Mai Mult a Miniaturomutu Ti Won
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Microarray Analysis of Gene Expression in the Ovarian Cancer Cell Line HO-8910 with Silencing of the ZNF217 Gene
MOLECULAR MEDICINE REPORTS 2: 851-855, 2009 851 Microarray analysis of gene expression in the ovarian cancer cell line HO-8910 with silencing of the ZNF217 gene GUIQIN SUN1, JINGXIA QIN1, YUWEN QIU1, YUNFEI GAO1, YANHONG YU1, QINKAI DENG2 and MEI ZHONG1 1Department of Obstetrics and Gynecology, Nanfang Hospital, Southern Medical University; 2Department of Bioinformatics, Southern Medical University, Guangzhou 510515, Guangdong, P.R. China Received April 30, 2009; Accepted July 7, 2009 DOI: 10.3892/mmr_00000183 Abstract. Zinc-finger protein 217 (ZNF217), which is over- late stages (stage III or IV) and have a 5-year survival rate of expressed during cancer progression, can promote tumor cell less than 30% (1). Ovarian cancer is a heterogeneous disease immortalization. To examine the function of ZNF217, a global with respect to its histopathology, molecular biology and expression profile was carried out using Affymetrix Gene clinical outcome. Though studies on the etiology of ovarian Chip analysis with HG-U133 plus 2.0 arrays in the ovarian cancer and susceptibility to the disease have shown that the cancer cell line HO-8910 after silencing of the ZNF217 gene. mechanisms of carcinogenesis and cancer development are The results were analyzed using the Gene Ontology program associated with genetic, epigenetic and environmental factors, to investigate the functional network affected by ZNF217 the pathogenesis of ovarian cancer remains unclear (2). in ovarian cancer cells. Changes in the mRNA expression Among the current methods for the identification of gene of the affected genes were confirmed by real-time reverse expression profiles in cancer, microarray analysis is the most transcriptase-polymerase chain reaction (RT-PCR). -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

1 UST College of Science Department of Biological Sciences
UST College of Science Department of Biological Sciences 1 Pharmacogenomics of Myofascial Pain Syndrome An Undergraduate Thesis Submitted to the Department of Biological Sciences College of Science University of Santo Tomas In Partial Fulfillment of the Requirements for the Degree of Bachelor of Science in Biology Jose Marie V. Lazaga Marc Llandro C. Fernandez May 2021 UST College of Science Department of Biological Sciences 2 PANEL APPROVAL SHEET This undergraduate research manuscript entitled: Pharmacogenomics of Myofascial Pain Syndrome prepared and submitted by Jose Marie V. Lazaga and Marc Llandro C. Fernandez, was checked and has complied with the revisions and suggestions requested by panel members after thorough evaluation. This final version of the manuscript is hereby approved and accepted for submission in partial fulfillment of the requirements for the degree of Bachelor of Science in Biology. Noted by: Asst. Prof. Marilyn G. Rimando, PhD Research adviser, Bio/MicroSem 602-603 Approved by: Bio/MicroSem 603 panel member Bio/MicroSem 603 panel member Date: Date: UST College of Science Department of Biological Sciences 3 DECLARATION OF ORIGINALITY We hereby affirm that this submission is our own work and that, to the best of our knowledge and belief, it contains no material previously published or written by another person nor material to which a substantial extent has been accepted for award of any other degree or diploma of a university or other institute of higher learning, except where due acknowledgement is made in the text. We also declare that the intellectual content of this undergraduate research is the product of our work, even though we may have received assistance from others on style, presentation, and language expression. -

