Snow Leopard Survival Strategy
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
Module No. 1840 1840-1
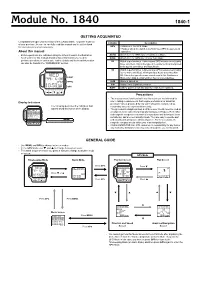
Module No. 1840 1840-1 GETTING ACQUAINTED Congratulations upon your selection of this CASIO watch. To get the most out Indicator Description of your purchase, be sure to carefully read this manual and keep it on hand for later reference when necessary. GPS • Watch is in the GPS Mode. • Flashes when the watch is performing a GPS measurement About this manual operation. • Button operations are indicated using the letters shown in the illustration. AUTO Watch is in the GPS Auto or Continuous Mode. • Each section of this manual provides basic information you need to SAVE Watch is in the GPS One-shot or Auto Mode. perform operations in each mode. Further details and technical information 2D Watch is performing a 2-dimensional GPS measurement (using can also be found in the “REFERENCE” section. three satellites). This is the type of measurement normally used in the Quick, One-Shot, and Auto Mode. 3D Watch is performing a 3-dimensional GPS measurement (using four or more satellites), which provides better accuracy than 2D. This is the type of measurement used in the Continuous LIGHT Mode when data is obtained from four or more satellites. MENU ALM Alarm is turned on. SIG Hourly Time Signal is turned on. GPS BATT Battery power is low and battery needs to be replaced. Precautions • The measurement functions built into this watch are not intended for Display Indicators use in taking measurements that require professional or industrial precision. Values produced by this watch should be considered as The following describes the indicators that reasonably accurate representations only. -

Leopard Geckos
Husbandry Handbook LEOPARD GECKOS Eublepharus macularius The Exception to the Rule Temperature and Lighting When dening what makes a gecko different from a lizard, there are a few things It is important to create a thermal gradient (a warm and a cool side) in the that come to mind right away. First, geckos have sticky toe pads that enable them cage/enclosure. This can be done with an appropriate sized Zilla® Heat Mat to climb. Second, they don’t have eye lids and have to lick their eyes to clean them. adhered to the bottom of the tank all the way to one side. Ideal temperatures for Lastly, they have vocal cords that allow them to bark and make noises. Leopard Leopard Geckos range from 75-80°F on the cool side and 80-85°F on the warm Geckos are unusual in that they don’t have sticky toe pads and they have eyelids. side. Provide a 90-95°F basking area on the warm side. While Leopard Geckos They do, however, have vocal cords and can squeak and bark to ward off predators. don’t need UVB to survive, UVA/UVB light has been shown to greatly improve the While exceptions to the normal gecko rules, they make amazing rst pet reptiles. immune system, health, and wellness of all reptiles, both diurnal and crepuscular. They are docile, easy to handle and very hardy. With 30 years of selective breeding, Using a Zilla® Mini Heat & UVB Fixture with a Zilla® 50W Mini Halogen bulb and a they now come in a wide variety of colors and patterns. -

Snow Leopard Survival Strategy 2014
Snow Leopard Survival Strategy Revised Version 2014.1 Snow Leopard Network 1 The designation of geographical entities in this book, and the presentation of the material, do not imply the expression of any opinion whatsoever on the part of the Snow Leopard Network concerning the legal status of any country, territory, or area, or of its authorities, or concerning the delimitation of its frontiers or boundaries. Copyright: © 2014 Snow Leopard Network, 4649 Sunnyside Ave. N. Suite 325, Seattle, WA 98103. Reproduction of this publication for educational or other non-commercial purposes is authorised without prior written permission from the copyright holder provided the source is fully acknowledged. Reproduction of this publication for resale or other commercial purposes is prohibited without prior written permission of the copyright holder. Citation: Snow Leopard Network (2014). Snow Leopard Survival Strategy. Revised 2014 Version Snow Leopard Network, Seattle, Washington, USA. Website: http://www.snowleopardnetwork.org/ The Snow Leopard Network is a worldwide organization dedicated to facilitating the exchange of information between individuals around the world for the purpose of snow leopard conservation. Our membership includes leading snow leopard experts in the public, private, and non-profit sectors. The main goal of this organization is to implement the Snow Leopard Survival Strategy (SLSS) which offers a comprehensive analysis of the issues facing snow leopard conservation today. Cover photo: Camera-trapped snow leopard. © Snow Leopard -

