Control Induction of Osmoprotective Genes in Euryhaline Fish
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

OSTF1 (AT9G4): Sc-517418
SAN TA C RUZ BI OTEC HNOL OG Y, INC . OSTF1 (AT9G4): sc-517418 BACKGROUND PRODUCT OSTF1 (osteoclast-stimulating factor 1) is a 214 amino acid cytoplasmic pro - Each vial contains 100 µg IgG 2a kappa light chain in 1.0 ml of PBS with tein that contains 3 ANK repeats and one SH3 domain. The SH3 domain < 0.1% sodium azide and 0.1% gelatin. mediates interaction of OSTF1 with SMN. In addition to SMN, OSTF1 inter - acts with Src and FAS-L. In osteoclasts, OSTF1 is thought to induce bone APPLICATIONS resorption most likely through a signaling cascade that results in the secre - OSTF1 (AT9G4) is recommended for detection of OSTF1 of mouse, rat and tion of factors that enhance osteoclast formation and activity. The gene that human origin by Western Blotting (starting dilution 1:200, dilution range encdoes OSTF1 consists of almost 59,000 bases and maps to human chro mo - 1:100-1:1000), immunoprecipitation [1-2 µg per 100-500 µg of total protein some 9q21.13. Housing over 900 genes, chromosome 9 comprises nearly 4% (1 ml of cell lysate)], immunofluorescence (starting dilution 1:50, dilution of the human genome. Hereditary hemorrhagic telangiectasia, which is char - range 1:50-1:500), flow cytometry (1 µg per 1 x 10 6 cells) and solid phase acterized by harmful vascular defects, and familial dysautonomia, are both ELISA (starting dilution 1:30, dilution range 1:30-1:3000). associated with chromosome 9. Notably, chromosome 9 encompasses the largest interferon family gene cluster. Suitable for use as control antibody for OSTF1 siRNA (h): sc-92800, OSTF1 siRNA (m): sc-151336, OSTF1 shRNA Plasmid (h): sc-92800-SH, OSTF1 REFERENCES shRNA Plasmid (m): sc-151336-SH, OSTF1 shRNA (h) Lentiviral Particles: sc-92800-V and OSTF1 shRNA (m) Lentiviral Particles: sc-151336-V. -

The Landscape of Human Mutually Exclusive Splicing
bioRxiv preprint doi: https://doi.org/10.1101/133215; this version posted May 2, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. The landscape of human mutually exclusive splicing Klas Hatje1,2,#,*, Ramon O. Vidal2,*, Raza-Ur Rahman2, Dominic Simm1,3, Björn Hammesfahr1,$, Orr Shomroni2, Stefan Bonn2§ & Martin Kollmar1§ 1 Group of Systems Biology of Motor Proteins, Department of NMR-based Structural Biology, Max-Planck-Institute for Biophysical Chemistry, Göttingen, Germany 2 Group of Computational Systems Biology, German Center for Neurodegenerative Diseases, Göttingen, Germany 3 Theoretical Computer Science and Algorithmic Methods, Institute of Computer Science, Georg-August-University Göttingen, Germany § Corresponding authors # Current address: Roche Pharmaceutical Research and Early Development, Pharmaceutical Sciences, Roche Innovation Center Basel, F. Hoffmann-La Roche Ltd., Basel, Switzerland $ Current address: Research and Development - Data Management (RD-DM), KWS SAAT SE, Einbeck, Germany * These authors contributed equally E-mail addresses: KH: [email protected], RV: [email protected], RR: [email protected], DS: [email protected], BH: [email protected], OS: [email protected], SB: [email protected], MK: [email protected] - 1 - bioRxiv preprint doi: https://doi.org/10.1101/133215; this version posted May 2, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. -

WO 2012/174282 A2 20 December 2012 (20.12.2012) P O P C T
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date WO 2012/174282 A2 20 December 2012 (20.12.2012) P O P C T (51) International Patent Classification: David [US/US]; 13539 N . 95th Way, Scottsdale, AZ C12Q 1/68 (2006.01) 85260 (US). (21) International Application Number: (74) Agent: AKHAVAN, Ramin; Caris Science, Inc., 6655 N . PCT/US20 12/0425 19 Macarthur Blvd., Irving, TX 75039 (US). (22) International Filing Date: (81) Designated States (unless otherwise indicated, for every 14 June 2012 (14.06.2012) kind of national protection available): AE, AG, AL, AM, AO, AT, AU, AZ, BA, BB, BG, BH, BR, BW, BY, BZ, English (25) Filing Language: CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, DO, Publication Language: English DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, HN, HR, HU, ID, IL, IN, IS, JP, KE, KG, KM, KN, KP, KR, (30) Priority Data: KZ, LA, LC, LK, LR, LS, LT, LU, LY, MA, MD, ME, 61/497,895 16 June 201 1 (16.06.201 1) US MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, 61/499,138 20 June 201 1 (20.06.201 1) US OM, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SC, SD, 61/501,680 27 June 201 1 (27.06.201 1) u s SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, TR, 61/506,019 8 July 201 1(08.07.201 1) u s TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. -