Distinct Genetic Alterations in Colorectal Cancer
Distinct Genetic Alterations in Colorectal Cancer Hassan Ashktorab1*, Alejandro A. Scha¨ffer2, Mohammad Daremipouran3, Duane T. Smoot3, Edward Lee3, Hassan Brim3 1 Department of Medicine and Cancer Center, Howard University, College of Medicine, Washington, DC, United States of America, 2 National Center for Biotechnology Information, National Institutes of Health (NIH), Department of Health and Human Services (DHHS), Bethesda, Maryland, United States of America, 3 Department of Pathology, Howard University, College of Medicine, Washington, DC, United States of America Abstract Background: Colon cancer (CRC) development often includes chromosomal instability (CIN) leading to amplifications and deletions of large DNA segments. Epidemiological, clinical, and cytogenetic studies showed that there are considerable differences between CRC tumors from African Americans (AAs) and Caucasian patients. In this study, we determined genomic copy number aberrations in sporadic CRC tumors from AAs, in order to investigate possible explanations for the observed disparities. Methodology/Principal Findings: We applied genome-wide array comparative genome hybridization (aCGH) using a 105k chip to identify copy number aberrations in samples from 15 AAs. In addition, we did a population comparative analysis with aCGH data in Caucasians as well as with a widely publicized list of colon cancer genes (CAN genes). There was an average of 20 aberrations per patient with more amplifications than deletions. Analysis of DNA copy number of frequently altered chromosomes revealed that deletions occurred primarily in chromosomes 4, 8 and 18. Chromosomal duplications occurred in more than 50% of cases on chromosomes 7, 8, 13, 20 and X. The CIN profile showed some differences when compared to Caucasian alterations. Conclusions/Significance: Chromosome X amplification in male patients and chromosomes 4, 8 and 18 deletions were prominent aberrations in AAs. -

Proteomic Analysis of HIV-1 Nef Cellular Binding Partners Reveals a Role for Exocyst Complex Proteins in Mediating Enhancement of Intercellular Nanotube Formation
Proteomic analysis of HIV-1 Nef cellular binding partners reveals a role for exocyst complex proteins in mediating enhancement of intercellular nanotube formation The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Mukerji, Joya, Kevin C Olivieri, Vikas Misra, Kristin A Agopian, and Dana Gabuzda. 2012. Proteomic analysis of hiv-1 nef cellular binding partners reveals a role for exocyst complex proteins in mediating enhancement of intercellular nanotube formation. Retrovirology 9: 33. Published Version doi:10.1186/1742-4690-9-33 Citable link http://nrs.harvard.edu/urn-3:HUL.InstRepos:10445557 Terms of Use This article was downloaded from Harvard University’s DASH repository, and is made available under the terms and conditions applicable to Other Posted Material, as set forth at http:// nrs.harvard.edu/urn-3:HUL.InstRepos:dash.current.terms-of- use#LAA Mukerji et al. Retrovirology 2012, 9:33 http://www.retrovirology.com/content/9/1/33 RESEARCH Open Access Proteomic analysis of HIV-1 Nef cellular binding partners reveals a role for exocyst complex proteins in mediating enhancement of intercellular nanotube formation Joya Mukerji1,2, Kevin C Olivieri1, Vikas Misra1, Kristin A Agopian1,2 and Dana Gabuzda1,2,3* Abstract Background: HIV-1 Nef protein contributes to pathogenesis via multiple functions that include enhancement of viral replication and infectivity, alteration of intracellular trafficking, and modulation of cellular signaling pathways. Nef stimulates formation of tunneling nanotubes and virological synapses, and is transferred to bystander cells via these intercellular contacts and secreted microvesicles. Nef associates with and activates Pak2, a kinase that regulates T-cell signaling and actin cytoskeleton dynamics, but how Nef promotes nanotube formation is unknown. -

Probabilistic Models for Collecting, Analyzing, and Modeling Expression Data
Probabilistic Models for Collecting, Analyzing, and Modeling Expression Data Hai-Son Phuoc Le May 2013 CMU-ML-13-101 Probabilistic Models for Collecting, Analyzing, and Modeling Expression Data Hai-Son Phuoc Le May 2013 CMU-ML-13-101 Machine Learning Department School of Computer Science Carnegie Mellon University Thesis Committee Ziv Bar-Joseph, Chair Christopher Langmead Roni Rosenfeld Quaid Morris Submitted in partial fulfillment of the requirements for the Degree of Doctor of Philosophy. Copyright @ 2013 Hai-Son Le This research was sponsored by the National Institutes of Health under grant numbers 5U01HL108642 and 1R01GM085022, the National Science Foundation under grant num- bers DBI0448453 and DBI0965316, and the Pittsburgh Life Sciences Greenhouse. The views and conclusions contained in this document are those of the author and should not be interpreted as representing the official policies, either expressed or implied, of any sponsoring institution, the U.S. government or any other entity. Keywords: genomics, gene expression, gene regulation, microarray, RNA-Seq, transcriptomics, error correction, comparative genomics, regulatory networks, cross-species, expression database, Gene Expression Omnibus, GEO, orthologs, microRNA, target prediction, Dirichlet Process, Indian Buffet Process, hidden Markov model, immune response, cancer. To Mom and Dad. i Abstract Advances in genomics allow researchers to measure the complete set of transcripts in cells. These transcripts include messenger RNAs (which encode for proteins) and microRNAs, short RNAs that play an important regulatory role in cellular networks. While this data is a great resource for reconstructing the activity of networks in cells, it also presents several computational challenges. These challenges include the data collection stage which often results in incomplete and noisy measurement, developing methods to integrate several experiments within and across species, and designing methods that can use this data to map the interactions and networks that are activated in specific conditions. -

