The Keap1–Nrf2 Pathway in Toxicology—Curtis Klaassen
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Thioredoxin Trx-1 Regulates the Major Oxidative Stress Response Transcription Factor, Skn-1, in Caenorhabditis Elegans
The Texas Medical Center Library DigitalCommons@TMC The University of Texas MD Anderson Cancer Center UTHealth Graduate School of The University of Texas MD Anderson Cancer Biomedical Sciences Dissertations and Theses Center UTHealth Graduate School of (Open Access) Biomedical Sciences 5-2016 THE THIOREDOXIN TRX-1 REGULATES THE MAJOR OXIDATIVE STRESS RESPONSE TRANSCRIPTION FACTOR, SKN-1, IN CAENORHABDITIS ELEGANS Katie C. McCallum Follow this and additional works at: https://digitalcommons.library.tmc.edu/utgsbs_dissertations Part of the Cellular and Molecular Physiology Commons, Medicine and Health Sciences Commons, Molecular Genetics Commons, and the Organismal Biological Physiology Commons Recommended Citation McCallum, Katie C., "THE THIOREDOXIN TRX-1 REGULATES THE MAJOR OXIDATIVE STRESS RESPONSE TRANSCRIPTION FACTOR, SKN-1, IN CAENORHABDITIS ELEGANS" (2016). The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences Dissertations and Theses (Open Access). 655. https://digitalcommons.library.tmc.edu/utgsbs_dissertations/655 This Dissertation (PhD) is brought to you for free and open access by the The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences at DigitalCommons@TMC. It has been accepted for inclusion in The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences Dissertations and Theses (Open Access) by an authorized administrator of DigitalCommons@TMC. For more information, please contact [email protected]. THE THIOREDOXIN TRX-1 REGULATES THE MAJOR OXIDATIVE STRESS RESPONSE TRANSCRIPTION FACTOR, SKN-1, IN CAENORHABDITIS ELEGANS A DISSERTATION Presented to the Faculty of The University of Texas Health Science Center at Houston and The University of Texas MD Anderson Cancer Center Graduate School of Biomedical Sciences in Partial Fulfillment of the Requirements for the Degree of DOCTOR OF PHILOSOPHY by Katie Carol McCallum, B.S. -

The Role of Glutathionperoxidase 4 (GPX4) in Hematopoiesis and Leukemia
The role of glutathionperoxidase 4 (GPX4) in hematopoiesis and leukemia von Kira Célénie Stahnke Inaugral- Dissertation zur Erlangung der Doktorwürde der TierärztliChen Fakultät der Ludwig-Maximilians-Universität MünChen The role of glutathionperoxidase 4 (GPX4) in hematopoiesis and leukemia von Kira Célénie Stahnke aus Dortmund München 2016 Aus dem VeterinärwissensChaftliChen Department der TierärztliChen Fakultät der Ludwig-Maximilians-Universität München Lehrstuhl für Molekulare Tierzucht und Biotechnologie Arbeit angefertigt unter der Leitung von Univ.-Prof. Dr. Eckhard Wolf Angefertigt am Institut für Experimentelle TumorforsChung Universitätsklinikum Ulm Mentor: Prof. Dr. Christian Buske GedruCkt mit Genehmigung der TierärztliChen Fakultät der Ludwig-Maximilians-Universität München Dekan: Univ.-Prof. Dr. JoaChim Braun Berichterstatter: Univ.-Prof. Dr. Eckard Wolf Korreferent/en: Priv.-Doz. Dr. Bianka Schulz Tag der Promotion: 06.02.2016 Table of contents List of Abbreviations..............................................................................2 1 Introduction ........................................................................................6 1.1 ROS and oxidative stress in physiology and pathology..................................................... 6 1.1.1 SourCes of Reactive Oxygen SpeCies..................................................................................................6 1.1.2 PhysiologiCal role of ROS........................................................................................................................8 -

