Pirb Inhibits Axonal Outgrowth Via the PI3K/Akt/Mtor Signaling Pathway
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Reticulons: a Family of Proteins with Diverse Functions Yvonne S Yang and Stephen M Strittmatter
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by PubMed Central Protein family review The reticulons: a family of proteins with diverse functions Yvonne S Yang and Stephen M Strittmatter Address: Program in Cellular Neuroscience, Neurodegeneration and Repair, Department of Neurology, Yale University School of Medicine, New Haven, CT 06536, USA. Correspondence: Stephen M Strittmatter. Email: [email protected] Published: 28 December 2007 Genome Biology 2007, 8:234 (doi:10.1186/gb-2007-8-12-234) The electronic version of this article is the complete one and can be found online at http://genomebiology.com/2007/8/12/234 © 2007 BioMed Central Ltd Summary The reticulon family is a large and diverse group of membrane-associated proteins found throughout the eukaryotic kingdom. All of its members contain a carboxy-terminal reticulon homology domain that consists of two hydrophobic regions flanking a hydrophilic loop of 60-70 amino acids, but reticulon amino-terminal domains display little or no similarity to each other. Reticulons principally localize to the endoplasmic reticulum, and there is evidence that they influence endoplasmic reticulum-Golgi trafficking, vesicle formation and membrane morphogenesis. However, mammalian reticulons have also been found on the cell surface and mammalian reticulon 4 expressed on the surface of oligodendrocytes is an inhibitor of axon growth both in culture and in vivo. There is also growing evidence that reticulons may be important in neurodegenerative diseases such as Alzheimer’s disease and amyotrophic lateral sclerosis. The diversity of structure, topology, localization and expression patterns of reticulons is reflected in their multiple, diverse functions in the cell. -

Functions of Nogo Proteins and Their Receptors in the Nervous System
REVIEWS Functions of Nogo proteins and their receptors in the nervous system Martin E. Schwab Abstract | The membrane protein Nogo-A was initially characterized as a CNS-specific inhibitor of axonal regeneration. Recent studies have uncovered regulatory roles of Nogo proteins and their receptors — in precursor migration, neurite growth and branching in the developing nervous system — as well as a growth-restricting function during CNS maturation. The function of Nogo in the adult CNS is now understood to be that of a negative regulator of neuronal growth, leading to stabilization of the CNS wiring at the expense of extensive plastic rearrangements and regeneration after injury. In addition, Nogo proteins interact with various intracellular components and may have roles in the regulation of endoplasmic reticulum (ER) structure, processing of amyloid precursor protein and cell survival. Nogo proteins were discovered, and have been exten- The amino-terminal segments of the proteins sively studied, in the context of injury and repair of fibre encoded by the different RTN genes have differing tracts in the CNS1 — a topic of great research interest lengths and there is no homology between them3,5. The and clinical relevance. However, much less in known N termini of the RTN4 products Nogo-A and Nogo-B are about the physiological functions of Nogo proteins in identical, consisting of a 172-amino acid sequence that development and in the intact adult organism, including is encoded by a single exon (exon 1) that is followed by in the brain. Through a number of recent publications, a short exon 2 and, in Nogo-A, by the very long exon Nogo proteins have emerged as important regulators of 3 encoding 800 amino acids2,6 (FIG. -

