Enzyme Activity in the Crowded Milieu
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Characterization of the Ergosterol Biosynthesis Pathway in Ceratocystidaceae
Journal of Fungi Article Characterization of the Ergosterol Biosynthesis Pathway in Ceratocystidaceae Mohammad Sayari 1,2,*, Magrieta A. van der Nest 1,3, Emma T. Steenkamp 1, Saleh Rahimlou 4 , Almuth Hammerbacher 1 and Brenda D. Wingfield 1 1 Department of Biochemistry, Genetics and Microbiology, Forestry and Agricultural Biotechnology Institute (FABI), University of Pretoria, Pretoria 0002, South Africa; [email protected] (M.A.v.d.N.); [email protected] (E.T.S.); [email protected] (A.H.); brenda.wingfi[email protected] (B.D.W.) 2 Department of Plant Science, University of Manitoba, 222 Agriculture Building, Winnipeg, MB R3T 2N2, Canada 3 Biotechnology Platform, Agricultural Research Council (ARC), Onderstepoort Campus, Pretoria 0110, South Africa 4 Department of Mycology and Microbiology, University of Tartu, 14A Ravila, 50411 Tartu, Estonia; [email protected] * Correspondence: [email protected]; Fax: +1-204-474-7528 Abstract: Terpenes represent the biggest group of natural compounds on earth. This large class of organic hydrocarbons is distributed among all cellular organisms, including fungi. The different classes of terpenes produced by fungi are mono, sesqui, di- and triterpenes, although triterpene ergosterol is the main sterol identified in cell membranes of these organisms. The availability of genomic data from members in the Ceratocystidaceae enabled the detection and characterization of the genes encoding the enzymes in the mevalonate and ergosterol biosynthetic pathways. Using Citation: Sayari, M.; van der Nest, a bioinformatics approach, fungal orthologs of sterol biosynthesis genes in nine different species M.A.; Steenkamp, E.T.; Rahimlou, S.; of the Ceratocystidaceae were identified. -

1 Metabolic Dysfunction Is Restricted to the Sciatic Nerve in Experimental
Page 1 of 255 Diabetes Metabolic dysfunction is restricted to the sciatic nerve in experimental diabetic neuropathy Oliver J. Freeman1,2, Richard D. Unwin2,3, Andrew W. Dowsey2,3, Paul Begley2,3, Sumia Ali1, Katherine A. Hollywood2,3, Nitin Rustogi2,3, Rasmus S. Petersen1, Warwick B. Dunn2,3†, Garth J.S. Cooper2,3,4,5* & Natalie J. Gardiner1* 1 Faculty of Life Sciences, University of Manchester, UK 2 Centre for Advanced Discovery and Experimental Therapeutics (CADET), Central Manchester University Hospitals NHS Foundation Trust, Manchester Academic Health Sciences Centre, Manchester, UK 3 Centre for Endocrinology and Diabetes, Institute of Human Development, Faculty of Medical and Human Sciences, University of Manchester, UK 4 School of Biological Sciences, University of Auckland, New Zealand 5 Department of Pharmacology, Medical Sciences Division, University of Oxford, UK † Present address: School of Biosciences, University of Birmingham, UK *Joint corresponding authors: Natalie J. Gardiner and Garth J.S. Cooper Email: [email protected]; [email protected] Address: University of Manchester, AV Hill Building, Oxford Road, Manchester, M13 9PT, United Kingdom Telephone: +44 161 275 5768; +44 161 701 0240 Word count: 4,490 Number of tables: 1, Number of figures: 6 Running title: Metabolic dysfunction in diabetic neuropathy 1 Diabetes Publish Ahead of Print, published online October 15, 2015 Diabetes Page 2 of 255 Abstract High glucose levels in the peripheral nervous system (PNS) have been implicated in the pathogenesis of diabetic neuropathy (DN). However our understanding of the molecular mechanisms which cause the marked distal pathology is incomplete. Here we performed a comprehensive, system-wide analysis of the PNS of a rodent model of DN. -