In Thyroid Cancer
Metere et al. Cancer Cell Int (2018) 18:7 https://doi.org/10.1186/s12935-018-0504-4 Cancer Cell International PRIMARY RESEARCH Open Access A possible role for selenoprotein glutathione peroxidase (GPx1) and thioredoxin reductases (TrxR1) in thyroid cancer: our experience in thyroid surgery Alessio Metere1* , Francesca Frezzotti1, Claire Elizabeth Graves2, Massimo Vergine1, Alessandro De Luca1, Donatella Pietraforte3 and Laura Giacomelli1 Abstract Background: Oxidative stress is responsible for some alterations in the chemical structure and, consequently, in the function of proteins, lipids, and DNA. Recent studies have linked oxidative stress to cancers, particularly thyroid cancer, but the mechanisms remain unclear. Here, we further characterize the role of oxidative stress in thyroid cancer by analyzing the expression of two selenium antioxidant molecules, glutathione peroxidase (GPx1) and thioredoxin reductase (TrxR1) in thyroid cancer cells. Methods: Samples of both healthy thyroid tissue and thyroid tumor were taken for analysis after total thyroidectomy. The expression of GPx1 and TrxR1 was revealed by Western blot analysis and quantifed by densitometric analy- ses, while the evaluation of free radicals was performed by Electron Paramagnetic Resonance (EPR)-spin trapping technique. Results: Our results show a decrease in the expression of GPx1 and TrxR1 ( 45.7 and 43.2% respectively, p < 0.01) in the thyroid cancer cells compared to the healthy cells. In addition, the EPR− technique− shows an increase of free radicals in tumor tissue, signifcantly higher than that found in healthy thyroid tissue ( 116.3%, p < 0.01). + Conclusions: Our fndings underscore the relationship between thyroid cancer and oxidative stress, showing the imbalance of the oxidant/antioxidant system in thyroid cancer tissue. -

Domain Structure of the Glucocorticoid Receptor Protein
Proc. Nati. Acad. Sci. USA Vol. 84, pp. 4437-4440, July 1987 Biochemistry Domain structure of the glucocorticoid receptor protein (proteolysis/steroid binding/DNA binding/protein sequence) JAN CARLSTEDT-DUKE*, PER-ERIK STROMSTEDT*, ORJAN WRANGEt, TOMAS BERGMANt, JAN-AKE GuSTAFSSON*, AND HANS JORNVALLf *Department of Medical Nutrition, Karolinska Institute, Huddinge University Hospital, F69, S-141 86 Huddinge, Sweden; and Departments of tMedical Cell Genetics and tChemistry, Karolinska Institute, Box 60400, S-104 01 Stockholm, Sweden Communicated by Viktor Mutt, March 26, 1987 (receivedfor review December 1, 1986) ABSTRACT The purified rat liver glucocorticoid receptor GR was eluted with 27.5 mM MgCl2 and further purified by protein was analyzed by limited proteolysis and amino acid chromatography on DEAE-Sepharose, eluted with a linear sequence determination. The NH2 terminus appears to be NaCl gradient. The receptor was detected by analysis for 3H blocked. The steroid-binding domain, defined by a unique radioactivity. Each purification resulted in a yield of '50 ,g tryptic cleavage site, corresponds to the COOH-terminal part of GR, starting from eight rat livers. The purified prepara- of the protein with the domain border in the region of residue tions of GR contained the 94-kDa GR, the 72-kDa GR- 518. The DNA-binding domain, defined by a region with associated protein that is unrelated to GR (12, 13), as well as chymotryptic cleavage sites, is immediately adjacent to the very small amounts of proteolytic GR fragments (usually steroid-binding domain and reflects another domain border in <5% of total protein according to densitometric analysis of the region of residues 410-414. -

