SCALE: Modeling Allele-Specific Gene Expression by Single-Cell RNA Sequencing Yuchao Jiang1, Nancy R
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Transcriptional Burst Frequency and Burst Size Are Equally Modulated Across the Human Genome
Transcriptional burst frequency and burst size are equally modulated across the human genome Roy D. Dara,b,c,1, Brandon S. Razookya,d,e,1, Abhyudai Singhd,2, Thomas V. Trimelonif, James M. McCollumf, Chris D. Coxg,h, Michael L. Simpsonb,I,3, and Leor S. Weinbergera,d,j,3 aGladstone Institutes, San Francisco, CA 94158; bCenter for Nanophase Materials Sciences, Oak Ridge National Laboratory, Oak Ridge, TN 37831; cDepartment of Physics and Astronomy, University of Tennessee, Knoxville, TN 37996; dDepartment of Chemistry and Biochemistry, University of California at San Diego, La Jolla, CA 92093; eBiophysics Graduate Group, University of California, San Francisco, CA 94158; gCenter for Environmental Biotechnology, University of Tennessee, Knoxville, TN 37996;hDepartment of Civil and Environmental Engineering, University of Tennessee, Knoxville, TN 37996; IDepartment of Materials Science and Engineering, University of Tennessee, Knoxville, TN 37996; jDepartment of Biochemistry and Biophysics, University of California, San Francisco, CA 94158; and fDepartment of Electrical and Computer Engineering, Virginia Commonwealth University, Richmond, VA 23284–3072 Edited by Jonathan S. Weissman, University of California, San Francisco, CA, and approved September 13, 2012 (received for review August 8, 2012) Gene expression occurs either as an episodic process, characterized autocorrelation time of expression fluctuations (as measured by pulsatile bursts, or as a constitutive process, characterized by by the noise autocorrelation time at half of its initial value, τ1∕2) a Poisson-like accumulation of gene products. It is not clear which (11, 12) (Fig. 1C). Although this three-dimensional noise space is mode of gene expression (constitutive versus bursty) predomi- impractical to analyze directly, different two-dimensional projec- nates across a genome or how transcriptional dynamics are influ- tions of noise space allow the quantification of rate parameters enced by genomic position and promoter sequence. -

Genome-Wide Analysis of Transcriptional Bursting-Induced Noise in Mammalian Cells
bioRxiv preprint doi: https://doi.org/10.1101/736207; this version posted August 15, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Title: Genome-wide analysis of transcriptional bursting-induced noise in mammalian cells Authors: Hiroshi Ochiai1*, Tetsutaro Hayashi2, Mana Umeda2, Mika Yoshimura2, Akihito Harada3, Yukiko Shimizu4, Kenta Nakano4, Noriko Saitoh5, Hiroshi Kimura6, Zhe Liu7, Takashi Yamamoto1, Tadashi Okamura4,8, Yasuyuki Ohkawa3, Itoshi Nikaido2,9* Affiliations: 1Graduate School of Integrated Sciences for Life, Hiroshima University, Higashi-Hiroshima, Hiroshima, 739-0046, Japan 2Laboratory for Bioinformatics Research, RIKEN BDR, Wako, Saitama, 351-0198, Japan 3Division of Transcriptomics, Medical Institute of Bioregulation, Kyushu University, Fukuoka, Fukuoka, 812-0054, Japan 4Department of Animal Medicine, National Center for Global Health and Medicine (NCGM), Tokyo, 812-0054, Japan 5Division of Cancer Biology, The Cancer Institute of JFCR, Tokyo, 135-8550, Japan 6Graduate School of Bioscience and Biotechnology, Tokyo Institute of Technology, Yokohama, Kanagawa, 226-8503, Japan 7Janelia Research Campus, Howard Hughes Medical Institute, Ashburn, VA, 20147, USA 8Section of Animal Models, Department of Infectious Diseases, National Center for Global Health and Medicine (NCGM), Tokyo, 812-0054, Japan 9Bioinformatics Course, Master’s/Doctoral Program in Life Science Innovation (T-LSI), School of Integrative and Global Majors (SIGMA), University of Tsukuba, Wako, 351-0198, Japan *Corresponding authors Corresponding authors e-mail addresses Hiroshi Ochiai, [email protected] Itoshi Nikaido, [email protected] bioRxiv preprint doi: https://doi.org/10.1101/736207; this version posted August 15, 2019. -