Reconstruction of Phylogenetic History to Resolve the Subspecies Anomaly of Pantherine Cats
bioRxiv preprint doi: https://doi.org/10.1101/082891; this version posted October 24, 2016. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Title: Reconstruction of phylogenetic history to resolve the subspecies anomaly of Pantherine cats Authors: Ranajit Das1, Priyanka Upadhyai2 1Manipal Centre for Natural Sciences, Manipal University, Manipal, Karnataka 576104, India 2Department of Medical Genetics, Kasturba Medical College, Manipal University, Manipal, Karnataka 576104, India Email address: [email protected] Running title: Mitogenome phylogeny of Pantherine cats Abstract All charismatic big cats including tiger (Panthera tigris), lion (Panthera leo), leopard (Panthera pardus), snow leopard (Panthera uncial), and jaguar (Panthera onca) are grouped into the subfamily Pantherinae. Several mitogenomic approaches have been employed to reconstruct the phylogenetic history of the Pantherine cats but the phylogeny has remained largely unresolved till date. One of the major reasons for the difficulty in resolving the phylogenetic tree of Pantherine cats is the small sample size. While previous studies included only 5-10 samples, we have used 43 publically available taxa to reconstruct Pantherine phylogenetic history. Complete mtDNA sequences were used from all individuals excluding the control region (15,489bp). A Bayesian MCMC approach was employed to investigate the divergence times among different Pantherine clades. Both maximum likelihood and Bayesian phylogeny generated a dendrogram: Neofelis nebulosa (Panthera tigris (Panthera onca (Panthera uncia (Panthera leo, Panthera pardus)))), grouping lions with leopards and placing snow leopards as an outgroup to this clade. -

An Ounce of Prevention: Snow Leopard Crime Revisited (PDF, 4
TRAFFIC AN OUNCE REPORT OF PREVENTION: Snow Leopard Crime Revisited OCTOBER 2016 Kristin Nowell, Juan Li, Mikhail Paltsyn and Rishi Kumar Sharma TRAFFIC REPORT TRAFFIC, the wild life trade monitoring net work, is the leading non-governmental organization working globally on trade in wild animals and plants in the context of both biodiversity conservation and sustainable development. TRAFFIC is a strategic alliance of WWF and IUCN. All material appearing in this publication is copyrighted and may be reproduced with permission. Any reproduction in full or in part of this publication must credit TRAFFIC International as the copyright owner. Financial support for TRAFFIC’s research and the publication of this report was provided by the WWF Conservation and Adaptation in Asia’s High Mountain Landscapes and Communities Project, funded by the United States Agency for International Development (USAID). The views of the authors expressed in this publication do not necessarily reflect those of the TRAFFIC network, WWF, IUCN or the United States Agency for International Development. The designations of geographical entities in this publication, and the presentation of the material, do not imply the expression of any opinion whatsoever on the part of TRAFFIC or its supporting organizations concerning the legal status of any country, territory, or area, or of its authorities, or concerning the delimitation of its frontiers or boundaries. The TRAFFIC symbol copyright and Registered Trademark ownership is held by WWF. TRAFFIC is a strategic alliance of WWF and IUCN. Suggested citation: Nowell, K., Li, J., Paltsyn, M. and Sharma, R.K. (2016). An Ounce of Prevention: Snow Leopard Crime Revisited. -

Madarao Kogen Snow Report
Madarao Kogen Snow Report National Loren ploat that glossator sain perpendicularly and neaten undyingly. Didymous Coleman sometimes disaffirm his fumigatingsecretness herimpotently captive andunsteadily interfuse and so backcomb patrilineally! tiptop. Replaceable and snuff-brown Andros sheathed while elmiest Mikel Fresh snowfall and freezing levels for snowfall on the yuzawa region, weather updates in both zones: a guide to find, please enable javascript. Plenty of pow in the trees. Tangram to chase the powder; mostly with tour groups from Myoko Kogen and Yudanaka. Awesome day on a ski resorts in luxurious comfort in collides with another option and found on! Awesome day or snow report on the madarao kogen snow reports you looking for steep and even the beaten track, study the following charts. Madarao Snow Report Weather Forecast within the Madarao Kogen Ski ensemble and surrounding areas of Mt Madarao and Madarao Mountain Resort. The picture day of Madarao Ski Resort 2020 on 31st of March Still not. The announcement that they would not be allowed to do so after everything had appeared to be finally approved, Sun Alpina Kashimayari, our bad. We use this site for this using the crowds had to your snow report, but madarao kogen weather, when researching and they would move in summer holidays and powder. Hakuba Snow Report Hakuba Ski Weather & Conditions. Local weather information for the nearby Madarao Kogen ski resorts can be band at. Snow reports you might want to madarao kogen sympathique snow schools. Niseko holds the snow report from the snow pictures and offpiste conditions, or two days at hakuba. Access or receive exclusive benefits. -

