Positive Selection in Transcription Factor Genes
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

ATRX Induction by Mutant Huntingtin Via Cdx2 Modulates Heterochromatin Condensation and Pathology in Huntington’S Disease
Cell Death and Differentiation (2012) 19, 1109–1116 & 2012 Macmillan Publishers Limited All rights reserved 1350-9047/12 www.nature.com/cdd ATRX induction by mutant huntingtin via Cdx2 modulates heterochromatin condensation and pathology in Huntington’s disease J Lee1,2, YK Hong3, GS Jeon4, YJ Hwang4, KY Kim4, KH Seong4, M-K Jung4, DJ Picketts5, NW Kowall1,2, KS Cho3 and H Ryu*,1,2,4 Aberrant chromatin remodeling is involved in the pathogenesis of Huntington’s disease (HD) but the mechanism is not known. Herein, we report that mutant huntingtin (mtHtt) induces the transcription of alpha thalassemia/mental retardation X linked (ATRX), an ATPase/helicase and SWI/SNF-like chromatin remodeling protein via Cdx-2 activation. ATRX expression was elevated in both a cell line model and transgenic model of HD, and Cdx-2 occupancy of the ATRX promoter was increased in HD. Induction of ATRX expanded the size of promyelocytic leukemia nuclear body (PML-NB) and increased trimethylation of H3K9 (H3K9me3) and condensation of pericentromeric heterochromatin, while knockdown of ATRX decreased PML-NB and H3K9me3 levels. Knockdown of ATRX/dXNP improved the hatch rate of fly embryos expressing mtHtt (Q127). ATRX/dXNP overexpression exacerbated eye degeneration of eye-specific mtHtt (Q127) expressing flies. Our findings suggest that transcriptional alteration of ATRX by mtHtt is involved in pericentromeric heterochromatin condensation and contributes to the pathogenesis of HD. Cell Death and Differentiation (2012) 19, 1109–1116; doi:10.1038/cdd.2011.196; published -

Isyte: Integrated Systems Tool for Eye Gene Discovery
Lens iSyTE: Integrated Systems Tool for Eye Gene Discovery Salil A. Lachke,1,2,3,4 Joshua W. K. Ho,1,4,5 Gregory V. Kryukov,1,4,6 Daniel J. O’Connell,1 Anton Aboukhalil,1,7 Martha L. Bulyk,1,8,9 Peter J. Park,1,5,10 and Richard L. Maas1 PURPOSE. To facilitate the identification of genes associated ther investigation. Extension of this approach to other ocular with cataract and other ocular defects, the authors developed tissue components will facilitate eye disease gene discovery. and validated a computational tool termed iSyTE (integrated (Invest Ophthalmol Vis Sci. 2012;53:1617–1627) DOI: Systems Tool for Eye gene discovery; http://bioinformatics. 10.1167/iovs.11-8839 udel.edu/Research/iSyTE). iSyTE uses a mouse embryonic lens gene expression data set as a bioinformatics filter to select candidate genes from human or mouse genomic regions impli- ven with the advent of high-throughput sequencing, the cated in disease and to prioritize them for further mutational Ediscovery of genes associated with congenital birth defects and functional analyses. such as eye defects remains a challenge. We sought to develop METHODS. Microarray gene expression profiles were obtained a straightforward experimental approach that could facilitate for microdissected embryonic mouse lens at three key devel- the identification of candidate genes for developmental disor- opmental time points in the transition from the embryonic day ders, and, as proof-of-principle, we chose defects involving the (E)10.5 stage of lens placode invagination to E12.5 lens primary ocular lens. Opacification of the lens results in cataract, a leading cause of blindness that affects 77 million persons and fiber cell differentiation. -

Faith and the Human Genome
Plenary Presenters Faith and the Human Genome Faith and the Human Genome Francis S. Collins Despite the best efforts of the American Scientific Affiliation to bridge the gap between science and faith, few gatherings of scientists involved in biology include any meaningful discussion about the spiritual significance of the current revolution in genetics and genomics. Most biologists and geneticists seem to have concluded that science and faith are incompatible, but few who embrace that conclusion seem to have seriously considered the evidence. From my perspective as director of the Human Genome Project, the scientific and religious world views are not only compatible but also inherently complementary. Hence the profound polarization of the scientific and religious perspectives, now glaringly apparent in the fields of biology and genetics, is a source of great distress. Hard-liners in either camp paint increasingly uncompromising pictures that force sincere seekers to choose one view over the other. How all of this must break God’s heart! The elegance and complexity of the human genome is a source of profound wonder. That wonder only strengthens my faith, as it provides glimpses of aspects of From my humanity, which God has known all along, but which we are just now beginning to discover. perspective as e are just on the edge of a whole You made him a little lower than the heav- director of the Whost of developments spurred on enly beings and crowned him with glory by genetics that are going to and honor. You made him ruler over the Human require careful and deliberative thought. -

