al. BMC Cancer (2017) 17:642 DOI 10.1186/s12885-017-3625-6 RESEARCH ARTICLE Open Access Involvement of DPP9 in gene fusions in serous ovarian carcinoma Marianne Lislerud Smebye1,2, Antonio Agostini1,2, Bjarne Johannessen2,3, Jim Thorsen1,2, Ben Davidson4,5, Claes Göran Tropé6, Sverre Heim1,2,5, Rolf Inge Skotheim2,3 and Francesca Micci1,2* Abstract Background: A fusion gene is a hybrid gene consisting of parts from two previously independent genes. Chromosomal rearrangements leading to gene breakage are frequent in high-grade serous ovarian carcinomas and have been reported as a common mechanism for inactivating tumor suppressor genes. However, no fusion genes have been repeatedly reported to be recurrent driver events in ovarian carcinogenesis. We combined genomic and transcriptomic information to identify novel fusion gene candidates and aberrantly expressed genes in ovarian carcinomas. Methods: Examined were 19 previously karyotyped ovarian carcinomas (18 of the serous histotype and one undifferentiated). First, karyotypic aberrations were compared to fusion gene candidates identified by RNA sequencing (RNA-seq). In addition, we used exon-level gene expression microarrays as a screening tool to identify aberrantly expressed genes possibly involved in gene fusion events, and compared the findings to the RNA-seq data. Results: We found a DPP9-PPP6R3 fusion transcript in one tumor showing a matching genomic 11;19-translocation. Another tumor had a rearrangement of DPP9 with PLIN3. Both rearrangements were associated with diminished expression of the 3′ end of DPP9 corresponding to the breakpoints identified by RNA-seq. For the exon-level expression analysis, candidate fusion partner genes were ranked according to deviating expression compared to the median of the sample set. -

Supporting Information
Supporting Information Biagioli et al. 10.1073/pnas.0813216106 SI Text and the procedure was repeated twice. Single cells were then RNA Isolation, Reverse Transcription, qPCR, Cloning, and Sequencing. resuspended in panning buffer and incubated on lectin-coated RNA was extracted from cell lines and blood by using TRIzol panning plates for 15 min at room temperature. Nonadherent reagent (Invitrogen) and following vendor instructions. cells were transferred to the next panning plate (four pannings, RNA was extracted from LCM- or FACS-purified cells with an 15 min each). Then, nonadherent cells were collected, centri- Absolutely RNA Nanoprep Kit (Stratagene). Single-strand fuged, and resuspended in serum-free neuronal medium or PBS. cDNA was obtained from purified RNA by using the iSCRIPT A similar procedure was also followed for the dissociation of cDNA Synhesis Kit (Bio-Rad) according to the manufacturer’s cortical and hippocampal astrocytes and oligodendrocytes. instructions. Quantitative RT-PCR was performed by using A cell strainer with 70-m nylon mesh was used to obtain a SYBER-Green PCR Master Mix and an iQ5 Real-Time PCR single-cell suspension (BD Falcon) before sorting. 7-AAD Detection System (Bio-Rad). (Beckman–Coulter) was added to the cell suspension to exclude Quantitative RT-PCR was performed with an iCycler IQ dead cells. A high-speed cell sorter (MoFlo) was used to sort (Bio-Rad); -actin was used as an endogenous control to nor- subpopulation of cells expressing GFP. Sorting parameters used malize the expression level of target genes. Primers were chosen for the three different populations are visualized in Fig. -