WO 2016/040794 Al 17 March 2016 (17.03.2016) P O P C T
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date WO 2016/040794 Al 17 March 2016 (17.03.2016) P O P C T (51) International Patent Classification: AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, C12N 1/19 (2006.01) C12Q 1/02 (2006.01) BZ, CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, C12N 15/81 (2006.01) C07K 14/47 (2006.01) DO, DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, HN, HR, HU, ID, IL, IN, IR, IS, JP, KE, KG, KN, KP, KR, (21) International Application Number: KZ, LA, LC, LK, LR, LS, LU, LY, MA, MD, ME, MG, PCT/US20 15/049674 MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, OM, (22) International Filing Date: PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SA, SC, 11 September 2015 ( 11.09.201 5) SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. (25) Filing Language: English (84) Designated States (unless otherwise indicated, for every (26) Publication Language: English kind of regional protection available): ARIPO (BW, GH, (30) Priority Data: GM, KE, LR, LS, MW, MZ, NA, RW, SD, SL, ST, SZ, 62/050,045 12 September 2014 (12.09.2014) US TZ, UG, ZM, ZW), Eurasian (AM, AZ, BY, KG, KZ, RU, TJ, TM), European (AL, AT, BE, BG, CH, CY, CZ, DE, (71) Applicant: WHITEHEAD INSTITUTE FOR BIOMED¬ DK, EE, ES, FI, FR, GB, GR, HR, HU, IE, IS, IT, LT, LU, ICAL RESEARCH [US/US]; Nine Cambridge Center, LV, MC, MK, MT, NL, NO, PL, PT, RO, RS, SE, SI, SK, Cambridge, Massachusetts 02142-1479 (US). -

UC San Diego Electronic Theses and Dissertations
UC San Diego UC San Diego Electronic Theses and Dissertations Title Cardiac Stretch-Induced Transcriptomic Changes are Axis-Dependent Permalink https://escholarship.org/uc/item/7m04f0b0 Author Buchholz, Kyle Stephen Publication Date 2016 Peer reviewed|Thesis/dissertation eScholarship.org Powered by the California Digital Library University of California UNIVERSITY OF CALIFORNIA, SAN DIEGO Cardiac Stretch-Induced Transcriptomic Changes are Axis-Dependent A dissertation submitted in partial satisfaction of the requirements for the degree Doctor of Philosophy in Bioengineering by Kyle Stephen Buchholz Committee in Charge: Professor Jeffrey Omens, Chair Professor Andrew McCulloch, Co-Chair Professor Ju Chen Professor Karen Christman Professor Robert Ross Professor Alexander Zambon 2016 Copyright Kyle Stephen Buchholz, 2016 All rights reserved Signature Page The Dissertation of Kyle Stephen Buchholz is approved and it is acceptable in quality and form for publication on microfilm and electronically: Co-Chair Chair University of California, San Diego 2016 iii Dedication To my beautiful wife, Rhia. iv Table of Contents Signature Page ................................................................................................................... iii Dedication .......................................................................................................................... iv Table of Contents ................................................................................................................ v List of Figures ................................................................................................................... -

