Proteomic Analysis of HIV-1 Nef Cellular Binding Partners Reveals a Role for Exocyst Complex Proteins in Mediating Enhancement of Intercellular Nanotube Formation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Transcriptomic Analysis of the Aquaporin (AQP) Gene Family
Pancreatology 19 (2019) 436e442 Contents lists available at ScienceDirect Pancreatology journal homepage: www.elsevier.com/locate/pan Transcriptomic analysis of the Aquaporin (AQP) gene family interactome identifies a molecular panel of four prognostic markers in patients with pancreatic ductal adenocarcinoma Dimitrios E. Magouliotis a, b, Vasiliki S. Tasiopoulou c, Konstantinos Dimas d, * Nikos Sakellaridis d, Konstantina A. Svokos e, Alexis A. Svokos f, Dimitris Zacharoulis b, a Division of Surgery and Interventional Science, Faculty of Medical Sciences, UCL, London, UK b Department of Surgery, University of Thessaly, Biopolis, Larissa, Greece c Faculty of Medicine, School of Health Sciences, University of Thessaly, Biopolis, Larissa, Greece d Department of Pharmacology, Faculty of Medicine, School of Health Sciences, University of Thessaly, Biopolis, Larissa, Greece e The Warren Alpert Medical School of Brown University, Providence, RI, USA f Riverside Regional Medical Center, Newport News, VA, USA article info abstract Article history: Background: This study aimed to assess the differential gene expression of aquaporin (AQP) gene family Received 14 October 2018 interactome in pancreatic ductal adenocarcinoma (PDAC) using data mining techniques to identify novel Received in revised form candidate genes intervening in the pathogenicity of PDAC. 29 January 2019 Method: Transcriptome data mining techniques were used in order to construct the interactome of the Accepted 9 February 2019 AQP gene family and to determine which genes members are differentially expressed in PDAC as Available online 11 February 2019 compared to controls. The same techniques were used in order to evaluate the potential prognostic role of the differentially expressed genes. Keywords: PDAC Results: Transcriptome microarray data of four GEO datasets were incorporated, including 142 primary Aquaporin tumor samples and 104 normal pancreatic tissue samples. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Distinct Genetic Alterations in Colorectal Cancer
Distinct Genetic Alterations in Colorectal Cancer Hassan Ashktorab1*, Alejandro A. Scha¨ffer2, Mohammad Daremipouran3, Duane T. Smoot3, Edward Lee3, Hassan Brim3 1 Department of Medicine and Cancer Center, Howard University, College of Medicine, Washington, DC, United States of America, 2 National Center for Biotechnology Information, National Institutes of Health (NIH), Department of Health and Human Services (DHHS), Bethesda, Maryland, United States of America, 3 Department of Pathology, Howard University, College of Medicine, Washington, DC, United States of America Abstract Background: Colon cancer (CRC) development often includes chromosomal instability (CIN) leading to amplifications and deletions of large DNA segments. Epidemiological, clinical, and cytogenetic studies showed that there are considerable differences between CRC tumors from African Americans (AAs) and Caucasian patients. In this study, we determined genomic copy number aberrations in sporadic CRC tumors from AAs, in order to investigate possible explanations for the observed disparities. Methodology/Principal Findings: We applied genome-wide array comparative genome hybridization (aCGH) using a 105k chip to identify copy number aberrations in samples from 15 AAs. In addition, we did a population comparative analysis with aCGH data in Caucasians as well as with a widely publicized list of colon cancer genes (CAN genes). There was an average of 20 aberrations per patient with more amplifications than deletions. Analysis of DNA copy number of frequently altered chromosomes revealed that deletions occurred primarily in chromosomes 4, 8 and 18. Chromosomal duplications occurred in more than 50% of cases on chromosomes 7, 8, 13, 20 and X. The CIN profile showed some differences when compared to Caucasian alterations. Conclusions/Significance: Chromosome X amplification in male patients and chromosomes 4, 8 and 18 deletions were prominent aberrations in AAs. -