Oxidative Stress Modulates the Expression Pattern of Peroxiredoxin-6 in Peripheral Blood Mononuclear Cells of Asthmatic Patients and Bronchial Epithelial Cells
Allergy Asthma Immunol Res. 2020 May;12(3):523-536 https://doi.org/10.4168/aair.2020.12.3.523 pISSN 2092-7355·eISSN 2092-7363 Original Article Oxidative Stress Modulates the Expression Pattern of Peroxiredoxin-6 in Peripheral Blood Mononuclear Cells of Asthmatic Patients and Bronchial Epithelial Cells Hyun Jae Shim ,1 So-Young Park ,2 Hyouk-Soo Kwon ,1 Woo-Jung Song ,1 Tae-Bum Kim ,1 Keun-Ai Moon ,1 Jun-Pyo Choi ,1 Sin-Jeong Kim ,1 You Sook Cho 1* 1Division of Allergy and Clinical Immunology, Department of Internal Medicine, Asan Medical Center, University of Ulsan College of Medicine, Seoul, Korea 2Division of Pulmonary, Allergy and Critical Care Medicine, Department of Internal Medicine, Konkuk University Medical Center, Seoul, Korea Received: Jul 8, 2019 ABSTRACT Revised: Nov 29, 2019 Accepted: Dec 23, 2019 Purpose: Reduction-oxidation reaction homeostasis is vital for regulating inflammatory Correspondence to conditions and its dysregulation may affect the pathogenesis of chronic airway inflammatory You Sook Cho, MD, PhD diseases such as asthma. Peroxiredoxin-6, an important intracellular anti-oxidant molecule, Division of Allergy and Clinical Immunology, Department of Internal Medicine, Asan is reported to be highly expressed in the airways and lungs. The aim of this study was to Medical Center, University of Ulsan College of analyze the expression pattern of peroxiredoxin-6 in the peripheral blood mononuclear cells Medicine, 88 Olympic-ro 43-gil, Songpa-gu, (PBMCs) of asthmatic patients and in bronchial epithelial cells (BECs). Seoul 05505, Korea. Methods: The expression levels and modifications of peroxiredoxin-6 were evaluated in Tel: +82-2-3010-3285 PBMCs from 22 asthmatic patients. -

Role of Thioredoxin-Interacting Protein in Diseases and Its Therapeutic Outlook
International Journal of Molecular Sciences Review Role of Thioredoxin-Interacting Protein in Diseases and Its Therapeutic Outlook Naila Qayyum 1,†, Muhammad Haseeb 1,† , Moon Suk Kim 1 and Sangdun Choi 1,2,* 1 Department of Molecular Science and Technology, Ajou University, Suwon 16499, Korea; [email protected] (N.Q.); [email protected] (M.H.); [email protected] (M.S.K.) 2 S&K Therapeutics, Woncheon Hall 135, Ajou University, Suwon 16499, Korea * Correspondence: [email protected] † These authors contributed equally to this work. Abstract: Thioredoxin-interacting protein (TXNIP), widely known as thioredoxin-binding protein 2 (TBP2), is a major binding mediator in the thioredoxin (TXN) antioxidant system, which involves a reduction-oxidation (redox) signaling complex and is pivotal for the pathophysiology of some diseases. TXNIP increases reactive oxygen species production and oxidative stress and thereby contributes to apoptosis. Recent studies indicate an evolving role of TXNIP in the pathogenesis of complex diseases such as metabolic disorders, neurological disorders, and inflammatory illnesses. In addition, TXNIP has gained significant attention due to its wide range of functions in energy metabolism, insulin sensitivity, improved insulin secretion, and also in the regulation of glucose and tumor suppressor activities in various cancers. This review aims to highlight the roles of TXNIP in the field of diabetology, neurodegenerative diseases, and inflammation. TXNIP is found to be a promising novel therapeutic target in the current review, not only in the aforementioned diseases but also in prolonged microvascular and macrovascular diseases. Therefore, TXNIP inhibitors hold promise for preventing the growing incidence of complications in relevant diseases. -