Content Based Search in Gene Expression Databases and a Meta-Analysis of Host Responses to Infection
Content Based Search in Gene Expression Databases and a Meta-analysis of Host Responses to Infection A Thesis Submitted to the Faculty of Drexel University by Francis X. Bell in partial fulfillment of the requirements for the degree of Doctor of Philosophy November 2015 c Copyright 2015 Francis X. Bell. All Rights Reserved. ii Acknowledgments I would like to acknowledge and thank my advisor, Dr. Ahmet Sacan. Without his advice, support, and patience I would not have been able to accomplish all that I have. I would also like to thank my committee members and the Biomed Faculty that have guided me. I would like to give a special thanks for the members of the bioinformatics lab, in particular the members of the Sacan lab: Rehman Qureshi, Daisy Heng Yang, April Chunyu Zhao, and Yiqian Zhou. Thank you for creating a pleasant and friendly environment in the lab. I give the members of my family my sincerest gratitude for all that they have done for me. I cannot begin to repay my parents for their sacrifices. I am eternally grateful for everything they have done. The support of my sisters and their encouragement gave me the strength to persevere to the end. iii Table of Contents LIST OF TABLES.......................................................................... vii LIST OF FIGURES ........................................................................ xiv ABSTRACT ................................................................................ xvii 1. A BRIEF INTRODUCTION TO GENE EXPRESSION............................. 1 1.1 Central Dogma of Molecular Biology........................................... 1 1.1.1 Basic Transfers .......................................................... 1 1.1.2 Uncommon Transfers ................................................... 3 1.2 Gene Expression ................................................................. 4 1.2.1 Estimating Gene Expression ............................................ 4 1.2.2 DNA Microarrays ...................................................... -

Supplementary Table 1 Double Treatment Vs Single Treatment
Supplementary table 1 Double treatment vs single treatment Probe ID Symbol Gene name P value Fold change TC0500007292.hg.1 NIM1K NIM1 serine/threonine protein kinase 1.05E-04 5.02 HTA2-neg-47424007_st NA NA 3.44E-03 4.11 HTA2-pos-3475282_st NA NA 3.30E-03 3.24 TC0X00007013.hg.1 MPC1L mitochondrial pyruvate carrier 1-like 5.22E-03 3.21 TC0200010447.hg.1 CASP8 caspase 8, apoptosis-related cysteine peptidase 3.54E-03 2.46 TC0400008390.hg.1 LRIT3 leucine-rich repeat, immunoglobulin-like and transmembrane domains 3 1.86E-03 2.41 TC1700011905.hg.1 DNAH17 dynein, axonemal, heavy chain 17 1.81E-04 2.40 TC0600012064.hg.1 GCM1 glial cells missing homolog 1 (Drosophila) 2.81E-03 2.39 TC0100015789.hg.1 POGZ Transcript Identified by AceView, Entrez Gene ID(s) 23126 3.64E-04 2.38 TC1300010039.hg.1 NEK5 NIMA-related kinase 5 3.39E-03 2.36 TC0900008222.hg.1 STX17 syntaxin 17 1.08E-03 2.29 TC1700012355.hg.1 KRBA2 KRAB-A domain containing 2 5.98E-03 2.28 HTA2-neg-47424044_st NA NA 5.94E-03 2.24 HTA2-neg-47424360_st NA NA 2.12E-03 2.22 TC0800010802.hg.1 C8orf89 chromosome 8 open reading frame 89 6.51E-04 2.20 TC1500010745.hg.1 POLR2M polymerase (RNA) II (DNA directed) polypeptide M 5.19E-03 2.20 TC1500007409.hg.1 GCNT3 glucosaminyl (N-acetyl) transferase 3, mucin type 6.48E-03 2.17 TC2200007132.hg.1 RFPL3 ret finger protein-like 3 5.91E-05 2.17 HTA2-neg-47424024_st NA NA 2.45E-03 2.16 TC0200010474.hg.1 KIAA2012 KIAA2012 5.20E-03 2.16 TC1100007216.hg.1 PRRG4 proline rich Gla (G-carboxyglutamic acid) 4 (transmembrane) 7.43E-03 2.15 TC0400012977.hg.1 SH3D19 -