(12) United States Patent (10) Patent No.: US 7,799,528 B2 Civin Et Al
US007799528B2 (12) United States Patent (10) Patent No.: US 7,799,528 B2 Civin et al. (45) Date of Patent: Sep. 21, 2010 (54) THERAPEUTIC AND DIAGNOSTIC Al-Hajj et al., “Prospective identification of tumorigenic breast can APPLICATIONS OF GENES cer cells.” Proc. Natl. Acad. Sci. U.S.A., 100(7):3983-3988 (2003). DIFFERENTIALLY EXPRESSED IN Bhatia et al., “A newly discovered class of human hematopoietic cells LYMPHO-HEMATOPOETC STEM CELLS with SCID-repopulating activity.” Nat. Med. 4(9)1038-45 (1998). (75) Inventors: Curt I. Civin, Baltimore, MD (US); Bonnet, D., “Normal and leukemic CD35-negative human Robert W. Georgantas, III, Towson, hematopoletic stem cells.” Rev. Clin. Exp. Hematol. 5:42-61 (2001). Cambot et al., “Human Immune Associated Nucleotide 1: a member MD (US) of a new guanosine triphosphatase family expressed in resting T and (73) Assignee: The Johns Hopkins University, B cells.” Blood, 99(9):3293-3301 (2002). Baltimore, MD (US) Chen et al., “Kruppel-like Factor 4 (Gut-enriched Kruppel-like Fac tor) Inhibits Cell Proliferation by Blocking G1/S Progression of the *) NotOt1Ce: Subjubject to anyy d1Sclaimer,disclai theh term off thisthi Cell Cycle.” J. Biol. Chem., 276(32):30423-30428 (2001). patent is extended or adjusted under 35 Chen et al., “Transcriptional profiling of Kurppel-like factor 4 reveals U.S.C. 154(b) by 168 days. a function in cell cycle regulation and epithelial differentiation.” J. Mol. Biol. 326(3):665-677 (2003). (21) Appl. No.: 11/199.665 Civin et al., “Highly purified CD34-positive cells reconstitute (22) Filed: Aug. 9, 2005 hematopoiesis,” J. -
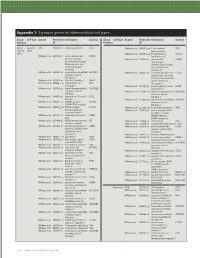
Technical Note, Appendix: an Analysis of Blood Processing Methods to Prepare Samples for Genechip® Expression Profiling (Pdf, 1
Appendix 1: Signature genes for different blood cell types. Blood Cell Type Source Probe Set Description Symbol Blood Cell Type Source Probe Set Description Symbol Fraction ID Fraction ID Mono- Lympho- GSK 203547_at CD4 antigen (p55) CD4 Whitney et al. 209813_x_at T cell receptor TRG nuclear cytes gamma locus cells Whitney et al. 209995_s_at T-cell leukemia/ TCL1A Whitney et al. 203104_at colony stimulating CSF1R lymphoma 1A factor 1 receptor, Whitney et al. 210164_at granzyme B GZMB formerly McDonough (granzyme 2, feline sarcoma viral cytotoxic T-lymphocyte- (v-fms) oncogene associated serine homolog esterase 1) Whitney et al. 203290_at major histocompatibility HLA-DQA1 Whitney et al. 210321_at similar to granzyme B CTLA1 complex, class II, (granzyme 2, cytotoxic DQ alpha 1 T-lymphocyte-associated Whitney et al. 203413_at NEL-like 2 (chicken) NELL2 serine esterase 1) Whitney et al. 203828_s_at natural killer cell NK4 (H. sapiens) transcript 4 Whitney et al. 212827_at immunoglobulin heavy IGHM Whitney et al. 203932_at major histocompatibility HLA-DMB constant mu complex, class II, Whitney et al. 212998_x_at major histocompatibility HLA-DQB1 DM beta complex, class II, Whitney et al. 204655_at chemokine (C-C motif) CCL5 DQ beta 1 ligand 5 Whitney et al. 212999_x_at major histocompatibility HLA-DQB Whitney et al. 204661_at CDW52 antigen CDW52 complex, class II, (CAMPATH-1 antigen) DQ beta 1 Whitney et al. 205049_s_at CD79A antigen CD79A Whitney et al. 213193_x_at T cell receptor beta locus TRB (immunoglobulin- Whitney et al. 213425_at Homo sapiens cDNA associated alpha) FLJ11441 fis, clone Whitney et al. 205291_at interleukin 2 receptor, IL2RB HEMBA1001323, beta mRNA sequence Whitney et al. -