Live Imaging of Nascent RNA Dynamics Reveals Distinct Types of Transcriptional Pulse Regulation
Live imaging of nascent RNA dynamics reveals distinct types of transcriptional pulse regulation Tetsuya Muramotoa,1, Danielle Cannona,2, Marek Gierlinski b,c, Adam Corrigana,2, Geoffrey J. Bartonb,c, and Jonathan R. Chubba,2,3 aDivision of Cell and Developmental Biology, bDivision of Biological Chemistry and Drug Discovery, and cWellcome Trust Centre for Gene Regulation and Expression, College of Life Sciences, University of Dundee, Dundee DD1 5EH, United Kingdom Edited by Sanjay Tyagi, University of Medicine and Dentistry of New Jersey, Newark, NJ, and accepted by the Editorial Board March 21, 2012 (received for review October 25, 2011) Transcription of genes can be discontinuous, occurring in pulses or have different fluctuation kinetics. However, available data on bursts. It is not clear how properties of transcriptional pulses vary transcription bursts and pulses imply active transcriptional states between different genes. We compared the pulsing of five house- last more in the range of minutes, even for strongly transcribed keeping and five developmentally induced genes by direct imaging genes (1). To describe the dynamics of transcription pulses it is of single gene transcriptional events in individual living Dictyoste- therefore necessary to directly observe pulses of RNA production, lium cells. Each gene displayed its own transcriptional signature, which requires using high-affinity RNA–protein interactions to differing in probability of firing and pulse duration, frequency, and deliver fluorescent signals to nascent RNA (15, 16). The resultant intensity. In contrast to the prevailing view from both prokary- accumulation of fluorescence at the site of transcription is viewed otes and eukaryotes that transcription displays binary behavior, under a microscope as a fluorescent spot, which appears and dis- strongly expressed housekeeping genes altered the magnitude of appears (pulses) at irregular intervals when imaged in living cells. -
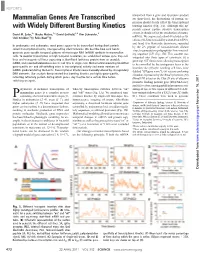
Mammalian Genes Are Transcribed with Widely Different Bursting Kinetics
REPORTS transcribed from a gene and its protein product Mammalian Genes Are Transcribed are short-lived, the fluctuations of protein ex- pression should closely reflect the transcriptional with Widely Different Bursting Kinetics bursting kinetics (Fig. 1A). Although this ap- proach cannot capture abortive transcription events, it should reflect the production of mature David M. Suter,1* Nacho Molina,2* David Gatfield,1,3 Kim Schneider,1 mRNAs. We engineered a short-lived nuclear lu- Ueli Schibler,1†‡ Felix Naef2†‡ ciferase (NLS-luc) encoded by a short-lived mRNA, and fused it to blasticidin deaminase, separated In prokaryotes and eukaryotes, most genes appear to be transcribed during short periods by the 2A peptide of foot-and-mouth disease called transcriptional bursts, interspersed by silent intervals. We describe how such bursts virus, to generate two polypeptides from one cod- generate gene-specific temporal patterns of messenger RNA (mRNA) synthesis in mammalian ing sequence (18) (Fig. 1B). This cassette was cells. To monitor transcription at high temporal resolution, we established various gene trap cell integrated into three types of constructs: (i) a lines and transgenic cell lines expressing a short-lived luciferase protein from an unstable gene trap (GT) lentivector, allowing transcription mRNA, and recorded bioluminescence in real time in single cells. Mathematical modeling identified to be controlled by the endogenous locus at the gene-specific on- and off-switching rates in transcriptional activity and mean numbers of insertion site (19) (the resulting cell lines were mRNAs produced during the bursts. Transcriptional kinetics were markedly altered by cis-regulatory dubbed “GT:gene name”); (ii) vectors conferring DNA elements. -

