RNA Pol II Length and Disorder Enable Cooperative Scaling of Transcriptional Bursting
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Transcriptional Burst Frequency and Burst Size Are Equally Modulated Across the Human Genome
Transcriptional burst frequency and burst size are equally modulated across the human genome Roy D. Dara,b,c,1, Brandon S. Razookya,d,e,1, Abhyudai Singhd,2, Thomas V. Trimelonif, James M. McCollumf, Chris D. Coxg,h, Michael L. Simpsonb,I,3, and Leor S. Weinbergera,d,j,3 aGladstone Institutes, San Francisco, CA 94158; bCenter for Nanophase Materials Sciences, Oak Ridge National Laboratory, Oak Ridge, TN 37831; cDepartment of Physics and Astronomy, University of Tennessee, Knoxville, TN 37996; dDepartment of Chemistry and Biochemistry, University of California at San Diego, La Jolla, CA 92093; eBiophysics Graduate Group, University of California, San Francisco, CA 94158; gCenter for Environmental Biotechnology, University of Tennessee, Knoxville, TN 37996;hDepartment of Civil and Environmental Engineering, University of Tennessee, Knoxville, TN 37996; IDepartment of Materials Science and Engineering, University of Tennessee, Knoxville, TN 37996; jDepartment of Biochemistry and Biophysics, University of California, San Francisco, CA 94158; and fDepartment of Electrical and Computer Engineering, Virginia Commonwealth University, Richmond, VA 23284–3072 Edited by Jonathan S. Weissman, University of California, San Francisco, CA, and approved September 13, 2012 (received for review August 8, 2012) Gene expression occurs either as an episodic process, characterized autocorrelation time of expression fluctuations (as measured by pulsatile bursts, or as a constitutive process, characterized by by the noise autocorrelation time at half of its initial value, τ1∕2) a Poisson-like accumulation of gene products. It is not clear which (11, 12) (Fig. 1C). Although this three-dimensional noise space is mode of gene expression (constitutive versus bursty) predomi- impractical to analyze directly, different two-dimensional projec- nates across a genome or how transcriptional dynamics are influ- tions of noise space allow the quantification of rate parameters enced by genomic position and promoter sequence. -

Genome-Wide Analysis of Transcriptional Bursting-Induced Noise in Mammalian Cells
bioRxiv preprint doi: https://doi.org/10.1101/736207; this version posted August 15, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Title: Genome-wide analysis of transcriptional bursting-induced noise in mammalian cells Authors: Hiroshi Ochiai1*, Tetsutaro Hayashi2, Mana Umeda2, Mika Yoshimura2, Akihito Harada3, Yukiko Shimizu4, Kenta Nakano4, Noriko Saitoh5, Hiroshi Kimura6, Zhe Liu7, Takashi Yamamoto1, Tadashi Okamura4,8, Yasuyuki Ohkawa3, Itoshi Nikaido2,9* Affiliations: 1Graduate School of Integrated Sciences for Life, Hiroshima University, Higashi-Hiroshima, Hiroshima, 739-0046, Japan 2Laboratory for Bioinformatics Research, RIKEN BDR, Wako, Saitama, 351-0198, Japan 3Division of Transcriptomics, Medical Institute of Bioregulation, Kyushu University, Fukuoka, Fukuoka, 812-0054, Japan 4Department of Animal Medicine, National Center for Global Health and Medicine (NCGM), Tokyo, 812-0054, Japan 5Division of Cancer Biology, The Cancer Institute of JFCR, Tokyo, 135-8550, Japan 6Graduate School of Bioscience and Biotechnology, Tokyo Institute of Technology, Yokohama, Kanagawa, 226-8503, Japan 7Janelia Research Campus, Howard Hughes Medical Institute, Ashburn, VA, 20147, USA 8Section of Animal Models, Department of Infectious Diseases, National Center for Global Health and Medicine (NCGM), Tokyo, 812-0054, Japan 9Bioinformatics Course, Master’s/Doctoral Program in Life Science Innovation (T-LSI), School of Integrative and Global Majors (SIGMA), University of Tsukuba, Wako, 351-0198, Japan *Corresponding authors Corresponding authors e-mail addresses Hiroshi Ochiai, [email protected] Itoshi Nikaido, [email protected] bioRxiv preprint doi: https://doi.org/10.1101/736207; this version posted August 15, 2019. -