Identification of Potential Key Genes and Pathway Linked with Sporadic Creutzfeldt-Jakob Disease Based on Integrated Bioinformatics Analyses
medRxiv preprint doi: https://doi.org/10.1101/2020.12.21.20248688; this version posted December 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. All rights reserved. No reuse allowed without permission. Identification of potential key genes and pathway linked with sporadic Creutzfeldt-Jakob disease based on integrated bioinformatics analyses Basavaraj Vastrad1, Chanabasayya Vastrad*2 , Iranna Kotturshetti 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karanataka, India. 3. Department of Ayurveda, Rajiv Gandhi Education Society`s Ayurvedic Medical College, Ron, Karnataka 562209, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India NOTE: This preprint reports new research that has not been certified by peer review and should not be used to guide clinical practice. medRxiv preprint doi: https://doi.org/10.1101/2020.12.21.20248688; this version posted December 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. All rights reserved. No reuse allowed without permission. Abstract Sporadic Creutzfeldt-Jakob disease (sCJD) is neurodegenerative disease also called prion disease linked with poor prognosis. The aim of the current study was to illuminate the underlying molecular mechanisms of sCJD. The mRNA microarray dataset GSE124571 was downloaded from the Gene Expression Omnibus database. Differentially expressed genes (DEGs) were screened. -
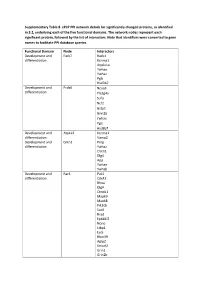
Supplementary Table 8. Cpcp PPI Network Details for Significantly Changed Proteins, As Identified in 3.2, Underlying Each of the Five Functional Domains
Supplementary Table 8. cPCP PPI network details for significantly changed proteins, as identified in 3.2, underlying each of the five functional domains. The network nodes represent each significant protein, followed by the list of interactors. Note that identifiers were converted to gene names to facilitate PPI database queries. Functional Domain Node Interactors Development and Park7 Rack1 differentiation Kcnma1 Atp6v1a Ywhae Ywhaz Pgls Hsd3b7 Development and Prdx6 Ncoa3 differentiation Pla2g4a Sufu Ncf2 Gstp1 Grin2b Ywhae Pgls Hsd3b7 Development and Atp1a2 Kcnma1 differentiation Vamp2 Development and Cntn1 Prnp differentiation Ywhaz Clstn1 Dlg4 App Ywhae Ywhab Development and Rac1 Pak1 differentiation Cdc42 Rhoa Dlg4 Ctnnb1 Mapk9 Mapk8 Pik3cb Sod1 Rrad Epb41l2 Nono Ltbp1 Evi5 Rbm39 Aplp2 Smurf2 Grin1 Grin2b Xiap Chn2 Cav1 Cybb Pgls Ywhae Development and Hbb-b1 Atp5b differentiation Hba Kcnma1 Got1 Aldoa Ywhaz Pgls Hsd3b4 Hsd3b7 Ywhae Development and Myh6 Mybpc3 differentiation Prkce Ywhae Development and Amph Capn2 differentiation Ap2a2 Dnm1 Dnm3 Dnm2 Atp6v1a Ywhab Development and Dnm3 Bin1 differentiation Amph Pacsin1 Grb2 Ywhae Bsn Development and Eef2 Ywhaz differentiation Rpgrip1l Atp6v1a Nphp1 Iqcb1 Ezh2 Ywhae Ywhab Pgls Hsd3b7 Hsd3b4 Development and Gnai1 Dlg4 differentiation Development and Gnao1 Dlg4 differentiation Vamp2 App Ywhae Ywhab Development and Psmd3 Rpgrip1l differentiation Psmd4 Hmga2 Development and Thy1 Syp differentiation Atp6v1a App Ywhae Ywhaz Ywhab Hsd3b7 Hsd3b4 Development and Tubb2a Ywhaz differentiation Nphp4 -