A Compendium of Co-Regulated Protein Complexes in Breast Cancer Reveals Collateral Loss Events
bioRxiv preprint doi: https://doi.org/10.1101/155333; this version posted June 26, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. A compendium of co-regulated protein complexes in breast cancer reveals collateral loss events Colm J. Ryan*1, Susan Kennedy1, Ilirjana Bajrami2, David Matallanas1, Christopher J. Lord2 1Systems Biology Ireland, School of Medicine, University College Dublin, Dublin 4, Ireland 2The Breast Cancer Now Toby Robins Breast Cancer Research Centre and CRUK Gene Function Laboratory, The Institute of Cancer Research, London, SW3 6JB, United Kingdom. * Correspondence: [email protected] Summary Protein complexes are responsible for the bulk of activities within the cell, but how their behavior and composition varies across tumors remains poorly understood. By combining proteomic profiles of breast tumors with a large-scale protein-protein interaction network, we have identified a set of 258 high-confidence protein complexes whose subunits have highly correlated protein abundance across tumor samples. We used this set to identify complexes that are reproducibly under- or over- expressed in specific breast cancer subtypes. We found that mutation or deletion of one subunit of a complex was often associated with a collateral reduction in protein expression of additional complex members. This collateral loss phenomenon was evident from proteomic, but not transcriptomic, profiles suggesting post- transcriptional control. Mutation of the tumor suppressor E-cadherin (CDH1) was associated with a collateral loss of members of the adherens junction complex, an effect we validated using an engineered model of E-cadherin loss. -

Microtubule Regulation in Cystic Fibrosis Pathophysiology
MICROTUBULE REGULATION IN CYSTIC FIBROSIS PATHOPHYSIOLOGY By: SHARON MARIE RYMUT Submitted in partial fulfillment of the requirements For the degree of Doctor of Philosophy Dissertation Advisor: Dr. Thomas J Kelley Department of Pharmacology CASE WESTERN RESERVE UNIVERSITY August 2015 CASE WESTERN RESERVE UNIVERSITY SCHOOL OF GRADUATE STUDIES We hereby approve the thesis/ dissertation of Sharon Marie Rymut candidate for the Doctor of Philosophy degree* Dissertation Advisor: Thomas J Kelley Committee Chair: Paul N MacDonald Committee Member: Ruth E Siegel Committee Member: Craig A Hodges Committee Member: Danny Manor Committee Member: Rebecca J Darrah Date of Defense: April 29, 2015 * We also certify that written approval has been obtained for any proprietary material contained therein. ii Dedication There are five chapters in this dissertation. To Mom, Dad, Joe, Marie and Susan, I dedicate one chapter to each of you. You can fight about which chapter you want later. iii Table of Contents Table of Contents................................................................................................................iv List of Tables .................................................................................................................... vii List of Figures .................................................................................................................. viii Acknowledgements ............................................................................................................. x List of Abbreviations ....................................................................................................... -
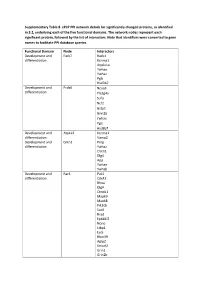
Supplementary Table 8. Cpcp PPI Network Details for Significantly Changed Proteins, As Identified in 3.2, Underlying Each of the Five Functional Domains