Recombinant Human Hemoglobin Subunit Theta-1/HBQ1 (N-6His)
9853 Pacific Heights Blvd. Suite D. San Diego, CA 92121, USA Tel: 858-263-4982 Email: [email protected] 32-8592: Recombinant Human Hemoglobin Subunit theta-1/HBQ1 (N-6His) Gene : HBQ1 Gene ID : 3049 Uniprot ID : P09105 Description Source: E.coli. MW :17.7kD. Recombinant Human Hemoglobin subunit theta-1 is produced by our E.coli expression system and the target gene encoding Met1-Arg142 is expressed with a 6His tag at the N-terminus. Hemoglobin subunit theta-1 is a protein that in humans is encoded by the HBQ1 gene. Theta-globin mRNA is originally found in human fetal erythroid tissue but not in adult erythroid or other nonerythroid tissue. Theta-1 is a member of the human alpha-globin gene cluster that includes five functional genes and two pseudogenes. Research supports a transcriptionally active role for the gene and a functional role for the peptide in specific cells, possibly those of early erythroid tissue. Hemoglobin has a quaternary structure characteristically composed of many multi- subunit globular proteins. Most of the amino acids in hemoglobin form alpha helices, connected by short non-helical segments. Hydrogen bonds stabilize the helical sections inside this protein, causing attractions within the molecule, folding each polypeptide chain into a specific shape. Hemoglobin's quaternary structure comes from its four subunits in roughly a tetrahedral arrangement. Product Info Amount : 10 µg / 50 µg Content : Lyophilized from a 0.2 µm filtered solution of 20mM PB, 150mM NaCl, pH 7.0. Lyophilized protein should be stored at -20°C, though stable at room temperature for 3 weeks. -

Simultaneous Detection of Target Cnvs and Snvs of Thalassemia by Multiplex PCR and Next‑Generation Sequencing
MOLECULAR MEDICINE REPORTS 19: 2837-2848, 2019 Simultaneous detection of target CNVs and SNVs of thalassemia by multiplex PCR and next‑generation sequencing DONG-MEI FAN1*, XU YANG2*, LI-MIN HUANG1, GUO-JUN OUYANG3, XUE-XI YANG1 and MING LI1 1Institute of Antibody Engineering, School of Laboratory Medicine and Biotechnology, Southern Medical University, Guangzhou, Guangdong 510515; 2Clinical Innovation and Research Center, Shenzhen Hospital of Southern Medical University, Shenzhen, Guangdong 518110; 3Guangzhou Darui Biotechnology Co., Ltd., Guangzhou, Guangdong 510663, P.R. China Received May 16, 2018; Accepted December 3, 2018 DOI: 10.3892/mmr.2019.9896 Abstract. Thalassemia is caused by complex mechanisms, Introduction including copy number variants (CNVs) and single nucleotide variants (SNVs). The CNV types of α-thalassemia are typi- Thalassemia is caused by copy number variants (CNVs) and cally detected by gap-polymerase chain reaction (PCR). The single nucleotide variants (SNVs) in the α-globin (HBA) or SNV types are detected by Sanger sequencing. In the present β-globin (HBB) genes that result in the absence or lack of α- or study, a novel method was developed that simultaneously β-globin chains, and ultimately hemolytic anemia. It is esti- detects CNVs and SNVs by multiplex PCR and next-genera- mated that ~7% of the world population carries the gene for the tion sequencing (NGS). To detect CNVs, 33 normal samples disease (1), and the birth rate of children with hemoglobin (Hb) were used as a cluster of control values to build a baseline, disorders is ≥2.4% per year (2). Thalassemia occurs most in the and the A, B, C, and D ratios were developed to evaluate-SEA, Mediterranean region, East South Asia, and the subcontinents -α4.2, -α3.7, and compound or homozygous CNVs, respectively. -