The Cyclin-Dependent Kinase 8 Module Sterically Blocks Mediator Interactions with RNA Polymerase II
The cyclin-dependent kinase 8 module sterically blocks Mediator interactions with RNA polymerase II Hans Elmlund*†, Vera Baraznenok‡, Martin Lindahl†, Camilla O. Samuelsen§, Philip J. B. Koeck*¶, Steen Holmberg§, Hans Hebert*ʈ, and Claes M. Gustafsson‡ʈ *Department of Biosciences and Nutrition, Karolinska Institutet and School of Technology and Health, Royal Institute of Technology, Novum, SE-141 87 Huddinge, Sweden; †Department of Molecular Biophysics, Lund University, P.O. Box 124, SE-221 00 Lund, Sweden; ‡Division of Metabolic Diseases, Karolinska Institutet, Novum, SE-141 86 Huddinge, Sweden; §Department of Genetics, Institute of Molecular Biology, Oester Farimagsgade 2A, DK-1353 Copenhagen K, Denmark; and ¶University College of Southern Stockholm, SE-141 57 Huddinge, Sweden Communicated by Roger D. Kornberg, Stanford University School of Medicine, Stanford, CA, August 28, 2006 (received for review February 21, 2006) CDK8 (cyclin-dependent kinase 8), along with CycC, Med12, and Here, we use the S. pombe system to investigate the molecular Med13, form a repressive module (the Cdk8 module) that prevents basis for the distinct functional properties of S and L Mediator. RNA polymerase II (pol II) interactions with Mediator. Here, we We find that the Cdk8 module binds to the pol II-binding cleft report that the ability of the Cdk8 module to prevent pol II of Mediator, where it sterically blocks interactions with the interactions is independent of the Cdk8-dependent kinase activity. polymerase. In contrast to earlier assumptions, the Cdk8 kinase We use electron microscopy and single-particle reconstruction to activity is dispensable for negative regulation of pol II interac- demonstrate that the Cdk8 module forms a distinct structural tions with Mediator. -

Functional Diversification of the Nleg Effector Family in Enterohemorrhagic Escherichia Coli
Functional diversification of the NleG effector family in enterohemorrhagic Escherichia coli Dylan Valleaua, Dustin J. Littleb, Dominika Borekc,d, Tatiana Skarinaa, Andrew T. Quailea, Rosa Di Leoa, Scott Houlistone,f, Alexander Lemake,f, Cheryl H. Arrowsmithe,f, Brian K. Coombesb, and Alexei Savchenkoa,g,1 aDepartment of Chemical Engineering and Applied Chemistry, University of Toronto, Toronto, ON M5S 3E5, Canada; bDepartment of Biochemistry and Biomedical Sciences, Michael G. DeGroote Institute for Infectious Disease Research, McMaster University, Hamilton, ON L8S 4K1, Canada; cDepartment of Biophysics, University of Texas Southwestern Medical Center, Dallas, TX 75390; dDepartment of Biochemistry, University of Texas Southwestern Medical Center, Dallas, TX 75390; ePrincess Margaret Cancer Centre, University of Toronto, Toronto, ON M5G 2M9, Canada; fDepartment of Medical Biophysics, University of Toronto, Toronto, ON M5G 1L7, Canada; and gDepartment of Microbiology, Immunology and Infectious Diseases, University of Calgary, Calgary, AB T2N 4N1, Canada Edited by Ralph R. Isberg, Howard Hughes Medical Institute and Tufts University School of Medicine, Boston, MA, and approved August 15, 2018 (receivedfor review November 6, 2017) The pathogenic strategy of Escherichia coli and many other gram- chains. Depending on the length and nature of the polyubiquitin negative pathogens relies on the translocation of a specific set of chain, it posttranslationally regulates the target protein’s locali- proteins, called effectors, into the eukaryotic host cell during in- zation, activation, or degradation. Ubiquitination is a multistep fection. These effectors act in concert to modulate host cell pro- process which begins with the ubiquitin-activating enzyme (E1) cesses in favor of the invading pathogen. Injected by the type III using ATP to “charge” ubiquitin, covalently binding the ubiquitin secretion system (T3SS), the effector arsenal of enterohemorrhagic C terminus by a thioester linkage. -