Supplementary Table 1
Supplementary Table 1. 492 genes are unique to 0 h post-heat timepoint. The name, p-value, fold change, location and family of each gene are indicated. Genes were filtered for an absolute value log2 ration 1.5 and a significance value of p ≤ 0.05. Symbol p-value Log Gene Name Location Family Ratio ABCA13 1.87E-02 3.292 ATP-binding cassette, sub-family unknown transporter A (ABC1), member 13 ABCB1 1.93E-02 −1.819 ATP-binding cassette, sub-family Plasma transporter B (MDR/TAP), member 1 Membrane ABCC3 2.83E-02 2.016 ATP-binding cassette, sub-family Plasma transporter C (CFTR/MRP), member 3 Membrane ABHD6 7.79E-03 −2.717 abhydrolase domain containing 6 Cytoplasm enzyme ACAT1 4.10E-02 3.009 acetyl-CoA acetyltransferase 1 Cytoplasm enzyme ACBD4 2.66E-03 1.722 acyl-CoA binding domain unknown other containing 4 ACSL5 1.86E-02 −2.876 acyl-CoA synthetase long-chain Cytoplasm enzyme family member 5 ADAM23 3.33E-02 −3.008 ADAM metallopeptidase domain Plasma peptidase 23 Membrane ADAM29 5.58E-03 3.463 ADAM metallopeptidase domain Plasma peptidase 29 Membrane ADAMTS17 2.67E-04 3.051 ADAM metallopeptidase with Extracellular other thrombospondin type 1 motif, 17 Space ADCYAP1R1 1.20E-02 1.848 adenylate cyclase activating Plasma G-protein polypeptide 1 (pituitary) receptor Membrane coupled type I receptor ADH6 (includes 4.02E-02 −1.845 alcohol dehydrogenase 6 (class Cytoplasm enzyme EG:130) V) AHSA2 1.54E-04 −1.6 AHA1, activator of heat shock unknown other 90kDa protein ATPase homolog 2 (yeast) AK5 3.32E-02 1.658 adenylate kinase 5 Cytoplasm kinase AK7 -

Centre for Arab Genomic Studies a Division of Sheikh Hamdan Award for Medical Sciences
Centre for Arab Genomic Studies A Division of Sheikh Hamdan Award for Medical Sciences The atalogue for ransmission enetics in rabs C T G A CTGA Database Exocyst Complex Component 4 Alternative Names defects. Individuals were diagnosed with MKS EXOC4 based on the presence of occipital encephalocele as SEC8, S. Cerevisiae, Homolog of well as any combination of liver fibrosis, cleft SEC8 palate, dysplastic kidneys, polydactyly and early KIAA1699 lethality. DNA from both affected and healthy members was obtained and an autozygome guided Record Category mutation analysis of known MKS genes was carried Gene locus out. However, some families did not have mutations in any of the known MKS genes. In such WHO-ICD cases, an exome sequencing was performed. N/A to gene loci Exomes were then searched for compound heterozygous mutations in known MKS genes. Incidence per 100,000 Live Births Failing that, all detected variants were filtered for N/A to gene loci homozygous novel changes within the autozygome. This resulted in the detection of a novel OMIM Number homozygous mutation c.1733A>G (p.Gln578Arg) 608185 in the EXOC4 gene in one of the affected families. This mutation was not found in dbSNP, 1000 Mode of Inheritance genomes or 200 Saudi controls. In-silico analysis N/A to gene loci by PolyPhen predicted this variant to be ‘probably damaging’ while SIFT predicted it to be Gene Map Locus deleterious. The authors noted that EXOC4 had not 7q33 previously been linked to MKS syndrome and that more studies were needed to confirm this Description association. The EXOC4 gene encodes a protein that forms the exocyst complex along with seven other EXOC References proteins. -