Obesity Is Associated with More Activated Neutrophils in African American Male Youth
International Journal of Obesity (2015) 39, 26–32 © 2015 Macmillan Publishers Limited All rights reserved 0307-0565/15 www.nature.com/ijo PEDIATRIC ORIGINAL ARTICLE Obesity is associated with more activated neutrophils in African American male youth XXu1,SSu1, X Wang1, V Barnes1, C De Miguel2, D Ownby3, J Pollock2, H Snieder4, W Chen5 and X Wang1 BACKGROUND: There is emerging evidence suggesting the role of peripheral blood leukocytes in the pathogenesis of obesity and related diseases. However, few studies have taken a genome-wide approach to investigating gene expression profiles in peripheral leukocytes between obese and lean individuals with the consideration of obesity-related shifts in leukocyte types. METHOD: We conducted this study in 95 African Americans (AAs) of both genders (age 14–20 years, 46 lean and 49 obese). Complete blood count with differential test (CBC) was performed in whole blood. Genome-wide gene expression analysis was obtained using the Illumina HumanHT-12 V4 Beadchip with RNA extracted from peripheral leukocytes. Out of the 95 participants, 64 had neutrophils stored. The validation study was based on real-time PCR with RNA extracted from purified neutrophils. RESULTS: CBC test suggested that, in males, obesity was associated with increased neutrophil percentage (P = 0.03). Genome-wide gene expression analysis showed that, in males, the majority of the most differentially expressed genes were related to neutrophil activation. Validation of the gene expression levels of ELANE (neutrophil elastase) and MPO (myeloperoxidase) in purified neutrophils demonstrated that the expression of these two genes—important biomarkers of neutrophils activation—were significantly elevated in obese males (P = 0.01 and P = 0.02, respectively). -

Role and Regulation of the P53-Homolog P73 in the Transformation of Normal Human Fibroblasts
Role and regulation of the p53-homolog p73 in the transformation of normal human fibroblasts Dissertation zur Erlangung des naturwissenschaftlichen Doktorgrades der Bayerischen Julius-Maximilians-Universität Würzburg vorgelegt von Lars Hofmann aus Aschaffenburg Würzburg 2007 Eingereicht am Mitglieder der Promotionskommission: Vorsitzender: Prof. Dr. Dr. Martin J. Müller Gutachter: Prof. Dr. Michael P. Schön Gutachter : Prof. Dr. Georg Krohne Tag des Promotionskolloquiums: Doktorurkunde ausgehändigt am Erklärung Hiermit erkläre ich, dass ich die vorliegende Arbeit selbständig angefertigt und keine anderen als die angegebenen Hilfsmittel und Quellen verwendet habe. Diese Arbeit wurde weder in gleicher noch in ähnlicher Form in einem anderen Prüfungsverfahren vorgelegt. Ich habe früher, außer den mit dem Zulassungsgesuch urkundlichen Graden, keine weiteren akademischen Grade erworben und zu erwerben gesucht. Würzburg, Lars Hofmann Content SUMMARY ................................................................................................................ IV ZUSAMMENFASSUNG ............................................................................................. V 1. INTRODUCTION ................................................................................................. 1 1.1. Molecular basics of cancer .......................................................................................... 1 1.2. Early research on tumorigenesis ................................................................................. 3 1.3. Developing -

ELA2 Monoclonal Antibody, Clone AOCI-5
ELA2 monoclonal antibody, clone Gene Symbol: ELA2 AOCI-5 Gene Alias: GE, HLE, HNE, NE, PMN-E Catalog Number: MAB20237 Gene Summary: Elastases form a subfamily of serine proteases that hydrolyze many proteins in addition to Regulatory Status: For research use only (RUO) elastin. Humans have six elastase genes which encode the structurally similar proteins. The product of this gene Product Description: Rabbit monoclonal antibody hydrolyzes proteins within specialized neutrophil raised against synthetic peptide of human ELA2. lysosomes, called azurophil granules, as well as proteins Clone Name: AOCI-5 of the extracellular matrix following the protein's release from activated neutrophils. The enzyme may play a role Immunogen: A synthetic peptide corresponding to in degenerative and inflammatory diseases by its human ELA2. proteolysis of collagen-IV and elastin of the extracellular matrix. This protein degrades the outer membrane Host: Rabbit protein A (OmpA) of E. coli as well as the virulence factors of such bacteria as Shigella, Salmonella and Reactivity: Human Yersinia. Mutations in this gene are associated with cyclic neutropenia and severe congenital neutropenia Applications: Flow Cyt, ICC, IF, IHC-P, WB-Ce (SCN). This gene is clustered with other serine protease (See our web site product page for detailed applications gene family members, azurocidin 1 and proteinase 3 information) genes, at chromosome 19pter. All 3 genes are expressed coordinately and their protein products are Protocols: See our web site at packaged together into azurophil granules during http://www.abnova.com/support/protocols.asp or product neutrophil differentiation. [provided by RefSeq] page for detailed protocols Form: Liquid Purification: Affinity purification Isotype: IgG Recommend Usage: Flow Cytometry (1:30) Immunocytochemistry (1:50-1:200) Immunofluorescence (1:50-1:200) Immunohistochemistry (1:100-1:500) Western Blot (1:500-1:2000) The optimal working dilution should be determined by the end user. -