Genetic Circuitry of Survival Motor Neuron, the Gene Underlying Spinal
Genetic circuitry of Survival motor neuron, the gene PNAS PLUS underlying spinal muscular atrophy Anindya Sena,1, Douglas N. Dimlicha,1, K. G. Guruharshaa,1, Mark W. Kankela,1, Kazuya Horia, Takakazu Yokokuraa,3, Sophie Brachatb,c, Delwood Richardsonb, Joseph Loureirob, Rajeev Sivasankaranb, Daniel Curtisb, Lance S. Davidowd, Lee L. Rubind, Anne C. Harte, David Van Vactora, and Spyros Artavanis-Tsakonasa,2 aDepartment of Cell Biology, Harvard Medical School, Boston, MA 02115; bDevelopmental and Molecular Pathways, Novartis Institutes for Biomedical Research, Cambridge, MA 02139; cMusculoskeletal Diseases, Novartis Institutes for Biomedical Research, CH-4002 Basel, Switzerland; dDepartment of Stem Cell and Regenerative Biology, Harvard Medical School, Boston, MA 02115; and eDepartment of Neuroscience, Brown University, Providence, RI 02912 Edited by Jeffrey C. Hall, University of Maine, Orono, ME, and approved May 7, 2013 (received for review February 13, 2013) The clinical severity of the neurodegenerative disorder spinal muscu- snRNP biogenesis, the molecular functionality that is most clearly lar atrophy (SMA) is dependent on the levels of functional Survival associated with SMN. Motor Neuron (SMN) protein. Consequently, current strategies for As the human disease state results from partial loss of SMN developing treatments for SMA generally focus on augmenting SMN function, we reasoned that a screening paradigm using a hypo- levels. To identify additional potential therapeutic avenues and morphic Smn background (as opposed to a background that achieve a greater understanding of SMN, we applied in vivo, in vitro, completely eliminates SMN function) would more closely resemble and in silico approaches to identify genetic and biochemical interac- the genetic condition in SMA. -

Transposon Mutagenesis Identifies Genetic Drivers of Brafv600e Melanoma
ARTICLES Transposon mutagenesis identifies genetic drivers of BrafV600E melanoma Michael B Mann1,2, Michael A Black3, Devin J Jones1, Jerrold M Ward2,12, Christopher Chin Kuan Yew2,12, Justin Y Newberg1, Adam J Dupuy4, Alistair G Rust5,12, Marcus W Bosenberg6,7, Martin McMahon8,9, Cristin G Print10,11, Neal G Copeland1,2,13 & Nancy A Jenkins1,2,13 Although nearly half of human melanomas harbor oncogenic BRAFV600E mutations, the genetic events that cooperate with these mutations to drive melanogenesis are still largely unknown. Here we show that Sleeping Beauty (SB) transposon-mediated mutagenesis drives melanoma progression in BrafV600E mutant mice and identify 1,232 recurrently mutated candidate cancer genes (CCGs) from 70 SB-driven melanomas. CCGs are enriched in Wnt, PI3K, MAPK and netrin signaling pathway components and are more highly connected to one another than predicted by chance, indicating that SB targets cooperative genetic networks in melanoma. Human orthologs of >500 CCGs are enriched for mutations in human melanoma or showed statistically significant clinical associations between RNA abundance and survival of patients with metastatic melanoma. We also functionally validate CEP350 as a new tumor-suppressor gene in human melanoma. SB mutagenesis has thus helped to catalog the cooperative molecular mechanisms driving BRAFV600E melanoma and discover new genes with potential clinical importance in human melanoma. Substantial sun exposure and numerous genetic factors, including including BrafV600E, recapitulate the genetic and histological hallmarks skin type and family history, are the most important melanoma risk of human melanoma. In these models, increased MEK-ERK signaling factors. Familial melanoma, which accounts for <10% of cases, is asso- initiates clonal expansion of melanocytes, which is limited by oncogene- ciated with mutations in CDKN2A1, MITF2 and POT1 (refs. -

START Lipid/Sterol-Binding Domains Are Amplified in Plants and Are
Open Access Research2004Schricket al.Volume 5, Issue 6, Article R41 START lipid/sterol-binding domains are amplified in plants and are comment predominantly associated with homeodomain transcription factors Kathrin Schrick*, Diana Nguyen*, Wojciech M Karlowski† and Klaus FX Mayer† Addresses: *Keck Graduate Institute of Applied Life Sciences, 535 Watson Drive, Claremont, CA 91711, USA. †Munich Information Center for Protein Sequences, Institute for Bioinformatics, GSF National Research Center for Environment and Health, Ingolstädter Landstrasse 1, 85764 Neuherberg, Germany. reviews Correspondence: Kathrin Schrick. E-mail: [email protected] Published: 27 May 2004 Received: 27 January 2004 Revised: 8 April 2004 Genome Biology 2004, 5:R41 Accepted: 30 April 2004 The electronic version of this article is the complete one and can be found online at http://genomebiology.com/2004/5/6/R41 reports © 2004 Schrick et al.; licensee BioMed Central Ltd. This is an Open Access article: verbatim copying and redistribution of this article are permitted in all media for any purpose, provided this notice is preserved along with the article's original URL. START<p>Aplants surveyand lipid/sterol-binding are of primarily proteins foundcontaining domains within lipid/sterol-binding are homeodomain amplified in (HD)plants StAR-rel transcription and areated predominantly lipid factors.</p> transfer (START)associated domains with homeodomain shows that they transcription are amplifi facted inors deposited research Abstract Background: In animals, steroid hormones regulate gene expression by binding to nuclear receptors. Plants lack genes for nuclear receptors, yet genetic evidence from Arabidopsis suggests developmental roles for lipids/sterols analogous to those in animals. In contrast to nuclear receptors, the lipid/sterol-binding StAR-related lipid transfer (START) protein domains are conserved, making them candidates for involvement in both animal and plant lipid/sterol signal transduction. -