Supplementary Table 8. cPCP PPI network details for significantly changed proteins, as identified in 3.2, underlying each of the five functional domains. The network nodes represent each significant protein, followed by the list of interactors. Note that identifiers were converted to gene names to facilitate PPI database queries. Functional Domain Node Interactors Development and Park7 Rack1 differentiation Kcnma1 Atp6v1a Ywhae Ywhaz Pgls Hsd3b7 Development and Prdx6 Ncoa3 differentiation Pla2g4a Sufu Ncf2 Gstp1 Grin2b Ywhae Pgls Hsd3b7 Development and Atp1a2 Kcnma1 differentiation Vamp2 Development and Cntn1 Prnp differentiation Ywhaz Clstn1 Dlg4 App Ywhae Ywhab Development and Rac1 Pak1 differentiation Cdc42 Rhoa Dlg4 Ctnnb1 Mapk9 Mapk8 Pik3cb Sod1 Rrad Epb41l2 Nono Ltbp1 Evi5 Rbm39 Aplp2 Smurf2 Grin1 Grin2b Xiap Chn2 Cav1 Cybb Pgls Ywhae Development and Hbb-b1 Atp5b differentiation Hba Kcnma1 Got1 Aldoa Ywhaz Pgls Hsd3b4 Hsd3b7 Ywhae Development and Myh6 Mybpc3 differentiation Prkce Ywhae Development and Amph Capn2 differentiation Ap2a2 Dnm1 Dnm3 Dnm2 Atp6v1a Ywhab Development and Dnm3 Bin1 differentiation Amph Pacsin1 Grb2 Ywhae Bsn Development and Eef2 Ywhaz differentiation Rpgrip1l Atp6v1a Nphp1 Iqcb1 Ezh2 Ywhae Ywhab Pgls Hsd3b7 Hsd3b4 Development and Gnai1 Dlg4 differentiation Development and Gnao1 Dlg4 differentiation Vamp2 App Ywhae Ywhab Development and Psmd3 Rpgrip1l differentiation Psmd4 Hmga2 Development and Thy1 Syp differentiation Atp6v1a App Ywhae Ywhaz Ywhab Hsd3b7 Hsd3b4 Development and Tubb2a Ywhaz differentiation Nphp4 -

Epigenome-Wide Association Study on Diffusing Capacity of the Lung: Replication and Meta- Analysis
EWAS of pulmonary diffusion 1 http://hdl.handle.net/1765/112075 Epigenome-wide association study on diffusing capacity Epigenome-wideof the lung: replication association and study on diffusingmeta-analysis capacity 3.4 of the lung: replication and meta- analysis 2 Erasmus Medical Center Rotterdam ABSTRACT Background: Epigenetics may play an important role in pathogenesis of lung diseases. However, little is known about the epigenetic factors that influence impaired gas exchange at the lungs. Aim: To identify the epigenetic signatures of the diffusing capacity of the lung measured by carbon monoxide uptake. Methods: Epigenome association study (EWAS) was performed on diffusing capacity, measured by carbon monoxide uptake (DLCO) and per alveolar volume (DLCO/VA) using the single-breath technique in 2,674 individuals from two population-based cohort studies, the Rotterdam Study (the discovery panel) and the Framingham Heart Study (the replication panel). We assessed the clinical relevance of our findings by investigating the identified sites in whole blood and lung tissue specific gene expression. Finally, we performed an exploratory epigenome-wide analysis, combining both cohorts. Results: We identified and replicated two CpG sites (cg05575921 and cg05951221) that were significantly associated with DLCO/VA and one (cg05575921) suggestively associated with DLCO. Furthermore, we found a positive association between AHRR (cg05575921) hypomethylation and gene expression of EXOC3 in whole blood. We confirmed that the expression of EXOC3 is positively associated with DLCO/VA and DLCO in lung tissue. Finally, several genome-wide associations were identified after meta-analysing the data of both cohorts. Conclusions: We report on epigenome-wide associations with diffusing capacity in the general population. -

Bioinformatics Analysis for the Identification of Differentially Expressed Genes and Related Signaling Pathways in H
Bioinformatics analysis for the identification of differentially expressed genes and related signaling pathways in H. pylori-CagA transfected gastric cancer cells Dingyu Chen*, Chao Li, Yan Zhao, Jianjiang Zhou, Qinrong Wang and Yuan Xie* Key Laboratory of Endemic and Ethnic Diseases , Ministry of Education, Guizhou Medical University, Guiyang, China * These authors contributed equally to this work. ABSTRACT Aim. Helicobacter pylori cytotoxin-associated protein A (CagA) is an important vir- ulence factor known to induce gastric cancer development. However, the cause and the underlying molecular events of CagA induction remain unclear. Here, we applied integrated bioinformatics to identify the key genes involved in the process of CagA- induced gastric epithelial cell inflammation and can ceration to comprehend the potential molecular mechanisms involved. Materials and Methods. AGS cells