Application of Microrna Database Mining in Biomarker Discovery and Identification of Therapeutic Targets for Complex Disease
Article Application of microRNA Database Mining in Biomarker Discovery and Identification of Therapeutic Targets for Complex Disease Jennifer L. Major, Rushita A. Bagchi * and Julie Pires da Silva * Department of Medicine, Division of Cardiology, University of Colorado Anschutz Medical Campus, Aurora, CO 80045, USA; [email protected] * Correspondence: [email protected] (R.A.B.); [email protected] (J.P.d.S.) Supplementary Tables Methods Protoc. 2021, 4, 5. https://doi.org/10.3390/mps4010005 www.mdpi.com/journal/mps Methods Protoc. 2021, 4, 5. https://doi.org/10.3390/mps4010005 2 of 25 Table 1. List of all hsa-miRs identified by Human microRNA Disease Database (HMDD; v3.2) analysis. hsa-miRs were identified using the term “genetics” and “circulating” as input in HMDD. Targets CAD hsa-miR-1 Targets IR injury hsa-miR-423 Targets Obesity hsa-miR-499 hsa-miR-146a Circulating Obesity Genetics CAD hsa-miR-423 hsa-miR-146a Circulating CAD hsa-miR-149 hsa-miR-499 Circulating IR Injury hsa-miR-146a Circulating Obesity hsa-miR-122 Genetics Stroke Circulating CAD hsa-miR-122 Circulating Stroke hsa-miR-122 Genetics Obesity Circulating Stroke hsa-miR-26b hsa-miR-17 hsa-miR-223 Targets CAD hsa-miR-340 hsa-miR-34a hsa-miR-92a hsa-miR-126 Circulating Obesity Targets IR injury hsa-miR-21 hsa-miR-423 hsa-miR-126 hsa-miR-143 Targets Obesity hsa-miR-21 hsa-miR-223 hsa-miR-34a hsa-miR-17 Targets CAD hsa-miR-223 hsa-miR-92a hsa-miR-126 Targets IR injury hsa-miR-155 hsa-miR-21 Circulating CAD hsa-miR-126 hsa-miR-145 hsa-miR-21 Targets Obesity hsa-mir-223 hsa-mir-499 hsa-mir-574 Targets IR injury hsa-mir-21 Circulating IR injury Targets Obesity hsa-mir-21 Targets CAD hsa-mir-22 hsa-mir-133a Targets IR injury hsa-mir-155 hsa-mir-21 Circulating Stroke hsa-mir-145 hsa-mir-146b Targets Obesity hsa-mir-21 hsa-mir-29b Methods Protoc. -

Cell Biology of Spinal Cord Injury and Repair
The Journal of Clinical Investigation REVIEW SERIES: GLIA AND NEURODEGENERATION Series Editors: Marco Colonna and David Holtzmann Cell biology of spinal cord injury and repair Timothy M. O’Shea, Joshua E. Burda, and Michael V. Sofroniew Department of Neurobiology, David Geffen School of Medicine, UCLA, Los Angeles, California, USA. Spinal cord injury (SCI) lesions present diverse challenges for repair strategies. Anatomically complete injuries require restoration of neural connectivity across lesions. Anatomically incomplete injuries may benefit from augmentation of spontaneous circuit reorganization. Here, we review SCI cell biology, which varies considerably across three different lesion- related tissue compartments: (a) non-neural lesion core, (b) astrocyte scar border, and (c) surrounding spared but reactive neural tissue. After SCI, axon growth and circuit reorganization are determined by neuron-cell-autonomous mechanisms and by interactions among neurons, glia, and immune and other cells. These interactions are shaped by both the presence and the absence of growth-modulating molecules, which vary markedly in different lesion compartments. The emerging understanding of how SCI cell biology differs across lesion compartments is fundamental to developing rationally targeted repair strategies. Introduction a central non-neural lesion core, often referred to as fibrotic scar Spinal cord injury (SCI) is a major cause of long-term physical (also mesenchymal or connective tissue scar); (b) an astroglial impairment. Current treatments are limited mostly to supportive scar border that intimately surrounds the lesion core; and (c) a sur- measures. Affected individuals often have life expectancies of rounding zone of viable neural tissue that is spared and function- decades with permanent disability (1–3). -