Junl;G.~~ Assistant Professor Department of Mechanical Engineering Rice University
RICE UNIVERSITY Novel Exogenous Agents for Improving Articular Cartilage Tissue Engineering by Donald Joseph Responte A THESIS SUBMITTED IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE Doctor of Philosophy APPROVED, THESIS COMMITTEE: Kyriacos thanas u, PH. ., Thesis Director Distinguis ed Professor Department of Biomedical Engineering University of California, Davis Antonios Mikos, Ph.D., Committee Chair Louis Calder Professor Department of Bioengineering Rice University Junl;G.~~ Assistant Professor Department of Mechanical Engineering Rice University HOUSTON, TEXAS OCTOBER 2011 ii ABSTRACT Novel Exogenous Agents for Improving Articular Cartilage Tissue Engineering by Donald Joseph Responte This thesis demonstrated the effects of exogenous stimuli on engineered articular cartilage constructs and elucidated mechanisms underlying the responses to these agents. In particular, a series of studies detailed the effects of chondroitinase-ABC (C-ABC), hyaluronic acid (HA), and TGF-~1 on the biochemical and biomechanical properties of self-assembled articular cartilage. Work with C-ABC showed that this catabolic agent can be employed to improve the tensile properties of constructs. When constructs were cultured for 6 weeks, treating with C-ABC at 2 weeks enhanced the tensile stiffness. Furthermore, treating at 2 and 4 weeks synergistically increased tensile properties and allowed compressive stiffness to recover to control levels. Another study showed that combining C-ABC and TGF-~1 synergistically enhanced the biochemical and biomechanical properties of neotissue. Microarray analysis demonstrated that TGF-~1 increased MAPK signaling in self-assembled neocartilage whereas C ABC had minimal effects on gene expression. SEM analysis showed that C-ABC increased collagen fibril diameter and fibril density, indicating that C-ABC potentially acts via a biophysical mechanism. -
Downloaded from the Mouse Lysosome Gene Database, Mlgdb
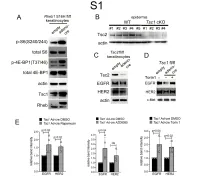
1 Supplemental Figure Legends 2 3 Supplemental Figure S1: Epidermal-specific mTORC1 gain-of-function models show 4 increased mTORC1 activation and down-regulate EGFR and HER2 protein expression in a 5 mTORC1-sensitive manner. (A) Immunoblotting of Rheb1 S16H flox/flox keratinocyte cultures 6 infected with empty or adenoviral cre recombinase for markers of mTORC1 (p-S6, p-4E-BP1) 7 activity. (B) Tsc1 cKO epidermal lysates also show decreased expression of TSC2 by 8 immunoblotting of the same experiment as in Figure 2A. (C) Immunoblotting of Tsc2 flox/flox 9 keratinocyte cultures infected with empty or adenoviral cre recombinase showing decreased EGFR 10 and HER2 protein expression. (D) Expression of EGFR and HER2 was decreased in Tsc1 cre 11 keratinocytes compared to empty controls, and up-regulated in response to Torin1 (1µM, 24 hrs), 12 by immunoblot analyses. Immunoblots are contemporaneous and parallel from the same biological 13 replicate and represent the same experiment as depicted in Figure 7B. (E) Densitometry 14 quantification of representative immunoblot experiments shown in Figures 2E and S1D (r≥3; error 15 bars represent STDEV; p-values by Student’s T-test). 16 17 18 19 20 21 22 23 Supplemental Figure S2: EGFR and HER2 transcription are unchanged with epidermal/ 24 keratinocyte Tsc1 or Rptor loss. Egfr and Her2 mRNA levels in (A) Tsc1 cKO epidermal lysates, 25 (B) Tsc1 cKO keratinocyte lysates and(C) Tsc1 cre keratinocyte lysates are minimally altered 26 compared to their respective controls. (r≥3; error bars represent STDEV; p-values by Student’s T- 27 test).