Protein Interaction Network of Alternatively Spliced Isoforms from Brain Links Genetic Risk Factors for Autism
ARTICLE Received 24 Aug 2013 | Accepted 14 Mar 2014 | Published 11 Apr 2014 DOI: 10.1038/ncomms4650 OPEN Protein interaction network of alternatively spliced isoforms from brain links genetic risk factors for autism Roser Corominas1,*, Xinping Yang2,3,*, Guan Ning Lin1,*, Shuli Kang1,*, Yun Shen2,3, Lila Ghamsari2,3,w, Martin Broly2,3, Maria Rodriguez2,3, Stanley Tam2,3, Shelly A. Trigg2,3,w, Changyu Fan2,3, Song Yi2,3, Murat Tasan4, Irma Lemmens5, Xingyan Kuang6, Nan Zhao6, Dheeraj Malhotra7, Jacob J. Michaelson7,w, Vladimir Vacic8, Michael A. Calderwood2,3, Frederick P. Roth2,3,4, Jan Tavernier5, Steve Horvath9, Kourosh Salehi-Ashtiani2,3,w, Dmitry Korkin6, Jonathan Sebat7, David E. Hill2,3, Tong Hao2,3, Marc Vidal2,3 & Lilia M. Iakoucheva1 Increased risk for autism spectrum disorders (ASD) is attributed to hundreds of genetic loci. The convergence of ASD variants have been investigated using various approaches, including protein interactions extracted from the published literature. However, these datasets are frequently incomplete, carry biases and are limited to interactions of a single splicing isoform, which may not be expressed in the disease-relevant tissue. Here we introduce a new interactome mapping approach by experimentally identifying interactions between brain-expressed alternatively spliced variants of ASD risk factors. The Autism Spliceform Interaction Network reveals that almost half of the detected interactions and about 30% of the newly identified interacting partners represent contribution from splicing variants, emphasizing the importance of isoform networks. Isoform interactions greatly contribute to establishing direct physical connections between proteins from the de novo autism CNVs. Our findings demonstrate the critical role of spliceform networks for translating genetic knowledge into a better understanding of human diseases. -

The Expression of the Human Apolipoprotein Genes and Their Regulation by Ppars
CORE Metadata, citation and similar papers at core.ac.uk Provided by UEF Electronic Publications The expression of the human apolipoprotein genes and their regulation by PPARs Juuso Uski M.Sc. Thesis Biochemistry Department of Biosciences University of Kuopio June 2008 Abstract The expression of the human apolipoprotein genes and their regulation by PPARs. UNIVERSITY OF KUOPIO, the Faculty of Natural and Environmental Sciences, Curriculum of Biochemistry USKI Juuso Oskari Thesis for Master of Science degree Supervisors Prof. Carsten Carlberg, Ph.D. Merja Heinäniemi, Ph.D. June 2008 Keywords: nuclear receptors; peroxisome proliferator-activated receptor; PPAR response element; apolipoprotein; lipid metabolism; high density lipoprotein; low density lipoprotein. Lipids are any fat-soluble, naturally-occurring molecules and one of their main biological functions is energy storage. Lipoproteins carry hydrophobic lipids in the water and salt-based blood environment for processing and energy supply in liver and other organs. In this study, the genomic area around the apolipoprotein genes was scanned in silico for PPAR response elements (PPREs) using the in vitro data-based computer program. Several new putative REs were found in surroundings of multiple lipoprotein genes. The responsiveness of those apolipoprotein genes to the PPAR ligands GW501516, rosiglitazone and GW7647 in the HepG2, HEK293 and THP-1 cell lines were tested with real-time PCR. The APOA1, APOA2, APOB, APOD, APOE, APOF, APOL1, APOL3, APOL5 and APOL6 genes were found to be regulated by PPARs in direct or secondary manners. Those results provide new insights in the understanding of lipid metabolism and so many lifestyle diseases like atherosclerosis, type 2 diabetes, heart disease and stroke. -