Integrative Meta-Analysis of Differential Gene Expression in Acute Myeloid Leukemia
Integrative Meta-Analysis of Differential Gene Expression in Acute Myeloid Leukemia Brady G. Miller1, John A. Stamatoyannopoulos2* 1 Department of Hematology, University of Washington, Seattle, Washington, United States of America, 2 Department of Genome Sciences, University of Washington, Seattle, Washington, United States of America Abstract Background: Acute myeloid leukemia (AML) is a heterogeneous disease with an overall poor prognosis. Gene expression profiling studies of patients with AML has provided key insights into disease pathogenesis while exposing potential diagnostic and prognostic markers and therapeutic targets. A systematic comparison of the large body of gene expression profiling studies in AML has the potential to test the extensibility of conclusions based on single studies and provide further insights into AML. Methodology/Principal Findings: In this study, we systematically compared 25 published reports of gene expression profiling in AML. There were a total of 4,918 reported genes of which one third were reported in more than one study. We found that only a minority of reported prognostically-associated genes (9.6%) were replicated in at least one other study. In a combined analysis, we comprehensively identified both gene sets and functional gene categories and pathways that exhibited significant differential regulation in distinct prognostic categories, including many previously unreported associations. Conclusions/Significance: We developed a novel approach for granular, cross-study analysis of gene-by-gene data and their relationships with established prognostic features and patient outcome. We identified many robust novel prognostic molecular features in AML that were undetected in prior studies, and which provide insights into AML pathogenesis with potential diagnostic, prognostic, and therapeutic implications. -

Stem Cells® Original Article
® Stem Cells Original Article Properties of Pluripotent Human Embryonic Stem Cells BG01 and BG02 XIANMIN ZENG,a TAKUMI MIURA,b YONGQUAN LUO,b BHASKAR BHATTACHARYA,c BRIAN CONDIE,d JIA CHEN,a IRENE GINIS,b IAN LYONS,d JOSEF MEJIDO,c RAJ K. PURI,c MAHENDRA S. RAO,b WILLIAM J. FREEDa aCellular Neurobiology Research Branch, National Institute on Drug Abuse, Department of Health and Human Services (DHHS), Baltimore, Maryland, USA; bLaboratory of Neuroscience, National Institute of Aging, DHHS, Baltimore, Maryland, USA; cLaboratory of Molecular Tumor Biology, Division of Cellular and Gene Therapies, Center for Biologics Evaluation and Research, Food and Drug Administration, Bethesda, Maryland, USA; dBresaGen Inc., Athens, Georgia, USA Key Words. Embryonic stem cells · Differentiation · Microarray ABSTRACT Human ES (hES) cell lines have only recently been compared with pooled human RNA. Ninety-two of these generated, and differences between human and mouse genes were also highly expressed in four other hES lines ES cells have been identified. In this manuscript we (TE05, GE01, GE09, and pooled samples derived from describe the properties of two human ES cell lines, GE01, GE09, and GE07). Included in the list are genes BG01 and BG02. By immunocytochemistry and reverse involved in cell signaling and development, metabolism, transcription polymerase chain reaction, undifferenti- transcription regulation, and many hypothetical pro- ated cells expressed markers that are characteristic of teins. Two focused arrays designed to examine tran- ES cells, including SSEA-3, SSEA-4, TRA-1-60, TRA-1- scripts associated with stem cells and with the 81, and OCT-3/4. Both cell lines were readily main- transforming growth factor-β superfamily were tained in an undifferentiated state and could employed to examine differentially expressed genes.