Mechanistic Implications for the Chorismatase Fkbo Based on the Crystal Structure
Mechanistic Implications for the Chorismatase FkbO Based on the Crystal Structure Puneet Juneja 1,†, Florian Hubrich 2,†, Kay Diederichs 1, Wolfram Welte 1 and Jennifer N. Andexer 2 1 - Department of Biology, University of Konstanz, D-78457 Konstanz, Germany 2 - Institute of Pharmaceutical Sciences, Albert-Ludwigs-University Freiburg, Albertstr 25, D-79104 Freiburg, Germany Correspondence to Jennifer N. Andexer: [email protected] http://dx.doi.org/10.1016/j.jmb.2013.09.006 Edited by M. Guss Abstract Chorismate-converting enzymes are involved in many biosynthetic pathways leading to natural products and can often be used as tools for the synthesis of chemical building blocks. Chorismatases such as FkbO from Streptomyces species catalyse the hydrolysis of chorismate yielding (dihydro)benzoic acid derivatives. In contrast to many other chorismate-converting enzymes, the structure and catalytic mechanism of a chorismatase had not been previously elucidated. Here we present the crystal structure of the chorismatase FkbO in complex with a competitive inhibitor at 1.08 Å resolution. FkbO is a monomer in solution and exhibits pseudo-3-fold symmetry; the structure of the individual domains indicates a possible connection to the trimeric RidA/YjgF family and related enzymes. The co-crystallised inhibitor led to the identification of FkbO's active site in the cleft between the central and the C-terminal domains. A mechanism for FkbO is proposed based on both interactions between the inhibitor and the surrounding amino acids and an FkbO structure with chorismate modelled in the active site. We suggest that the methylene group of the chorismate enol ether takes up a proton from an active-site glutamic acid residue, thereby initiating chorismate hydrolysis. -

Phza/B Catalyzes the Formation of the Tricycle in Phenazine Biosynthesis Ekta G
Subscriber access provided by DigiTop | USDA's Digital Desktop Library Article PhzA/B Catalyzes the Formation of the Tricycle in Phenazine Biosynthesis Ekta G. Ahuja, Petra Janning, Matthias Mentel, Almut Graebsch, Rolf Breinbauer, Wolf Hiller, Burkhard Costisella, Linda S. Thomashow, Dmitri V. Mavrodi, and Wulf Blankenfeldt J. Am. Chem. Soc., 2008, 130 (50), 17053-17061 • DOI: 10.1021/ja806325k • Publication Date (Web): 17 November 2008 Downloaded from http://pubs.acs.org on January 15, 2009 More About This Article Additional resources and features associated with this article are available within the HTML version: • Supporting Information • Access to high resolution figures • Links to articles and content related to this article • Copyright permission to reproduce figures and/or text from this article Journal of the American Chemical Society is published by the American Chemical Society. 1155 Sixteenth Street N.W., Washington, DC 20036 Published on Web 11/17/2008 PhzA/B Catalyzes the Formation of the Tricycle in Phenazine Biosynthesis Ekta G. Ahuja,† Petra Janning,† Matthias Mentel,‡,§ Almut Graebsch,‡ Rolf Breinbauer,†,‡,§,| Wolf Hiller,‡ Burkhard Costisella,‡ Linda S. Thomashow,⊥,# Dmitri V. Mavrodi,⊥ and Wulf Blankenfeldt*,† Max-Planck-Institute of Molecular Physiology, Otto-Hahn-Strasse 11, 44227 Dortmund, Germany, Technical UniVersity of Dortmund, Faculty of Chemistry, Otto-Hahn-Strasse 6, 44221 Dortmund, Germany, UniVersity of Leipzig, Institute of Organic Chemistry, Johannisallee 29, 04103 Leipzig, Germany, Graz UniVersity of -

Catalase: Bioinformatics Analyses of One of the Key Enzymes in Hydrogen Peroxide Metabolism
6–71RYHPEHU 2019, Brno, Czech Republic Catalase: Bioinformatics analyses of one of the key enzymes in hydrogen peroxide metabolism Michaela Kameniarova, Romana Kopecka Department of Molecular Biology and Radiobiology Mendel University in Brno Zemedelska 1, 613 00 Brno CZECH REPUBLIC [email protected] Abstract: Catalases (CAT) are family of important antioxidant enzymes present in different isoforms and responsible for scavenging of hydrogen peroxide in almost all aerobically living organisms. In plants, they were found both in unicellular and multicellular species. Here, we performed bioinformatics analysis and analysed catalase evolutionary relationship and expression patterns. By comparing expression profiles of CATs and expression profiles of genes related to abiotic stimuli we found that almost 50% of light signalling genes were co-expressed with CATs. Further, by datamining in available resources and structural modelling we pinpointed candidate amino acid residues responsible for CAT thermostability. Key Words: catalase, H2O2, phylogenetics, abiotic stress, stability INTRODUCTION Plants contain several types of enzymes involved in H2O2 metabolism. These include catalases, ascorbate peroxidases, peroxiredoxins, glutathione/thioredoxin peroxidases, and glutathione S- transferases, but only catalases do not require additional cellular reductants (Mhamdi et al. 2010). Catalases are present almost in all aerobically respiring organisms. Within the cell environment, catalases are localized in all major sites of H2O2 production, mostly at peroxisomes, but they were detected in cytosol, mitochondria, and chloroplasts as well (Sharma and Ahmad 2014). Catalases (CAT, 1.11.1.6) are antioxidant enzymes that catalyse decomposition of hydrogen peroxide to water and molecular oxygen (2H2O2 → 2H2O + O2), an important process in defending cells against oxidative damage caused by reactive oxygen species (ROS; Alfonso-Prieto et al. -