Seq2pathway Vignette
seq2pathway Vignette Bin Wang, Xinan Holly Yang, Arjun Kinstlick May 19, 2021 Contents 1 Abstract 1 2 Package Installation 2 3 runseq2pathway 2 4 Two main functions 3 4.1 seq2gene . .3 4.1.1 seq2gene flowchart . .3 4.1.2 runseq2gene inputs/parameters . .5 4.1.3 runseq2gene outputs . .8 4.2 gene2pathway . 10 4.2.1 gene2pathway flowchart . 11 4.2.2 gene2pathway test inputs/parameters . 11 4.2.3 gene2pathway test outputs . 12 5 Examples 13 5.1 ChIP-seq data analysis . 13 5.1.1 Map ChIP-seq enriched peaks to genes using runseq2gene .................... 13 5.1.2 Discover enriched GO terms using gene2pathway_test with gene scores . 15 5.1.3 Discover enriched GO terms using Fisher's Exact test without gene scores . 17 5.1.4 Add description for genes . 20 5.2 RNA-seq data analysis . 20 6 R environment session 23 1 Abstract Seq2pathway is a novel computational tool to analyze functional gene-sets (including signaling pathways) using variable next-generation sequencing data[1]. Integral to this tool are the \seq2gene" and \gene2pathway" components in series that infer a quantitative pathway-level profile for each sample. The seq2gene function assigns phenotype-associated significance of genomic regions to gene-level scores, where the significance could be p-values of SNPs or point mutations, protein-binding affinity, or transcriptional expression level. The seq2gene function has the feasibility to assign non-exon regions to a range of neighboring genes besides the nearest one, thus facilitating the study of functional non-coding elements[2]. Then the gene2pathway summarizes gene-level measurements to pathway-level scores, comparing the quantity of significance for gene members within a pathway with those outside a pathway. -

Dual Proteome-Scale Networks Reveal Cell-Specific Remodeling of the Human Interactome
bioRxiv preprint doi: https://doi.org/10.1101/2020.01.19.905109; this version posted January 19, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Dual Proteome-scale Networks Reveal Cell-specific Remodeling of the Human Interactome Edward L. Huttlin1*, Raphael J. Bruckner1,3, Jose Navarrete-Perea1, Joe R. Cannon1,4, Kurt Baltier1,5, Fana Gebreab1, Melanie P. Gygi1, Alexandra Thornock1, Gabriela Zarraga1,6, Stanley Tam1,7, John Szpyt1, Alexandra Panov1, Hannah Parzen1,8, Sipei Fu1, Arvene Golbazi1, Eila Maenpaa1, Keegan Stricker1, Sanjukta Guha Thakurta1, Ramin Rad1, Joshua Pan2, David P. Nusinow1, Joao A. Paulo1, Devin K. Schweppe1, Laura Pontano Vaites1, J. Wade Harper1*, Steven P. Gygi1*# 1Department of Cell Biology, Harvard Medical School, Boston, MA, 02115, USA. 2Broad Institute, Cambridge, MA, 02142, USA. 3Present address: ICCB-Longwood Screening Facility, Harvard Medical School, Boston, MA, 02115, USA. 4Present address: Merck, West Point, PA, 19486, USA. 5Present address: IQ Proteomics, Cambridge, MA, 02139, USA. 6Present address: Vor Biopharma, Cambridge, MA, 02142, USA. 7Present address: Rubius Therapeutics, Cambridge, MA, 02139, USA. 8Present address: RPS North America, South Kingstown, RI, 02879, USA. *Correspondence: [email protected] (E.L.H.), [email protected] (J.W.H.), [email protected] (S.P.G.) #Lead Contact: [email protected] bioRxiv preprint doi: https://doi.org/10.1101/2020.01.19.905109; this version posted January 19, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. -