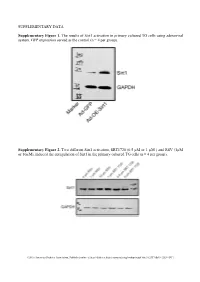
SUPPLEMENTARY DATA Supplementary Figure 1. The
SUPPLEMENTARY DATA Supplementary Figure 1. The results of Sirt1 activation in primary cultured TG cells using adenoviral system. GFP expression served as the control (n = 4 per group). Supplementary Figure 2. Two different Sirt1 activators, SRT1720 (0.5 µM or 1 µM ) and RSV (1µM or 10µM), induced the upregulation of Sirt1 in the primary cultured TG cells (n = 4 per group). ©2016 American Diabetes Association. Published online at http://diabetes.diabetesjournals.org/lookup/suppl/doi:10.2337/db15-1283/-/DC1 SUPPLEMENTARY DATA Supplementary Table 1. Primers used in qPCR Gene Name Primer Sequences Product Size (bp) Sirt1 F: tgccatcatgaagccagaga 241 (NM_001159589) R: aacatcgcagtctccaagga NOX4 F: tgtgcctttattgtgcggag 172 (NM_001285833.1) R: gctgatacactggggcaatg Supplementary Table 2. Antibodies used in Western blot or Immunofluorescence Antibody Company Cat. No Isotype Dilution Sirt1 Santa Cruz * sc-15404 Rabbit IgG 1/200 NF200 Sigma** N5389 Mouse IgG 1/500 Tubulin R&D# MAB1195 Mouse IgG 1/500 NOX4 Abcam† Ab133303 Rabbit IgG 1/500 NOX2 Abcam Ab129068 Rabbit IgG 1/500 phospho-AKT CST‡ #4060 Rabbit IgG 1/500 EGFR CST #4267 Rabbit IgG 1/500 Ki67 Santa Cruz sc-7846 Goat IgG 1/500 * Santa Cruz Biotechnology, Santa Cruz, CA, USA ** Sigma aldrich, Shanghai, China # R&D Systems Inc, Minneapolis, MN, USA † Abcam, Inc., Cambridge, MA, USA ‡ Cell Signaling Technology, Inc., Danvers, MA, USA ©2016 American Diabetes Association. Published online at http://diabetes.diabetesjournals.org/lookup/suppl/doi:10.2337/db15-1283/-/DC1 SUPPLEMENTARY DATA Supplementary -

Role of Peroxiredoxin 6 in Human Melanoma
Role of Peroxiredoxin 6 in human melanoma Dissertation zur Erlangung des naturwissenschaftlichen Doktorgrades der Bayerischen Julius-Maximilians-Universität Würzburg vorgelegt von Alexandra Schmitt aus Würzburg Würzburg 2015 Eingereicht am:__________________________ Mitglieder der Promotionskommission: Vorsitzender:____________________________ Gutachter:______________________________ Gutachter:______________________________ Tag des Promotionskolloquiums:___________________ Doktorurkunde ausgehändigt am:___________________ Eidesstattliche Erklärung Gemäß §4, Abs. 3, Ziff. 3, 5 und 8 der Promotionsordnung der Fakultät für Biologie der Bayerischen Julius-Maximilians-Universität Würzburg Hiermit erkläre ich ehrenwörtlich, dass ich die vorliegende Dissertation selbständig angefertigt und keine anderen als die angegebenen Quellen und Hilfsmittel verwendet habe. Ich erkläre weiterhin, dass die vorliegende Dissertation weder in gleicher, noch in ähnlicher Form bereits in einem anderen Prüfungsverfahren vorgelegen hat. Weiterhin erkläre ich, dass ich außer den mit dem Zulassungsantrag urkundlich vorgelegten Graden keine weiteren akademischen Grade erworben oder zu erwerben versucht habe. Würzburg, Januar 2015 ______________________________________ Alexandra Schmitt Table of contents 1. Abstract ................................................................................................................. 1 2. Zusammenfassung ............................................................................................... 3 3. Introduction .......................................................................................................... -

GPX4 at the Crossroads of Lipid Homeostasis and Ferroptosis Giovanni C