The Route to Transcription Initiation Determines the Mode of Transcriptional Bursting in E
The route to transcription initiation determines the mode of transcriptional bursting in E. coli. Christoph Engl1,5,6, Goran Jovanovic2,3,4,5, Rowan D Brackston3,5, Ioly Kotta-Loizou3, Martin Buck3,6 1 School of Biological & Chemical Sciences, Queen Mary University of London, London E1 4NS, UK. 2 Faculty of Medicine, Department of Medicine, Imperial College London, London SW7 2AZ, UK. 3 Faculty of Natural Sciences, Department of Life Sciences, Imperial College London, London SW7 2AZ, UK. 4 present address: Institute of Molecular Genetics and Genetic Engineering, University of Belgrade, Vojvode Stepe 444a, 11042, Belgrade, Serbia 5 these authors contributed equally to this work. 6 corresponding authors: Dr Christoph Engl, [email protected]; Prof Martin Buck, [email protected] Abstract Transcription is fundamentally noisy, leading to significant heterogeneity across bacterial populations. Noise is often attributed to burstiness, but the underlying mechanisms and their dependence on the mode of promotor regulation remain unclear. Here, we measure E. coli single cell mRNA levels for two stress responses that depend on bacterial sigma factors with different mode of transcription initiation (70 and 54). By fitting a stochastic model to the observed mRNA distributions, we show that the transition from low to high expression of the 70-controlled stress response is regulated via the burst size, while that of the 54-controlled stress response is regulated via the burst frequency. Therefore, transcription initiation involving 54 differs from other bacterial systems, and yields bursting kinetics characteristic of eukaryotic systems. Keywords Transcription; noise; burst kinetics; enhancer; sigma factor; bacteria; stress response; RNA fish; 1 Introduction Transcription is a series of discrete interactions of transcription factors and RNA polymerase with the promoter alongside any biochemical steps that these factors undertake1,2. -

Transcriptional Bursting Shape Autosomal Dynamic Random Monoallelic Expression in Pre-Gastrulation Embryos Naik C H, Chandel D, Mandal S, and Gayen S*
bioRxiv preprint doi: https://doi.org/10.1101/2020.09.18.303776; this version posted September 19, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. Transcriptional bursting shape autosomal dynamic random monoallelic expression in pre-gastrulation embryos Naik C H, Chandel D, Mandal S, and Gayen S* Department of Molecular Reproduction, Development and Genetics, Indian Institute of Science, Bangalore-560012, India. *Correspondence: [email protected] Abstract Recent years, allele-specific single cell RNA-seq (scRNA-seq) analysis have demonstrated wide-spread dynamic random monoallelic expression of autosomal genes (aRME). However, the origin of dynamic aRME remains poorly understood. It is believed that dynamic aRME is originated from discrete transcriptional burst of two alleles. Here, for the first time, we have profiled genome-wide pattern of dynamic aRME and allele-specific burst kinetics in mouse pre-gastrulation embryos. We found wide-spread dynamic aRME across the different lineages of pre-gastrulation embryos and which is linked to the allelic burst kinetics. Specially, we found that expression level and burst frequency are the key determinants of dynamic aRME. Altogether, our study provides significant insight about the origin of prevalent dynamic aRME and cell to cell expression heterogeneity during the early mammalian development. Keywords: Autosomal random monoallelic expression (aRME), Transcriptional burst, RNA, Pre-gastrulation, Epiblast, Visceral endoderm (VE), Extraembryonic ectoderm (ExE), Single cell RNA-Seq. Introduction Recent advances on allele-specific single cell RNA-seq (scRNA-seq) have revealed cell to cell dramatic variation of allelic gene expression pattern (Deng et al., 2014; Gendrel et al., 2016; Gregg, 2017; Reinius and Sandberg, 2015; Reinius et al., 2016). -

Using Dynamics and Theory to Uncover Mechanisms of Transcriptional Bursting
A matter of time: Using dynamics and theory to uncover mechanisms of transcriptional bursting Nicholas C Lammers1, Yang Joon Kim1,∗, Jiaxi Zhao2,∗, Hernan G Garcia1,2,3,4,∗∗ Abstract Eukaryotic transcription generally occurs in bursts of activity lasting minutes to hours; however, state-of-the-art measurements have revealed that many of the molec- ular processes that underlie bursting, such as transcription factor binding to DNA, unfold on timescales of seconds. This temporal disconnect lies at the heart of a broader challenge in physical biology of predicting transcriptional outcomes and cel- lular decision-making from the dynamics of underlying molecular processes. Here, we review how new dynamical information about the processes underlying transcrip- tional control can be combined with theoretical models that predict not only averaged transcriptional dynamics, but also their variability, to formulate testable hypotheses about the molecular mechanisms underlying transcriptional bursting and control. Keywords: Live imaging, Transcriptional bursting, Gene regulation, Transcriptional dynamics, Theoretical models of transcription, Non-equilibrium models of transcription, Waiting time distributions 1. A disconnect between transcriptional bursting and its underlying molec- ular processes Over the past two decades, new technologies have revealed that transcription is a fundamentally discontinuous process characterized by transient bursts of transcrip- ∗These authors contributed equally to this work. arXiv:2008.09225v2 [q-bio.SC] 17 Dec 2020 ∗∗For correspondence: [email protected] (HGG) 1Biophysics Graduate Group, University of California at Berkeley, Berkeley, California 2Department of Physics, University of California at Berkeley, Berkeley, California 3Department of Molecular and Cell Biology, University of California at Berkeley, Berkeley, California 4Institute for Quantitative Biosciences-QB3, University of California at Berkeley, Berkeley, California tional activity interspersed with periods of quiescence. -