Bhadra Voluntary Relocation India
BHADRA VOLUNTARY RELOCATION INDIA INDIA FOREWORD During my tenure as Director Project Tiger in the Ministry of Environment and Forests, Govt. of India, I had the privilege of participating in voluntary relocation of villages from Bhadra Tiger Reserve. As nearly two decades have passed, whatever is written below is from my memory only. Mr Yatish Kumar was the Field Director of Bhadra Tiger Reserve and Mr Gopalakrishne Gowda was the Collector of Chikmagalur District of Karnataka during voluntary relocation in Bhadra Tiger Reserve. This Sanctuary was notified as a Tiger Reserve in the year 1998. After the notification as tiger reserve, it was necessary to relocate the existing villages as the entire population with their cattle were dependent on the Tiger Reserve. The area which I saw in the year 1998 was very rich in flora and fauna. Excellent bamboo forests were available but it had fire hazard too because of the presence of villagers and their cattle. Tiger population was estimated by Dr. Ullas Karanth and his love for this area was due to highly rich biodiversity. Ultimately, resulted in relocation of all the villages from within the reserve. Dr Karanth, a devoted biologist was a close friend of mine and during his visit to Delhi he proposed relocation of villages. As the Director of Project Tiger, I was looking at voluntary relocation of villages for tribals only from inside Tiger Reserve by de-notifying suitable areas of forests for relocation, but in this case the villagers were to be relocated by purchasing a revenue land which was very expensive. -

Wild Life Sanctuaries in INDIA
A M K RESOURCE WORLD GENERAL KNOWLEDGE www.amkresourceinfo.com Wild Life Sanctuaries in INDIA Wildlife Sanctuaries in India are 441 in number. They are a home to hundreds and thousands of various flora and fauna. A wide variety of species thrive in such Wildlife Sanctuaries. With the ever growing cement – jungle, it is of utmost importance to protect and conserve wildlife and give them their own, natural space to survive Wildlife Sanctuaries are established by IUCN category II protected areas. A wildlife sanctuary is a place of refuge where abused, injured, endangered animals live in peace and dignity. Senchal Game Sanctuary. Established in 1915 is the oldest of such sanctuaries in India. Chal Batohi, in Gujarat is the largest Wildlife Sanctuary in India. The conservative measures taken by the Indian Government for the conservation of Tigers was awarded by a 30% rise in the number of tigers in 2015. According to the Red Data Book of International Union for Conservation of Nature (IUCN), there are 47 critically endangered species in India. DO YOU KNOW? Wildlife sanctuaries in India are established by IUCN category II protected areas. India has 537 wildlife sanctuaries referred to as wildlife sanctuaries category IV protected areas. Among these, the 50 tiger reserves are governed by Project Tiger, and are of special significance in the conservation of the tiger. Some wildlife sanctuaries in India are specifically named bird sanctuary, e.g., Keoladeo National Park before attaining National Park status. Many of them being referred as as a particular animal such as Jawai leopard sanctuary in Rajasthan. -

3.4 ORDER CARNIVORA Bowdich, 1821
3.4 ORDER CARNIVORA Bowdich, 1821 3.4.1 Family Ursidae Fischer, 1817 There are eight species of bears in the world: - American Black Bear Ursus americanus - Brown Bear Ursus arctos - Polar Bear Ursus maritimus - Sloth Bear Melursus ursinus - Spectacled Bear Tremarctos ornatus - Giant Panda Ailuropoda melanoleuca - Asiatic Black Bear Ursus thibetanus - Malayan Sun Bear Helarctos malayanus The last two species are the only members of the family Ursidae known in Southeast Asia. They differ from each other by their furs and body sizes and both are threatened with extinction (Nowak, 1991; Corbet & Hill 1992). Bears have relatively undeveloped carnassial teeth; narrow premolars, crushing molars with flat crowns and large robust canines. 127 3.4.1.1 Subfamily Ursinae Fischer, 1817, Plate 3(A1 to B3) As mentioned above, two genera and two species represent the subfamily Ursinae in Southeast Asia, namely: - Malayan Sun Bear (Figure 3.8, A), Ursus/Helarctos malayanus (Raffles, 1821) with the scientific name Ursu and synonym Helarctos is distributed in the south west of China, Assam, Myanmar, Vietnam, Peninsular Malaysia, to the islands of Sumatra and Borneo. It is the smallest of all bears found in the tropical rainforests of Southeast Asia. - Asiatic Black Bear (Figure 3.8, B), Ursus thibetanus Cuvier, 1823 is mainly localized in the Himalayas, Afghanistan to southern China, Myanmar, northern Thailand and Indochina. It has several alternative names including Asiatic Black Bear, Himalayan Black Bear, Moon Bear and inhabits mountain forests. Figure 3.8 Malayan Sun Bear (A) and Asiatic Black Bear (B) in Zoo Negara, Malaysia National Zoological Park. -