BRENDA Tutorial
BRENDA Tutorial Introduction to the Enzyme Information System Facts about BRENDA (BRaunschweig ENzyme DAtabase) • one of the most comprehensive enzyme information repositories • BRENDA is a member of de.NBI (German Network for Bioinformatics Structure, since 2015) • BRENDA is an ELIXIR Core Data Resource (since 2018) • all enzymes, classified by the Enzyme Nomenclature (IUBMB) • data of molecular biology, biochemistry, medical research, and biotechnology • furthermore BRENDA includes data from interconnected databases containing results from text mining methods and bioinformatic approaches • BRENDA is freely available to the scientific community • more than 80,000 visits of the BRENDA website each month • major updates of the data in BRENDA are performed twice a year History and major developments of BRENDA • BRENDA was created at the former German National Research Center for Biotechnology (GBF, now HZI, Helmholtz Zentrum für Infektionsforschung, Braunschweig, Germany) in 1987 • BRENDA was originally published as a series of book o 1st Edition 1990-1997 (Enzyme Handbook) o 2nd Edition 2001-2013 (Handbook of Enzymes) • BRENDA moved to the University of Cologne, Germany • First online version in 1998 via the SRS system at the EBI • First website of BRENDA in Cologne • Transfer of BRENDA into a fully relational database system • BRENDA moved back to Braunschweig in 2007 • BRENDA is now maintained and further developed at the BRICS - TU Braunschweig Facts about BRENDA The main categories are based on the Enzymes and the Metabolites / Ligands Enzyme-related data encompasses information on: • Enzyme and ligand nomenclature • Organism • Reaction and specificity • Kinetic properties • Structure and role of the ligands • Stability information • Ligand-enzyme information • Enzyme sequence and structure • Mutants and disease • Occurrence, isolation, and preparation • Pathways BRENDA is the most comprehensive information system on: • 7862 EC Numbers (July 2019) • more than 2 Mill. -

Supporting Information High-Throughput Virtual Screening
Supporting Information High-Throughput Virtual Screening of Proteins using GRID Molecular Interaction Fields Simone Sciabola, Robert V. Stanton, James E. Mills, Maria M. Flocco, Massimo Baroni, Gabriele Cruciani, Francesca Perruccio and Jonathan S. Mason Contents Table S1 S2-S21 Figure S1 S22 * To whom correspondence should be addressed: Simone Sciabola, Pfizer Research Technology Center, Cambridge, 02139 MA, USA Phone: +1-617-551-3327; Fax: +1-617-551-3117; E-mail: [email protected] S1 Table S1. Description of the 990 proteins used as decoy for the Protein Virtual Screening analysis. PDB ID Protein family Molecule Res. (Å) 1n24 ISOMERASE (+)-BORNYL DIPHOSPHATE SYNTHASE 2.3 1g4h HYDROLASE 1,3,4,6-TETRACHLORO-1,4-CYCLOHEXADIENE HYDROLASE 1.8 1cel HYDROLASE(O-GLYCOSYL) 1,4-BETA-D-GLUCAN CELLOBIOHYDROLASE I 1.8 1vyf TRANSPORT PROTEIN 14 KDA FATTY ACID BINDING PROTEIN 1.85 1o9f PROTEIN-BINDING 14-3-3-LIKE PROTEIN C 2.7 1t1s OXIDOREDUCTASE 1-DEOXY-D-XYLULOSE 5-PHOSPHATE REDUCTOISOMERASE 2.4 1t1r OXIDOREDUCTASE 1-DEOXY-D-XYLULOSE 5-PHOSPHATE REDUCTOISOMERASE 2.3 1q0q OXIDOREDUCTASE 1-DEOXY-D-XYLULOSE 5-PHOSPHATE REDUCTOISOMERASE 1.9 1jcy LYASE 2-DEHYDRO-3-DEOXYPHOSPHOOCTONATE ALDOLASE 1.9 1fww LYASE 2-DEHYDRO-3-DEOXYPHOSPHOOCTONATE ALDOLASE 1.85 1uk7 HYDROLASE 2-HYDROXY-6-OXO-7-METHYLOCTA-2,4-DIENOATE 1.7 1v11 OXIDOREDUCTASE 2-OXOISOVALERATE DEHYDROGENASE ALPHA SUBUNIT 1.95 1x7w OXIDOREDUCTASE 2-OXOISOVALERATE DEHYDROGENASE ALPHA SUBUNIT 1.73 1d0l TRANSFERASE 35KD SOLUBLE LYTIC TRANSGLYCOSYLASE 1.97 2bt4 LYASE 3-DEHYDROQUINATE DEHYDRATASE -