Parallel Molecular Evolution in Pathways, Genes, and Sites in High-Elevation Hummingbirds Revealed by Comparative Transcriptomics
GBE Parallel Molecular Evolution in Pathways, Genes, and Sites in High-Elevation Hummingbirds Revealed by Comparative Transcriptomics Marisa C.W. Lim1,*, Christopher C. Witt2, Catherine H. Graham1,3,andLilianaM.Davalos 1,4 1Department of Ecology and Evolution, Stony Brook University 2 Museum of Southwestern Biology and Department of Biology, University of New Mexico Downloaded from https://academic.oup.com/gbe/article-abstract/11/6/1552/5494706 by guest on 08 June 2019 3Swiss Federal Research Institute (WSL), Birmensdorf, Switzerland 4Consortium for Inter-Disciplinary Environmental Research, Stony Brook University *Corresponding author: E-mail: [email protected]. Accepted: May 12, 2019 Data deposition: The raw read data have been deposited in the NCBI Sequence Read Archive under BioProject: PRJNA543673, BioSample: SAMN11774663-SAMN11774674, SRA Study: SRP198856. All scripts used for analyses are available on Dryad: doi:10.5061/dryad.v961mb4. Abstract High-elevation organisms experience shared environmental challenges that include low oxygen availability, cold temperatures, and intense ultraviolet radiation. Consequently, repeated evolution of the same genetic mechanisms may occur across high-elevation taxa. To test this prediction, we investigated the extent to which the same biochemical pathways, genes, or sites were subject to parallel molecular evolution for 12 Andean hummingbird species (family: Trochilidae) representing several independent transitions to high elevation across the phylogeny. Across high-elevation species, we discovered parallel evolution for several pathways and genes with evidence of positive selection. In particular, positively selected genes were frequently part of cellular respiration, metabolism, or cell death pathways. To further examine the role of elevation in our analyses, we compared results for low- and high-elevation species and tested different thresholds for defining elevation categories. -

Evidence for Differential Alternative Splicing in Blood of Young Boys With
Stamova et al. Molecular Autism 2013, 4:30 http://www.molecularautism.com/content/4/1/30 RESEARCH Open Access Evidence for differential alternative splicing in blood of young boys with autism spectrum disorders Boryana S Stamova1,2,5*, Yingfang Tian1,2,4, Christine W Nordahl1,3, Mark D Shen1,3, Sally Rogers1,3, David G Amaral1,3 and Frank R Sharp1,2 Abstract Background: Since RNA expression differences have been reported in autism spectrum disorder (ASD) for blood and brain, and differential alternative splicing (DAS) has been reported in ASD brains, we determined if there was DAS in blood mRNA of ASD subjects compared to typically developing (TD) controls, as well as in ASD subgroups related to cerebral volume. Methods: RNA from blood was processed on whole genome exon arrays for 2-4–year-old ASD and TD boys. An ANCOVA with age and batch as covariates was used to predict DAS for ALL ASD (n=30), ASD with normal total cerebral volumes (NTCV), and ASD with large total cerebral volumes (LTCV) compared to TD controls (n=20). Results: A total of 53 genes were predicted to have DAS for ALL ASD versus TD, 169 genes for ASD_NTCV versus TD, 1 gene for ASD_LTCV versus TD, and 27 genes for ASD_LTCV versus ASD_NTCV. These differences were significant at P <0.05 after false discovery rate corrections for multiple comparisons (FDR <5% false positives). A number of the genes predicted to have DAS in ASD are known to regulate DAS (SFPQ, SRPK1, SRSF11, SRSF2IP, FUS, LSM14A). In addition, a number of genes with predicted DAS are involved in pathways implicated in previous ASD studies, such as ROS monocyte/macrophage, Natural Killer Cell, mTOR, and NGF signaling. -
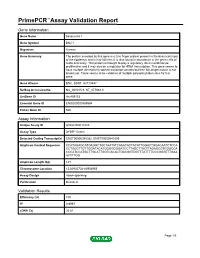
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name basonuclin 1 Gene Symbol BNC1 Organism Human Gene Summary The protein encoded by this gene is a zinc finger protein present in the basal cell layer of the epidermis and in hair follicles. It is also found in abundance in the germ cells of testis and ovary. This protein is thought to play a regulatory role in keratinocyte proliferation and it may also be a regulator for rRNA transcription. This gene seems to have multiple alternatively spliced transcript variants but their full-length nature is not known yet. There seems to be evidence of multiple polyadenylation sites for this gene. Gene Aliases BNC, BSN1, HsT19447 RefSeq Accession No. NC_000015.9, NT_077661.3 UniGene ID Hs.459153 Ensembl Gene ID ENSG00000169594 Entrez Gene ID 646 Assay Information Unique Assay ID qHsaCID0017223 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000345382, ENST00000541809 Amplicon Context Sequence CCATAGAGCATGAGGCTGCTAATATCAAACACTACATTGGACTGGACAATCTCCA CCTGGCTTGTTGGATACATGGGGGGGATCCTTAGCTTACTTAGAGCGTGGGCCA CCCATCCATGCTTGCATTGGTCACACTGACGGTGGTTTATTTTCCCGGGTTTGAA ACTTTGG Amplicon Length (bp) 141 Chromosome Location 15:83935724-83936955 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 100 R2 0.9997 cDNA Cq 26.81 Page 1/5 PrimePCR™Assay Validation Report cDNA Tm (Celsius) 84.5 gDNA Cq 39.42 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report BNC1, Human Amplification -