REVIEW GPX4 www.proteomics-journal.com GPX4 at the Crossroads of Lipid Homeostasis and Ferroptosis Giovanni C. Forcina and Scott J. Dixon* formation of toxic radicals (e.g., R-O•).[5] Oxygen is necessary for aerobic metabolism but can cause the harmful The eight mammalian GPX proteins fall oxidation of lipids and other macromolecules. Oxidation of cholesterol and into three clades based on amino acid phospholipids containing polyunsaturated fatty acyl chains can lead to lipid sequence similarity: GPX1 and GPX2; peroxidation, membrane damage, and cell death. Lipid hydroperoxides are key GPX3, GPX5, and GPX6; and GPX4, GPX7, and GPX8.[6] GPX1–4 and 6 (in intermediates in the process of lipid peroxidation. The lipid hydroperoxidase humans) are selenoproteins that contain glutathione peroxidase 4 (GPX4) converts lipid hydroperoxides to lipid an essential selenocysteine in the enzyme + alcohols, and this process prevents the iron (Fe2 )-dependent formation of active site, while GPX5, 6 (in mouse and toxic lipid reactive oxygen species (ROS). Inhibition of GPX4 function leads to rats), 7, and 8 use an active site cysteine lipid peroxidation and can result in the induction of ferroptosis, an instead. Unlike other family members, GPX4 (PHGPx) can act as a phospholipid iron-dependent, non-apoptotic form of cell death. This review describes the hydroperoxidase to reduce lipid perox- formation of reactive lipid species, the function of GPX4 in preventing ides to lipid alcohols.[7,8] Thus,GPX4ac- oxidative lipid damage, and the link between GPX4 dysfunction, lipid tivity is essential to maintain lipid home- oxidation, and the induction of ferroptosis. ostasis in the cell, prevent the accumula- tion of toxic lipid ROS and thereby block the onset of an oxidative, iron-dependent, non-apoptotic mode of cell death termed 1. -

Is Glyceraldehyde-3-Phosphate Dehydrogenase a Central Redox Mediator?
1 Is glyceraldehyde-3-phosphate dehydrogenase a central redox mediator? 2 Grace Russell, David Veal, John T. Hancock* 3 Department of Applied Sciences, University of the West of England, Bristol, 4 UK. 5 *Correspondence: 6 Prof. John T. Hancock 7 Faculty of Health and Applied Sciences, 8 University of the West of England, Bristol, BS16 1QY, UK. 9 [email protected] 10 11 SHORT TITLE | Redox and GAPDH 12 13 ABSTRACT 14 D-Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) is an immensely important 15 enzyme carrying out a vital step in glycolysis and is found in all living organisms. 16 Although there are several isoforms identified in many species, it is now recognized 17 that cytosolic GAPDH has numerous moonlighting roles and is found in a variety of 18 intracellular locations, but also is associated with external membranes and the 19 extracellular environment. The switch of GAPDH function, from what would be 20 considered as its main metabolic role, to its alternate activities, is often under the 21 influence of redox active compounds. Reactive oxygen species (ROS), such as 22 hydrogen peroxide, along with reactive nitrogen species (RNS), such as nitric oxide, 23 are produced by a variety of mechanisms in cells, including from metabolic 24 processes, with their accumulation in cells being dramatically increased under stress 25 conditions. Overall, such reactive compounds contribute to the redox signaling of the 26 cell. Commonly redox signaling leads to post-translational modification of proteins, 27 often on the thiol groups of cysteine residues. In GAPDH the active site cysteine can 28 be modified in a variety of ways, but of pertinence, can be altered by both ROS and 29 RNS, as well as hydrogen sulfide and glutathione. -

Thiol Peroxidases Mediate Specific Genome-Wide Regulation of Gene Expression in Response to Hydrogen Peroxide