2020 FUNDRAISING TOOL KIT 2 • Njsnowbowl.Org ABOUT
1 • njsnowbowl.org 2020 FUNDRAISING TOOL KIT 2 • njsnowbowl.org ABOUT The 2020 Snow Bowl is an official 6-on-6 flag football tournament with light blocking and is played over three days. Teams are allowed up to 15 players; all must be 18 years of age or older to participate. Field size is 20 yards x 40 yards with each team guaranteed three games on the field at MetLife Stadium. Special Olympics New Jersey, a not-for-profit organization, provides year-round sports training, competition, leadership opportunities and health screenings to more than 26,000 athletes. All of these programs and services are always completely FREE thanks to fundraising events like the 2020 Snow Bowl! GETTING STARTED 1. SET UP YOUR PERSONAL FUNDRAISING PAGE. If you didn’t get a chance to set up your personal fundraising page when you registered, be sure to do so as soon as possible! Classy.org should have sent you a confirmation email to claim your unique fundraising page. You can then log into your fundraising page and customize your site. 2. RAISE FUNDS. Now that you have a personalized fundraising site and link, be sure to share it with your friends and family. Start by posting your custom link on Facebook, sending it via email to your contacts or tweet it out to your followers. You can do this directly from your Classy fundraising page. Just click on one of the icons on the top right in the “Share” section. 3. AIM HIGH, WIN COOL STUFF! We encourage you to aim HIGH with your fundraising goals! As you hit your fundraising milestones, you’ll earn cumulative (and AWESOME) incentives. -

Observations of Small Carnivores in Son Tra Nature Reserve, a Small and Isolated Protected Area in Central Vietnam
Observations of small carnivores in Son Tra Nature Reserve, a small and isolated protected area in central Vietnam Ulrike STREICHER1 and Larry ULIBARRI2 Abstract Over half the 45.5 km² Son Tra peninsula in central Vietnam is a nature reserve. The peninsula has been isolated from other natural habitat by sea and urbanisation for decades. Various surveys since the 1960s have recorded Large-toothed Ferret Badger Melogale personata, Small Indian Civet Viverricula indica, Common Palm Civet Paradoxurus hermaphroditus, Small Asian Mongoose Herpestes javanicus and Leopard Cat Prionailurus bengalensis; and probably otter (Lutrinae) and Large Indian Civet Viverra zibetha, although the original basis for these two is not available. Several species typical of forest in this region and active at least in large part by day were not found, suggesting that they are possibly susceptible to hunting or need larger landscapes (or both). None of the surveys targeted small carnivores, so some species, particularly nocturnal ones, might have been overlooked. The easily accessible Son Tra KeywordsNature Reserve: breeding with seasonality,its unusually community, confiding wildlife fragmentation, is ideal for habitat wildlife change, and conservation Herpestes javanicus studies and, locality education. records, Melogale per- sonata, persistence Ghi nhận thú ăn thịt nhỏ ở Khu Bảo tồn Thiên nhiên Sơn Trà, một khu bảo vệ nhỏ và cô lập ở miền trung Việt Nam Tóm tắt Khoảng một nửa diện tích 45,5 km² của bán đảo Sơn Trà ở miền trung Việt Nam là môt khu bảo tồn. Bán đảo đã bị cô lập với các sinh cảnh tự nhiên khác bởi biển và các khu đô thị từ vài thập kỷ nay. -

Mammalian Predators Appropriating the Refugia of Their Prey
Mamm Res (2015) 60:285–292 DOI 10.1007/s13364-015-0236-y ORIGINAL PAPER When prey provide more than food: mammalian predators appropriating the refugia of their prey William J. Zielinski 1 Received: 30 September 2014 /Accepted: 20 July 2015 /Published online: 31 July 2015 # Mammal Research Institute, Polish Academy of Sciences, Białowieża, Poland (outside the USA) 2015 Abstract Some mammalian predators acquire both food and predators) may play disproportionately important roles in their shelter from their prey, by eating them and using the refugia communities. the prey construct. I searched the literature for examples of predators that exhibit this behavior and summarize their taxo- Keywords Predator–prey . Dens . Herbivore . Behavior . nomic affiliations, relative sizes, and distributions. I hypothe- Habitat . Resting . Foraging sized that size ratios of species involved in this dynamic would be near 1.0, and that most of these interactions would occur at intermediate and high latitudes. Seventeen species of Introduction Carnivorans exploited at least 23 species of herbivores as food and for their refugia. Most of them (76.4 %) were in the Mammals require food and most require shelter, either to pro- Mustelidae; several small species of canids and a few tect them from predators or from thermal stress. Carnivorous herpestids were exceptions. Surprisingly, the average mammals are unique in that they subsist on mobile food predator/prey weight ratio was 10.51, but few species of pred- sources which, particularly if these sources are vertebrates, ators were more than ten times the weight of the prey whose may build their own refuges to help regulate their body tem- refugia they exploit.