FBXL19 Recruits CDK-Mediator to Cpg Islands of Developmental Genes Priming Them for Activation During Lineage Commitment
RESEARCH ARTICLE FBXL19 recruits CDK-Mediator to CpG islands of developmental genes priming them for activation during lineage commitment Emilia Dimitrova1, Takashi Kondo2†, Angelika Feldmann1†, Manabu Nakayama3, Yoko Koseki2, Rebecca Konietzny4‡, Benedikt M Kessler4, Haruhiko Koseki2,5, Robert J Klose1* 1Department of Biochemistry, University of Oxford, Oxford, United Kingdom; 2Laboratory for Developmental Genetics, RIKEN Center for Integrative Medical Sciences, Yokohama, Japan; 3Department of Technology Development, Kazusa DNA Research Institute, Kisarazu, Japan; 4Nuffield Department of Medicine, TDI Mass Spectrometry Laboratory, Target Discovery Institute, University of Oxford, Oxford, United Kingdom; 5CREST, Japan Science and Technology Agency, Kawaguchi, Japan Abstract CpG islands are gene regulatory elements associated with the majority of mammalian promoters, yet how they regulate gene expression remains poorly understood. Here, we identify *For correspondence: FBXL19 as a CpG island-binding protein in mouse embryonic stem (ES) cells and show that it [email protected] associates with the CDK-Mediator complex. We discover that FBXL19 recruits CDK-Mediator to †These authors contributed CpG island-associated promoters of non-transcribed developmental genes to prime these genes equally to this work for activation during cell lineage commitment. We further show that recognition of CpG islands by Present address: ‡Agilent FBXL19 is essential for mouse development. Together this reveals a new CpG island-centric Technologies, Waldbronn, mechanism for CDK-Mediator recruitment to developmental gene promoters in ES cells and a Germany requirement for CDK-Mediator in priming these developmental genes for activation during cell lineage commitment. Competing interests: The DOI: https://doi.org/10.7554/eLife.37084.001 authors declare that no competing interests exist. -

Live Imaging of Nascent RNA Dynamics Reveals Distinct Types of Transcriptional Pulse Regulation
Live imaging of nascent RNA dynamics reveals distinct types of transcriptional pulse regulation Tetsuya Muramotoa,1, Danielle Cannona,2, Marek Gierlinski b,c, Adam Corrigana,2, Geoffrey J. Bartonb,c, and Jonathan R. Chubba,2,3 aDivision of Cell and Developmental Biology, bDivision of Biological Chemistry and Drug Discovery, and cWellcome Trust Centre for Gene Regulation and Expression, College of Life Sciences, University of Dundee, Dundee DD1 5EH, United Kingdom Edited by Sanjay Tyagi, University of Medicine and Dentistry of New Jersey, Newark, NJ, and accepted by the Editorial Board March 21, 2012 (received for review October 25, 2011) Transcription of genes can be discontinuous, occurring in pulses or have different fluctuation kinetics. However, available data on bursts. It is not clear how properties of transcriptional pulses vary transcription bursts and pulses imply active transcriptional states between different genes. We compared the pulsing of five house- last more in the range of minutes, even for strongly transcribed keeping and five developmentally induced genes by direct imaging genes (1). To describe the dynamics of transcription pulses it is of single gene transcriptional events in individual living Dictyoste- therefore necessary to directly observe pulses of RNA production, lium cells. Each gene displayed its own transcriptional signature, which requires using high-affinity RNA–protein interactions to differing in probability of firing and pulse duration, frequency, and deliver fluorescent signals to nascent RNA (15, 16). The resultant intensity. In contrast to the prevailing view from both prokary- accumulation of fluorescence at the site of transcription is viewed otes and eukaryotes that transcription displays binary behavior, under a microscope as a fluorescent spot, which appears and dis- strongly expressed housekeeping genes altered the magnitude of appears (pulses) at irregular intervals when imaged in living cells. -