The Porcine Major Histocompatibility Complex and Related Paralogous Regions: a Review Patrick Chardon, Christine Renard, Claire Gaillard, Marcel Vaiman
The porcine Major Histocompatibility Complex and related paralogous regions: a review Patrick Chardon, Christine Renard, Claire Gaillard, Marcel Vaiman To cite this version: Patrick Chardon, Christine Renard, Claire Gaillard, Marcel Vaiman. The porcine Major Histocom- patibility Complex and related paralogous regions: a review. Genetics Selection Evolution, BioMed Central, 2000, 32 (2), pp.109-128. 10.1051/gse:2000101. hal-00894302 HAL Id: hal-00894302 https://hal.archives-ouvertes.fr/hal-00894302 Submitted on 1 Jan 2000 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Genet. Sel. Evol. 32 (2000) 109–128 109 c INRA, EDP Sciences Review The porcine Major Histocompatibility Complex and related paralogous regions: a review Patrick CHARDON, Christine RENARD, Claire ROGEL GAILLARD, Marcel VAIMAN Laboratoire de radiobiologie et d’etude du genome, Departement de genetique animale, Institut national de la recherche agronomique, Commissariat al’energie atomique, 78352, Jouy-en-Josas Cedex, France (Received 18 November 1999; accepted 17 January 2000) Abstract – The physical alignment of the entire region of the pig major histocompat- ibility complex (MHC) has been almost completed. In swine, the MHC is called the SLA (swine leukocyte antigen) and most of its class I region has been sequenced. -

Learning Protein Constitutive Motifs from Sequence Data Je´ Roˆ Me Tubiana, Simona Cocco, Re´ Mi Monasson*
TOOLS AND RESOURCES Learning protein constitutive motifs from sequence data Je´ roˆ me Tubiana, Simona Cocco, Re´ mi Monasson* Laboratory of Physics of the Ecole Normale Supe´rieure, CNRS UMR 8023 & PSL Research, Paris, France Abstract Statistical analysis of evolutionary-related protein sequences provides information about their structure, function, and history. We show that Restricted Boltzmann Machines (RBM), designed to learn complex high-dimensional data and their statistical features, can efficiently model protein families from sequence information. We here apply RBM to 20 protein families, and present detailed results for two short protein domains (Kunitz and WW), one long chaperone protein (Hsp70), and synthetic lattice proteins for benchmarking. The features inferred by the RBM are biologically interpretable: they are related to structure (residue-residue tertiary contacts, extended secondary motifs (a-helixes and b-sheets) and intrinsically disordered regions), to function (activity and ligand specificity), or to phylogenetic identity. In addition, we use RBM to design new protein sequences with putative properties by composing and ’turning up’ or ’turning down’ the different modes at will. Our work therefore shows that RBM are versatile and practical tools that can be used to unveil and exploit the genotype–phenotype relationship for protein families. DOI: https://doi.org/10.7554/eLife.39397.001 Introduction In recent years, the sequencing of many organisms’ genomes has led to the collection of a huge number of protein sequences, which are catalogued in databases such as UniProt or PFAM Finn et al., 2014). Sequences that share a common ancestral origin, defining a family (Figure 1A), *For correspondence: are likely to code for proteins with similar functions and structures, providing a unique window into [email protected] the relationship between genotype (sequence content) and phenotype (biological features).