Exocyst Components Promote an Incompatible Interaction Between Glycine Max (Soybean) and Heterodera Glycines (The Soybean Cyst Nematode) Keshav Sharma1,7, Prakash M
www.nature.com/scientificreports OPEN Exocyst components promote an incompatible interaction between Glycine max (soybean) and Heterodera glycines (the soybean cyst nematode) Keshav Sharma1,7, Prakash M. Niraula1,8, Hallie A. Troell1, Mandeep Adhikari1, Hamdan Ali Alshehri2, Nadim W. Alkharouf3, Kathy S. Lawrence4 & Vincent P. Klink1,5,6* Vesicle and target membrane fusion involves tethering, docking and fusion. The GTPase SECRETORY4 (SEC4) positions the exocyst complex during vesicle membrane tethering, facilitating docking and fusion. Glycine max (soybean) Sec4 functions in the root during its defense against the parasitic nematode Heterodera glycines as it attempts to develop a multinucleate nurse cell (syncytium) serving to nourish the nematode over its 30-day life cycle. Results indicate that other tethering proteins are also important for defense. The G. max exocyst is encoded by 61 genes: 5 EXOC1 (Sec3), 2 EXOC2 (Sec5), 5 EXOC3 (Sec6), 2 EXOC4 (Sec8), 2 EXOC5 (Sec10) 6 EXOC6 (Sec15), 31 EXOC7 (Exo70) and 8 EXOC8 (Exo84) genes. At least one member of each gene family is expressed within the syncytium during the defense response. Syncytium-expressed exocyst genes function in defense while some are under transcriptional regulation by mitogen-activated protein kinases (MAPKs). The exocyst component EXOC7-H4-1 is not expressed within the syncytium but functions in defense and is under MAPK regulation. The tethering stage of vesicle transport has been demonstrated to play an important role in defense in the G. max-H. glycines pathosystem, with some of the spatially and temporally regulated exocyst components under transcriptional control by MAPKs. Abbreviations DCM Detection call methodology wr Whole root system pg Per gram SAR Systemic acquired resistance During their defense against pathogen infection, plants employ cellular processes to detect and amplify signals derived from the activities of those pathogens. -

Polymorphisms Near EXOC4 and LRGUK on Chromosome 7Q32 Are Associated with Type 2 Diabetes and Fasting Glucose; the NHLBI Family Heart Study Jason M
Washington University School of Medicine Digital Commons@Becker Open Access Publications 2008 Polymorphisms near EXOC4 and LRGUK on chromosome 7q32 are associated with Type 2 Diabetes and fasting glucose; The NHLBI Family Heart Study Jason M. Laramie Boston University Jemma B. Wilk Boston University Sally L. Williamson Boston University Michael W. Nagle Boston University Jeanne C. Latourelle Boston University See next page for additional authors Follow this and additional works at: https://digitalcommons.wustl.edu/open_access_pubs Part of the Medicine and Health Sciences Commons Recommended Citation Laramie, Jason M.; Wilk, Jemma B.; Williamson, Sally L.; Nagle, Michael W.; Latourelle, Jeanne C.; Tobin, Jennifer E.; Province, Michael A.; Borecki, Ingrid B.; and Myers, Richard H., ,"Polymorphisms near EXOC4 and LRGUK on chromosome 7q32 are associated with Type 2 Diabetes and fasting glucose; The NHLBI Family Heart Study." BMC Medical Genetics.,. 46. (2008). https://digitalcommons.wustl.edu/open_access_pubs/269 This Open Access Publication is brought to you for free and open access by Digital Commons@Becker. It has been accepted for inclusion in Open Access Publications by an authorized administrator of Digital Commons@Becker. For more information, please contact [email protected]. Authors Jason M. Laramie, Jemma B. Wilk, Sally L. Williamson, Michael W. Nagle, Jeanne C. Latourelle, Jennifer E. Tobin, Michael A. Province, Ingrid B. Borecki, and Richard H. Myers This open access publication is available at Digital Commons@Becker: https://digitalcommons.wustl.edu/open_access_pubs/269