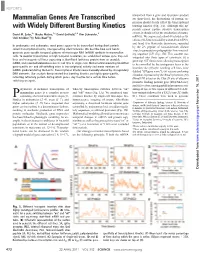
Mammalian Genes Are Transcribed with Widely Different Bursting Kinetics
REPORTS transcribed from a gene and its protein product Mammalian Genes Are Transcribed are short-lived, the fluctuations of protein ex- pression should closely reflect the transcriptional with Widely Different Bursting Kinetics bursting kinetics (Fig. 1A). Although this ap- proach cannot capture abortive transcription events, it should reflect the production of mature David M. Suter,1* Nacho Molina,2* David Gatfield,1,3 Kim Schneider,1 mRNAs. We engineered a short-lived nuclear lu- Ueli Schibler,1†‡ Felix Naef2†‡ ciferase (NLS-luc) encoded by a short-lived mRNA, and fused it to blasticidin deaminase, separated In prokaryotes and eukaryotes, most genes appear to be transcribed during short periods by the 2A peptide of foot-and-mouth disease called transcriptional bursts, interspersed by silent intervals. We describe how such bursts virus, to generate two polypeptides from one cod- generate gene-specific temporal patterns of messenger RNA (mRNA) synthesis in mammalian ing sequence (18) (Fig. 1B). This cassette was cells. To monitor transcription at high temporal resolution, we established various gene trap cell integrated into three types of constructs: (i) a lines and transgenic cell lines expressing a short-lived luciferase protein from an unstable gene trap (GT) lentivector, allowing transcription mRNA, and recorded bioluminescence in real time in single cells. Mathematical modeling identified to be controlled by the endogenous locus at the gene-specific on- and off-switching rates in transcriptional activity and mean numbers of insertion site (19) (the resulting cell lines were mRNAs produced during the bursts. Transcriptional kinetics were markedly altered by cis-regulatory dubbed “GT:gene name”); (ii) vectors conferring DNA elements. -

The Route to Transcription Initiation Determines the Mode of Transcriptional Bursting in E
The route to transcription initiation determines the mode of transcriptional bursting in E. coli. Christoph Engl1,5,6, Goran Jovanovic2,3,4,5, Rowan D Brackston3,5, Ioly Kotta-Loizou3, Martin Buck3,6 1 School of Biological & Chemical Sciences, Queen Mary University of London, London E1 4NS, UK. 2 Faculty of Medicine, Department of Medicine, Imperial College London, London SW7 2AZ, UK. 3 Faculty of Natural Sciences, Department of Life Sciences, Imperial College London, London SW7 2AZ, UK. 4 present address: Institute of Molecular Genetics and Genetic Engineering, University of Belgrade, Vojvode Stepe 444a, 11042, Belgrade, Serbia 5 these authors contributed equally to this work. 6 corresponding authors: Dr Christoph Engl, [email protected]; Prof Martin Buck, [email protected] Abstract Transcription is fundamentally noisy, leading to significant heterogeneity across bacterial populations. Noise is often attributed to burstiness, but the underlying mechanisms and their dependence on the mode of promotor regulation remain unclear. Here, we measure E. coli single cell mRNA levels for two stress responses that depend on bacterial sigma factors with different mode of transcription initiation (70 and 54). By fitting a stochastic model to the observed mRNA distributions, we show that the transition from low to high expression of the 70-controlled stress response is regulated via the burst size, while that of the 54-controlled stress response is regulated via the burst frequency. Therefore, transcription initiation involving 54 differs from other bacterial systems, and yields bursting kinetics characteristic of eukaryotic systems. Keywords Transcription; noise; burst kinetics; enhancer; sigma factor; bacteria; stress response; RNA fish; 1 Introduction Transcription is a series of discrete interactions of transcription factors and RNA polymerase with the promoter alongside any biochemical steps that these factors undertake1,2. -