Thiol peroxidases mediate specific genome-wide regulation of gene expression in response to hydrogen peroxide Dmitri E. Fomenkoa,1,2, Ahmet Koca,1, Natalia Agishevaa, Michael Jacobsena,b, Alaattin Kayaa,c, Mikalai Malinouskia,c, Julian C. Rutherfordd, Kam-Leung Siue, Dong-Yan Jine, Dennis R. Winged, and Vadim N. Gladysheva,c,2 aDepartment of Biochemistry, University of Nebraska, Lincoln, NE 68588-0664; bDepartment of Life Sciences, Wayne State College, Wayne, NE 68787; dDepartment of Medicine, University of Utah Health Sciences Center, Salt Lake City, UT 84132; eDepartment of Biochemistry, University of Hong Kong, Hong Kong, China; and cDivision of Genetics, Department of Medicine, Brigham and Women’s Hospital and Harvard Medical School, Boston, MA 02115 Edited by Joan Selverstone Valentine, University of California, Los Angeles, CA, and approved December 22, 2010 (received for review July 21, 2010) Hydrogen peroxide is thought to regulate cellular processes by and could withstand significant oxidative stress. It responded to direct oxidation of numerous cellular proteins, whereas antioxi- several redox stimuli by robust transcriptional reprogramming. dants, most notably thiol peroxidases, are thought to reduce However, it was unable to transcriptionally respond to hydrogen peroxides and inhibit H2O2 response. However, thiol peroxidases peroxide. The data suggested that thiol peroxidases transfer have also been implicated in activation of transcription factors oxidative signals from peroxides to target proteins, thus activating and signaling. It remains unclear if these enzymes stimulate or various transcriptional programs. This study revealed a previously inhibit redox regulation and whether this regulation is widespread undescribed function of these proteins, in addition to their roles or limited to a few cellular components. -

The C718T Polymorphism in the 3″-Untranslated
Hypertension Research (2012) 35, 507–512 & 2012 The Japanese Society of Hypertension All rights reserved 0916-9636/12 www.nature.com/hr ORIGINAL ARTICLE The C718T polymorphism in the 3¢-untranslated region of glutathione peroxidase-4 gene is a predictor of cerebral stroke in patients with essential hypertension Alexey V Polonikov1, Ekaterina K Vialykh2, Mikhail I Churnosov3, Thomas Illig4, Maxim B Freidin5, Oksana V Vasil¢eva1, Olga Yu Bushueva1, Valentina N Ryzhaeva1, Irina V Bulgakova1 and Maria A Solodilova1 In the present study we have investigated the association of three single nucleotide polymorphisms in glutathione peroxidase (GPx) genes GPX1 rs1050450 (P198L), GPX3 rs2070593 (G930A) and GPX4 rs713041 (T718C) with the risk of cerebral stroke (CS) in patients with essential hypertension (EH). A total of 667 unrelated EH patients of Russian origin, including 306 hypertensives (the EH–CS group) who suffered from CS and 361 people (the EH–CS group) who did not have cerebrovascular accidents, were enrolled in the study. The variant allele 718C of the GPX4 gene was found to be significantly associated with an increased risk of CS in hypertensive patients (odds ratio (OR) 1.53, 95% confidence interval (CI) 1.23–1.90, Padj¼0.0003). The prevalence of the 718TC and 718CC genotypes of the GPX4 gene was higher in the EH–CS group than the EH-alone group (OR¼2.12, 95%CI 1.42–3.16, Padj¼0.0018). The association of the variant GPX4 genotypes with the increased risk of CS in hypertensives remained statistically significant after adjusting for confounding variables such as sex, body mass index (BMI), blood pressure and antihypertensive medication use (OR¼2.18, 95%CI 1.46–3.27, P¼0.0015).