Chorismate Mutase and Isochorismatase, Two Potential
Received: 28 June 2020 | Revised: 7 September 2020 | Accepted: 7 September 2020 DOI: 10.1111/mpp.13003 ORIGINAL ARTICLE Chorismate mutase and isochorismatase, two potential effectors of the migratory nematode Hirschmanniella oryzae, increase host susceptibility by manipulating secondary metabolite content of rice Lander Bauters 1 | Tina Kyndt 1 | Tim De Meyer2 | Kris Morreel3,4 | Wout Boerjan3,4 | Hannes Lefevere1 | Godelieve Gheysen 1 1Department of Biotechnology, Faculty of Bioscience Engineering, Ghent University, Abstract Ghent, Belgium Hirschmanniella oryzae is one of the most devastating nematodes on rice, leading to 2 Department of Data Analysis and substantial yield losses. Effector proteins aid the nematode during the infection pro- Mathematical Modelling, Faculty of Bioscience Engineering, Ghent University, cess by subduing plant defence responses. In this research we characterized two po- Ghent, Belgium tential H. oryzae effector proteins, chorismate mutase (HoCM) and isochorismatase 3VIB-UGent Center for Plant Systems Biology, Ghent, Belgium (HoICM), and investigated their enzymatic activity and their role in plant immunity. 4Department of Plant Biotechnology and Both HoCM and HoICM proved to be enzymatically active in complementation tests Bioinformatics, Faculty of Sciences, Ghent in mutant Escherichia coli strains. Infection success by the migratory nematode H. ory- University, Ghent, Belgium zae was significantly higher in transgenic rice lines constitutively expressing HoCM Correspondence or HoICM. Expression of HoCM, but not HoICM, increased rice susceptibility against Godelieve Gheysen, Department of Biotechnology, Faculty of Bioscience the sedentary nematode Meloidogyne graminicola also. Transcriptome and metabo- Engineering, Ghent University, Ghent, lome analyses indicated reductions in secondary metabolites in the transgenic rice Belgium. Email: [email protected] plants expressing the potential nematode effectors. -

Biological Role of Conceptus Derived Factors During Early Pregnancy In
Biological Role of Conceptus Derived Factors During Early Pregnancy in Ruminants A dissertation submitted in partial fulfillment of the requirements for the degree of DOCTOR OF PHILOSOPHY IN ANIMAL SCIENCES UNIVERSITY OF MISSOURI- COLUMBIA Division of Animal Science By KELSEY BROOKS Dr. Thomas Spencer, Dissertation Supervisor August 2016 The undersigned have examined the dissertation entitled, BIOLOGICAL ROLE OF CONCEPTUS DERIVED FACTORS DURING EARLY PREGNANCY IN RUMINANTS presented by Kelsey Brooks, a candidate for the degree of doctor of philosophy, and hereby certify that, in their opinion, it is worthy of acceptance. __________________________________ Chair, Dr. Thomas Spencer ___________________________________ Dr. Rodney Geisert ___________________________________ Dr. Randall Prather ___________________________________ Dr. Laura Schulz ACKNOWLEDGMENTS I would like to acknowledge all the students, faculty and staff at Washington State University and the University of Missouri for their help and support throughout my doctoral program. I am grateful for the opportunity to work with Dr. Thomas Spencer, and thank him for his input and guidance not only in planning experiments and completing projects but for helping me turn my love of science into a career in research. I would also like to acknowledge the members of my graduate committee at Washington State University for their help and input during the first 3 years of my studies. A special thanks to Dr. Jim Pru and Cindy Pru for providing unlimited entertainment, and the occasional missing reagent. Thank you to my committee members at the University of Missouri for adopting me late in my program and helping shape my future as an independent scientist. Thanks are also extended to members of the Prather lab and Wells lab for letting me in on the secrets of success using the CRISPR/Cas9 system. -
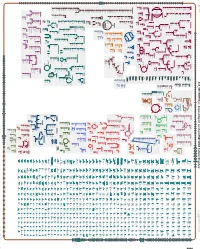
Generate Metabolic Map Poster
Authors: Pallavi Subhraveti Ron Caspi Quang Ong Peter D Karp An online version of this diagram is available at BioCyc.org. Biosynthetic pathways are positioned in the left of the cytoplasm, degradative pathways on the right, and reactions not assigned to any pathway are in the far right of the cytoplasm. Transporters and membrane proteins are shown on the membrane. Ingrid Keseler Periplasmic (where appropriate) and extracellular reactions and proteins may also be shown. Pathways are colored according to their cellular function. Gcf_900114035Cyc: Amycolatopsis sacchari DSM 44468 Cellular Overview Connections between pathways are omitted for legibility. -