were transected with pcDNA3.1 and pcDNA3.1::CagA for 24 h. The transfected cells were subjected to transcriptome sequencing to obtain the expressed genes. Differentially expressed genes (DEG) with adjusted P value < 0.05, | logFC |> 2 were screened, and the R package was applied for gene ontology (GO) enrichment and the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis. The differential gene protein–protein interaction (PPI) network was constructed using the STRING Cytoscape application, which conducted visual analysis to create the key function networks and identify the key genes. Next, the Submitted 20 August 2020 Kaplan–Meier plotter survival analysis tool was employed to analyze the survival of the Accepted 11 March 2021 key genes derived from the PPI network. Further analysis of the key gene expressions Published 15 April 2021 in gastric cancer and normal tissues were performed based on The Cancer Genome Corresponding author Atlas (TCGA) database and RT-qPCR verification. -

Centre for Arab Genomic Studies a Division of Sheikh Hamdan Award for Medical Sciences
Centre for Arab Genomic Studies A Division of Sheikh Hamdan Award for Medical Sciences The atalogue for ransmission enetics in rabs C T G A CTGA Database Exocyst Complex Component 4 Alternative Names defects. Individuals were diagnosed with MKS EXOC4 based on the presence of occipital encephalocele as SEC8, S. Cerevisiae, Homolog of well as any combination of liver fibrosis, cleft SEC8 palate, dysplastic kidneys, polydactyly and early KIAA1699 lethality. DNA from both affected and healthy members was obtained and an autozygome guided Record Category mutation analysis of known MKS genes was carried Gene locus out. However, some families did not have mutations in any of the known MKS genes. In such WHO-ICD cases, an exome sequencing was performed. N/A to gene loci Exomes were then searched for compound heterozygous mutations in known MKS genes. Incidence per 100,000 Live Births Failing that, all detected variants were filtered for N/A to gene loci homozygous novel changes within the autozygome. This resulted in the detection of a novel OMIM Number homozygous mutation c.1733A>G (p.Gln578Arg) 608185 in the EXOC4 gene in one of the affected families. This mutation was not found in dbSNP, 1000 Mode of Inheritance genomes or 200 Saudi controls. In-silico analysis N/A to gene loci by PolyPhen predicted this variant to be ‘probably damaging’ while SIFT predicted it to be Gene Map Locus deleterious. The authors noted that EXOC4 had not 7q33 previously been linked to MKS syndrome and that more studies were needed to confirm this Description association. The EXOC4 gene encodes a protein that forms the exocyst complex along with seven other EXOC References proteins. -

Exocyst Components Promote an Incompatible Interaction Between Glycine Max (Soybean) and Heterodera Glycines (The Soybean Cyst Nematode) Keshav Sharma1,7, Prakash M
www.nature.com/scientificreports OPEN Exocyst components promote an incompatible interaction between Glycine max (soybean) and Heterodera glycines (the soybean cyst nematode) Keshav Sharma1,7, Prakash M. Niraula1,8, Hallie A. Troell1, Mandeep Adhikari1, Hamdan Ali Alshehri2, Nadim W. Alkharouf3, Kathy S. Lawrence4 & Vincent P. Klink1,5,6* Vesicle and target membrane fusion involves tethering, docking and fusion. The GTPase SECRETORY4 (SEC4) positions the exocyst complex during vesicle membrane tethering, facilitating docking and fusion. Glycine max (soybean) Sec4 functions in the root during its defense against the parasitic nematode Heterodera glycines as it attempts to develop a multinucleate nurse cell (syncytium) serving to nourish the nematode over its 30-day life cycle. Results indicate that other tethering proteins are also important for defense. The G. max exocyst is encoded by 61 genes: 5 EXOC1 (Sec3), 2 EXOC2 (Sec5), 5 EXOC3 (Sec6), 2 EXOC4 (Sec8), 2 EXOC5 (Sec10) 6 EXOC6 (Sec15), 31 EXOC7 (Exo70) and 8 EXOC8 (Exo84) genes. At least one member of each gene family is expressed within the syncytium during the defense response. Syncytium-expressed exocyst genes function in defense while some are under