MED26 Antibody
Product Datasheet MED26 Antibody Catalog No: #33633 Package Size: #33633-1 50ul #33633-2 100ul Orders: [email protected] Support: [email protected] Description Product Name MED26 Antibody Host Species Rabbit Clonality Polyclonal Purification The antibody was affinity-purified from rabbit antiserum by affinity-chromatography using epitope-specific immunogen. Applications WB Species Reactivity Hu Ms Specificity The antibody detects endogenous levels of total MED26 protein. Immunogen Type Peptide Immunogen Description Synthesized peptide derived from N-terminal of human MED26. Target Name MED26 Other Names ARC70; CRSP complex 7; CRSP70; MED26; activator-recruited cofactor 70 kDa component Accession No. Swiss-Prot: O95402NCBI Gene ID: 9441 SDS-PAGE MW 65kd Concentration 1.0mg/ml Formulation Rabbit IgG in phosphate buffered saline (without Mg2+ and Ca2+), pH 7.4, 150mM NaCl, 0.02% sodium azide and 50% glycerol. Storage Store at -20°C Application Details Western blotting: 1:500~1:3000 Images Western blot analysis of extracts from LOVO cells, using MED26 antibody #33633. Background Component of the Mediator complex, a coactivator involved in the regulated transcription of nearly all RNA polymerase II-dependent genes. Mediator Address: 8400 Baltimore Ave., Suite 302, College Park, MD 20740, USA http://www.sabbiotech.com 1 functions as a bridge to convey information from gene-specific regulatory proteins to the basal RNA polymerase II transcription machinery. Mediator is recruited to promoters by direct interactions with regulatory proteins and serves as a scaffold for the assembly of a functional preinitiation complex with RNA polymerase II and the general transcription factors. Ryu S., Nature 397:446-450(1999). Naeaer A.M., Nature 398:828-832(1999). -

Cbx3 Maintains Lineage Specificity During Neural Differentiation
Downloaded from genesdev.cshlp.org on September 27, 2021 - Published by Cold Spring Harbor Laboratory Press RESEARCH COMMUNICATION et al. 2013). However, Cbx3 is also enriched at promoters Cbx3 maintains lineage of mouse embryonic fibroblast (MEF)-derived pre-iPSCs specificity during neural (preinduced pluripotent stem cells), where it does not cor- relate with H3K9 methylation (Sridharan et al. 2013). differentiation RNAi knockdown of Cbx3 promotes reprogramming of fi- broblasts to iPSCs (Sridharan et al. 2013). Chengyang Huang,1,2 Trent Su,1 Yong Xue,1 1 3 1 Cbx3 plays important roles in developmental processes Chen Cheng, Fides D. Lay, Robin A. McKee, (Morikawa et al. 2013). In model systems, Cbx3 promotes Meiyang Li,2 Ajay Vashisht,1 James Wohlschlegel,1 neuronal maturation, kidney development, differentia- Bennett G. Novitch,4 Kathrin Plath,1 tion of ESCs to smooth muscle in culture, and embryonic Siavash K. Kurdistani,1 and Michael Carey1 arteriogenesis (Xiao et al. 2011; Dihazi et al. 2015; Oshiro et al. 2015). Little is known of how Cbx3 functions in dif- 1Department of Biological Chemistry, Eli and Edythe Broad ferentiation, but its promoter localization suggested that Center for Regenerative Medicine and Stem Cell Research, David it might directly affect transcription through the preinitia- Geffen School of Medicine, University of California at Los tion complex (PIC) (Grunberg and Hahn 2013; Allen and Angeles, Los Angeles California 90095, USA; 2Department of Taatjes 2015). The PIC is assembled in response to activa- Neurobiology, Shantou University Medical College, Shantou tors and requires the Mediator coactivator complex to re- 515041, China; 3Department of Molecular, Cell, and cruit the general transcription factors and Pol II (Chen Developmental Biology, University of California at Los Angeles, et al. -