Gene Networks with Transcriptional Bursting Recapitulate Rare Transient Coordinated Expression States in Cancer
bioRxiv preprint doi: https://doi.org/10.1101/704247; this version posted July 18, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Gene networks with transcriptional bursting recapitulate rare transient coordinated expression states in cancer 1,2,3 4 1 1 Lea Schuh , Michael Saint-Antoine , Eric Sanford , Benjamin L. Emert , Abhyudai 4 2 1,* 1,*,** Singh , Carsten Marr , Yogesh Goyal , Arjun Raj 1 Department of Bioengineering, University of Pennsylvania, Philadelphia, Pennsylvania 19104, USA 2 Institute of Computational Biology, Helmholtz Zentrum München, Neuherberg, 85764, Germany 3 Department of Mathematics, Technical University of Munich, Garching, 85748, Germany 4 Electrical and Computer Engineering, University of Delaware, Newark, Delaware 19716, USA *Authors for correspondence: [email protected], [email protected], **Lead contact: [email protected] SUMMARY Non-genetic transcriptional variability at the single-cell level is a potential mechanism for therapy resistance in melanoma. Specifically, rare subpopulations of melanoma cells occupy a transient pre-resistant state characterized by coordinated high expression of several genes. Importantly, these rare cells are able to survive drug treatment and develop resistance. How might these extremely rare states arise and disappear within the population? It is unclear whether the canonical stochastic models of probabilistic transcriptional pulsing can explain this behavior, or if it requires special, hitherto unidentified molecular mechanisms. Here we use mathematical modeling to show that a minimal network comprising of transcriptional bursting and interactions between genes can give rise to rare coordinated high states. -

Tunable Stochastic Pulsing in the Escherichia Coli Multiple Antibiotic Resistance Network from Interlinked Positive and Negative Feedback Loops
Tunable Stochastic Pulsing in the Escherichia coli Multiple Antibiotic Resistance Network from Interlinked Positive and Negative Feedback Loops Javier Garcia-Bernardo, Mary J. Dunlop* School of Engineering, University of Vermont, Burlington, Vermont, United States of America Abstract Cells live in uncertain, dynamic environments and have many mechanisms for sensing and responding to changes in their surroundings. However, sudden fluctuations in the environment can be catastrophic to a population if it relies solely on sensory responses, which have a delay associated with them. Cells can reconcile these effects by using a tunable stochastic response, where in the absence of a stressor they create phenotypic diversity within an isogenic population, but use a deterministic response when stressors are sensed. Here, we develop a stochastic model of the multiple antibiotic resistance network of Escherichia coli and show that it can produce tunable stochastic pulses in the activator MarA. In particular, we show that a combination of interlinked positive and negative feedback loops plays an important role in setting the dynamics of the stochastic pulses. Negative feedback produces a pulsatile response that is tunable, while positive feedback serves to amplify the effect. Our simulations show that the uninduced native network is in a parameter regime that is of low cost to the cell (taxing resistance mechanisms are expressed infrequently) and also elevated noise strength (phenotypic variability is high). The stochastic pulsing can be tuned by MarA induction such that variability is decreased once stresses are sensed, avoiding the detrimental effects of noise when an optimal MarA concentration is needed. We further show that variability in the expression of MarA can act as a bet hedging mechanism, allowing for survival in time-varying stress environments, however this effect is tunable to allow for a fully induced, deterministic response in the presence of a stressor. -

Diverse Spatial Expression Patterns Emerge from Common Transcription Bursting Kinetics
Diverse spatial expression patterns emerge from common transcription bursting kinetics Benjamin Zollera∗, Shawn C. Littleb∗, Thomas Gregoray aJoseph Henry Laboratories of Physics and the Lewis Sigler Institute for Integrative Genomics, Princeton University, Princeton, NJ 08544, USA bDepartment of Cell and Developmental Biology, University of Pennsylvania Perelman School of Medicine, Philadelphia, PA 19143, USA (Dated: December 25, 2017) In early development, regulation of transcription results in precisely positioned and highly repro- ducible expression patterns that specify cellular identities. How transcription, a fundamentally noisy molecular process, is regulated to achieve reliable embryonic patterning remains unclear. In par- ticular, it is unknown how gene-specific regulation mechanisms affect kinetic rates of transcription, and whether there are common, global features that govern these rates across a genetic network. Here, we measure nascent transcriptional activity in the gap gene network of early Drosophila em- bryos and characterize the variability in absolute activity levels across expression boundaries. We demonstrate that boundary formation follows a common transcriptional principle: a single control parameter determines the distribution of transcriptional activity, regardless of gene identity, bound- ary position, or enhancer-promoter architecture. By employing a minimalist model of transcription, we infer kinetic rates of transcriptional bursting for these patterning genes; we find that the key regulatory parameter is the fraction of time a gene is in an actively transcribing state, while the rate of Pol II loading appears globally conserved. These results point to a universal simplicity under- lying the apparently complex transcriptional processes responsible for early embryonic patterning and indicate a path to general rules in transcriptional regulation. I. -