The Estimation of Kinetic Parameters in Systems Biology by Comparing Molecular Interaction Fields of Enzymes
237 Beilstein-Institut ESCEC, March 19th –23rd, 2006, Rdesheim/Rhein, Germany The Estimation of Kinetic Parameters in Systems Biology by Comparing Molecular Interaction Fields of Enzymes Matthias Stein, Razif R. Gabdoulline, Bruno Besson, Rebecca C. Wade EML Research gGmbH, Molecular and Cellular Modeling Group, Schloss-Wolfsbrunnenweg 33, 69118 Heidelberg, Germany E-Mail: [email protected] Received: 16th June 2006 / Published: 31st August 2007 Abstract The kinetic modelling of biochemical pathways requires a consistent set of enzymatic kinetic parameters. We report results from software development to assist the user in systems biology, allowing the retrie- val of heterogeneous protein sequence, structural and kinetic data. For the simulation of biological networks, missing enzymatic kinetic para- meters can be calculated using a similarity analysis of the enzymes’ molecular interaction fields. The quantitative PIPSA (qPIPSA) meth- odology relates changes in the molecular interaction fields of the enzymes with variations in the enzymatic rate constants or binding affinities. As an illustrative example, this approach is used to predict kinetic parameters for glucokinases from Escherichia coli based on experimental values for a test set of enzymes. The best correlation of the electrostatic potentials with kinetic parameters is found for the open form of the glucokinases. The similarity analysis was extended to a large set of glucokinases from various organisms. http://www.beilstein-institut.de/escec2006/proceedings/Stein/Stein.pdf 238 Stein, M. et al. Introduction One of the aims of systems biology is to provide a mathematical description of metabolic or signalling protein networks. This can be achieved by constructing a set of differential equations describing changes in concentrations of compounds with time [1]. -

The Crystal Structure of Novel Chondroitin Lyase ODV-E66, A
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector FEBS Letters 587 (2013) 3943–3948 journal homepage: www.FEBSLetters.org The crystal structure of novel chondroitin lyase ODV-E66, a baculovirus envelope protein ⇑ Yoshirou Kawaguchi a, Nobuo Sugiura b, Koji Kimata c, Makoto Kimura a,d, Yoshimitsu Kakuta a,d, a Laboratory of Structural Biology, Graduate School of System Life Sciences, Kyushu University, 6-10-1 Hakozaki, Fukuoka 812-8581, Japan b Institute for Molecular Science of Medicine, Aichi Medical University, 1-1 Yazakokarimata, Nagakute, Aichi 480-1195, Japan c Research Complex for the Medicine Frontiers, Aichi Medical University, 1-1 Yazakokarimata, Nagakute, Aichi 480-1195, Japan d Faculty of Agriculture, Kyushu University, 6-10-1 Hakozaki, Fukuoka 812-8581, Japan article info abstract Article history: Chondroitin lyases have been known as pathogenic bacterial enzymes that degrade chondroitin. Received 5 August 2013 Recently, baculovirus envelope protein ODV-E66 was identified as the first reported viral chondroi- Revised 1 October 2013 tin lyase. ODV-E66 has low sequence identity with bacterial lyases at <12%, and unique characteris- Accepted 15 October 2013 tics reflecting the life cycle of baculovirus. To understand ODV-E66’s structural basis, the crystal Available online 26 October 2013 structure was determined and it was found that the structural fold resembled that of polysaccharide Edited by Christian Griesinger lyase 8 proteins and that the catalytic residues were also conserved. This structure enabled discus- sion of the unique substrate specificity and the stability of ODV-E66 as well as the host specificity of baculovirus.