Transcriptional Bursting Shape Autosomal Dynamic Random Monoallelic Expression in Pre-Gastrulation Embryos Naik C H, Chandel D, Mandal S, and Gayen S*
bioRxiv preprint doi: https://doi.org/10.1101/2020.09.18.303776; this version posted September 19, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. Transcriptional bursting shape autosomal dynamic random monoallelic expression in pre-gastrulation embryos Naik C H, Chandel D, Mandal S, and Gayen S* Department of Molecular Reproduction, Development and Genetics, Indian Institute of Science, Bangalore-560012, India. *Correspondence: [email protected] Abstract Recent years, allele-specific single cell RNA-seq (scRNA-seq) analysis have demonstrated wide-spread dynamic random monoallelic expression of autosomal genes (aRME). However, the origin of dynamic aRME remains poorly understood. It is believed that dynamic aRME is originated from discrete transcriptional burst of two alleles. Here, for the first time, we have profiled genome-wide pattern of dynamic aRME and allele-specific burst kinetics in mouse pre-gastrulation embryos. We found wide-spread dynamic aRME across the different lineages of pre-gastrulation embryos and which is linked to the allelic burst kinetics. Specially, we found that expression level and burst frequency are the key determinants of dynamic aRME. Altogether, our study provides significant insight about the origin of prevalent dynamic aRME and cell to cell expression heterogeneity during the early mammalian development. Keywords: Autosomal random monoallelic expression (aRME), Transcriptional burst, RNA, Pre-gastrulation, Epiblast, Visceral endoderm (VE), Extraembryonic ectoderm (ExE), Single cell RNA-Seq. Introduction Recent advances on allele-specific single cell RNA-seq (scRNA-seq) have revealed cell to cell dramatic variation of allelic gene expression pattern (Deng et al., 2014; Gendrel et al., 2016; Gregg, 2017; Reinius and Sandberg, 2015; Reinius et al., 2016). -

The Mammalian Mediator Complex
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector FEBS Letters 579 (2005) 904–908 FEBS 29061 Minireview The mammalian Mediator complex Joan Weliky Conawaya,b,c,*, Laurence Florensa, Shigeo Satoa, Chieri Tomomori-Satoa, Tari J. Parmelya, Tingting Yaoa, Selene K. Swansona, Charles A.S. Banksa, Michael P. Washburna, Ronald C. Conawaya,b a Stowers Institute for Medical Research, Kansas City, MO 64110, USA b Department of Biochemistry and Molecular Biology, Kansas University Medical Center, Kansas City, KS 66160, USA c Department of Biochemistry and Molecular Biology, University of Oklahoma Health Sciences Center, Oklahoma City, OK 73190, USA Received 25 October 2004; accepted 2 November 2004 Available online 24 November 2004 Edited by Gunnar von Heijne and Anders Liljas expressed in eukaryotes from yeast to man, is composed of Abstract The multiprotein Mediator (Med) complex is an evo- lutionarily conserved transcriptional regulator that plays impor- more than twenty subunits and has been named Mediator tant roles in activation and repression of RNA polymerase II (Med) for its role in mediating transcriptional signals from transcription. Prior studies identified a set of more than twenty DNA binding transcription factors bound at upstream pro- distinct polypeptides that compose the Saccharomyces cerevisiae moter elements and enhancers to RNA polymerase II and Mediator. Here we discuss efforts to characterize the subunit the general initiation factors bound at the core promoter sur- composition and associated activities of the mammalian Med rounding the transcriptional start site. complex. Ó 2004 Federation of European Biochemical Societies. Published by Elsevier B.V.