Transcriptional Burst Frequency and Burst Size Are Equally Modulated Across the Human Genome
Transcriptional burst frequency and burst size are equally modulated across the human genome Roy D. Dara,b,c,1, Brandon S. Razookya,d,e,1, Abhyudai Singhd,2, Thomas V. Trimelonif, James M. McCollumf, Chris D. Coxg,h, Michael L. Simpsonb,I,3, and Leor S. Weinbergera,d,j,3 aGladstone Institutes, San Francisco, CA 94158; bCenter for Nanophase Materials Sciences, Oak Ridge National Laboratory, Oak Ridge, TN 37831; cDepartment of Physics and Astronomy, University of Tennessee, Knoxville, TN 37996; dDepartment of Chemistry and Biochemistry, University of California at San Diego, La Jolla, CA 92093; eBiophysics Graduate Group, University of California, San Francisco, CA 94158; gCenter for Environmental Biotechnology, University of Tennessee, Knoxville, TN 37996;hDepartment of Civil and Environmental Engineering, University of Tennessee, Knoxville, TN 37996; IDepartment of Materials Science and Engineering, University of Tennessee, Knoxville, TN 37996; jDepartment of Biochemistry and Biophysics, University of California, San Francisco, CA 94158; and fDepartment of Electrical and Computer Engineering, Virginia Commonwealth University, Richmond, VA 23284–3072 Edited by Jonathan S. Weissman, University of California, San Francisco, CA, and approved September 13, 2012 (received for review August 8, 2012) Gene expression occurs either as an episodic process, characterized autocorrelation time of expression fluctuations (as measured by pulsatile bursts, or as a constitutive process, characterized by by the noise autocorrelation time at half of its initial value, τ1∕2) a Poisson-like accumulation of gene products. It is not clear which (11, 12) (Fig. 1C). Although this three-dimensional noise space is mode of gene expression (constitutive versus bursty) predomi- impractical to analyze directly, different two-dimensional projec- nates across a genome or how transcriptional dynamics are influ- tions of noise space allow the quantification of rate parameters enced by genomic position and promoter sequence. -

A Biophysical Model of Supercoiling Dependent Transcription Predicts a Structural Aspect to Gene Regulation Christopher H
Bohrer and Roberts BMC Biophysics (2016) 9:2 DOI 10.1186/s13628-016-0027-0 RESEARCH ARTICLE Open Access A biophysical model of supercoiling dependent transcription predicts a structural aspect to gene regulation Christopher H. Bohrer and Elijah Roberts* Abstract Background: Transcription in Escherichia coli generates positive supercoiling in the DNA, which is relieved by the enzymatic activity of gyrase. Recently published experimental evidence suggests that transcription initiation and elongation are inhibited by the buildup of positive supercoiling. It has therefore been proposed that intermittent binding of gyrase plays a role in transcriptional bursting. Considering that transcription is one of the most fundamental cellular processes, it is desirable to be able to account for the buildup and release of positive supercoiling in models of transcription. Results: Here we present a detailed biophysical model of gene expression that incorporates the effects of supercoiling due to transcription. By directly linking the amount of positive supercoiling to the rate of transcription, the model predicts that highly transcribed genes’ mRNA distributions should substantially deviate from Poisson distributions, with enhanced density at low mRNA copy numbers. Additionally, the model predicts a high degree of correlation between expression levels of genes inside the same supercoiling domain. Conclusions: Our model, incorporating the supercoiling state of the gene, makes specific predictions that differ from previous models of gene expression. Genes in the same supercoiling domain influence the expression level of neighboring genes. Such structurally dependent regulation predicts correlations between genes in the same supercoiling domain. The topology of the chromosome therefore creates a higher level of gene regulation, which has broad implications for understanding the evolution and organization of bacterial genomes.