transcriptional regulation by mitogen-activated protein kinases (MAPKs). The exocyst component EXOC7-H4-1 is not expressed within the syncytium but functions in defense and is under MAPK regulation. The tethering stage of vesicle transport has been demonstrated to play an important role in defense in the G. max-H. glycines pathosystem, with some of the spatially and temporally regulated exocyst components under transcriptional control by MAPKs. Abbreviations DCM Detection call methodology wr Whole root system pg Per gram SAR Systemic acquired resistance During their defense against pathogen infection, plants employ cellular processes to detect and amplify signals derived from the activities of those pathogens. -

Polymorphisms Near EXOC4 and LRGUK on Chromosome 7Q32 Are Associated with Type 2 Diabetes and Fasting Glucose; the NHLBI Family Heart Study Jason M
Washington University School of Medicine Digital Commons@Becker Open Access Publications 2008 Polymorphisms near EXOC4 and LRGUK on chromosome 7q32 are associated with Type 2 Diabetes and fasting glucose; The NHLBI Family Heart Study Jason M. Laramie Boston University Jemma B. Wilk Boston University Sally L. Williamson Boston University Michael W. Nagle Boston University Jeanne C. Latourelle Boston University See next page for additional authors Follow this and additional works at: https://digitalcommons.wustl.edu/open_access_pubs Part of the Medicine and Health Sciences Commons Recommended Citation Laramie, Jason M.; Wilk, Jemma B.; Williamson, Sally L.; Nagle, Michael W.; Latourelle, Jeanne C.; Tobin, Jennifer E.; Province, Michael A.; Borecki, Ingrid B.; and Myers, Richard H., ,"Polymorphisms near EXOC4 and LRGUK on chromosome 7q32 are associated with Type 2 Diabetes and fasting glucose; The NHLBI Family Heart Study." BMC Medical Genetics.,. 46. (2008). https://digitalcommons.wustl.edu/open_access_pubs/269 This Open Access Publication is brought to you for free and open access by Digital Commons@Becker. It has been accepted for inclusion in Open Access Publications by an authorized administrator of Digital Commons@Becker. For more information, please contact [email protected]. Authors Jason M. Laramie, Jemma B. Wilk, Sally L. Williamson, Michael W. Nagle, Jeanne C. Latourelle, Jennifer E. Tobin, Michael A. Province, Ingrid B. Borecki, and Richard H. Myers This open access publication is available at Digital Commons@Becker: https://digitalcommons.wustl.edu/open_access_pubs/269 -

A Deep Learning Approach for Predicting Antidepressant Response in Major Depression Using Clinical and Genetic Biomarkers
ORIGINAL RESEARCH published: 06 July 2018 doi: 10.3389/fpsyt.2018.00290 A Deep Learning Approach for Predicting Antidepressant Response in Major Depression Using Clinical and Genetic Biomarkers Eugene Lin 1,2*, Po-Hsiu Kuo 3, Yu-Li Liu 4, Younger W.-Y. Yu 5, Albert C. Yang 6,7,8,9 and Shih-Jen Tsai 6,7,9* 1 Department of Electrical Engineering, University of Washington, Seattle, WA, United States, 2 Graduate Institute of Biomedical Sciences, China Medical University, Taichung, Taiwan, 3 Department of Public Health, Institute of Epidemiology and Preventive Medicine, National Taiwan University, Taipei, Taiwan, 4 Center for Neuropsychiatric Research, National Health Research Institutes, Miaoli County, Taiwan, 5 Yu’s Psychiatric Clinic, Kaohsiung, Taiwan, 6 Department of Psychiatry, Taipei Veterans General Hospital, Taipei, Taiwan, 7 Division of Psychiatry, National Yang-Ming University, Taipei, Taiwan, 8 Division of Interdisciplinary Medicine and Biotechnology, Beth Israel Deaconess Medical Center/Harvard Medical School, Boston, MA, United States, 9 Institute of Brain Science, National Yang-Ming University, Taipei, Taiwan Edited by: In the wake of recent advances in scientific research, personalized medicine using Chad A. Bousman, University of Calgary, Canada deep learning techniques represents a new paradigm. In this work, our goal was to Reviewed by: establish deep learning models which distinguish responders from non-responders, Eva E. Redei, and also to predict possible antidepressant treatment outcomes in major depressive Northwestern