Strand Breaks for P53 Exon 6 and 8 Among Different Time Course of Folate Depletion Or Repletion in the Rectosigmoid Mucosa
SUPPLEMENTAL FIGURE COLON p53 EXONIC STRAND BREAKS DURING FOLATE DEPLETION-REPLETION INTERVENTION Supplemental Figure Legend Strand breaks for p53 exon 6 and 8 among different time course of folate depletion or repletion in the rectosigmoid mucosa. The input of DNA was controlled by GAPDH. The data is shown as ΔCt after normalized to GAPDH. The higher ΔCt the more strand breaks. The P value is shown in the figure. SUPPLEMENT S1 Genes that were significantly UPREGULATED after folate intervention (by unadjusted paired t-test), list is sorted by P value Gene Symbol Nucleotide P VALUE Description OLFM4 NM_006418 0.0000 Homo sapiens differentially expressed in hematopoietic lineages (GW112) mRNA. FMR1NB NM_152578 0.0000 Homo sapiens hypothetical protein FLJ25736 (FLJ25736) mRNA. IFI6 NM_002038 0.0001 Homo sapiens interferon alpha-inducible protein (clone IFI-6-16) (G1P3) transcript variant 1 mRNA. Homo sapiens UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 15 GALNTL5 NM_145292 0.0001 (GALNT15) mRNA. STIM2 NM_020860 0.0001 Homo sapiens stromal interaction molecule 2 (STIM2) mRNA. ZNF645 NM_152577 0.0002 Homo sapiens hypothetical protein FLJ25735 (FLJ25735) mRNA. ATP12A NM_001676 0.0002 Homo sapiens ATPase H+/K+ transporting nongastric alpha polypeptide (ATP12A) mRNA. U1SNRNPBP NM_007020 0.0003 Homo sapiens U1-snRNP binding protein homolog (U1SNRNPBP) transcript variant 1 mRNA. RNF125 NM_017831 0.0004 Homo sapiens ring finger protein 125 (RNF125) mRNA. FMNL1 NM_005892 0.0004 Homo sapiens formin-like (FMNL) mRNA. ISG15 NM_005101 0.0005 Homo sapiens interferon alpha-inducible protein (clone IFI-15K) (G1P2) mRNA. SLC6A14 NM_007231 0.0005 Homo sapiens solute carrier family 6 (neurotransmitter transporter) member 14 (SLC6A14) mRNA. -

A Single Legionella Effector Catalyzes a Multistep Ubiquitination Pathway to Rearrange Tubular Endoplasmic Reticulum for Replication
Article A Single Legionella Effector Catalyzes a Multistep Ubiquitination Pathway to Rearrange Tubular Endoplasmic Reticulum for Replication Graphical Abstract Authors Kristin M. Kotewicz, Vinay Ramabhadran, Nicole Sjoblom, ..., Jessica Behringer, Rebecca A. Scheck, Ralph R. Isberg Correspondence [email protected] In Brief Intracellular pathogens, including Legionella, target host organelles for replication. Kotewicz et al. show that Legionella generates an ER- encompassed replication compartment via Sde protein-mediated ubiquitination of host reticulon 4. Ubiquitination is mediated by sequential action of the ADP-ribosyltransferase and nucleotidase activities of Sde and independent of the host ubiquitination system. Highlights d Legionella Sde proteins remodel tubular ER to initiate intracellular replication d The tubular ER protein reticulon 4 is targeted by Sde proteins d ER remodeling requires ubiquitin transfer by Sde proteins d Ubiquitin transfer involves ADP-ribosyltransferase and nucleotidase activities Kotewicz et al., 2017, Cell Host & Microbe 21, 169–181 February 8, 2017 ª 2016 Elsevier Inc. http://dx.doi.org/10.1016/j.chom.2016.12.007 Cell Host & Microbe Article ASingleLegionella Effector Catalyzes a Multistep Ubiquitination Pathway to Rearrange Tubular Endoplasmic Reticulum for Replication Kristin M. Kotewicz,1,6 Vinay Ramabhadran,1,3,6 Nicole Sjoblom,2 Joseph P. Vogel,4 Eva Haenssler,1,7 Mengyun Zhang,1 Jessica Behringer,5 Rebecca A. Scheck,2 and Ralph R. Isberg1,3,8,* 1Department of Molecular Biology and Microbiology, Tufts University School of Medicine, 150 Harrison Ave., Boston, MA 02111, USA 2Department of Chemistry, Tufts University, 62 Talbot Ave., Medford, MA 02155, USA 3Howard Hughes Medical Institute, Tufts University School of Medicine, 150 Harrison Ave., Boston, MA 02111, USA 4Department of Molecular Microbiology, Washington University School of Medicine, St. -

Transdifferentiation of Human Mesenchymal Stem Cells
Transdifferentiation of Human Mesenchymal Stem Cells Dissertation zur Erlangung des naturwissenschaftlichen Doktorgrades der Julius-Maximilians-Universität Würzburg vorgelegt von Tatjana Schilling aus San Miguel de Tucuman, Argentinien Würzburg, 2007 Eingereicht am: Mitglieder der Promotionskommission: Vorsitzender: Prof. Dr. Martin J. Müller Gutachter: PD Dr. Norbert Schütze Gutachter: Prof. Dr. Georg Krohne Tag des Promotionskolloquiums: Doktorurkunde ausgehändigt am: Hiermit erkläre ich ehrenwörtlich, dass ich die vorliegende Dissertation selbstständig angefertigt und keine anderen als die von mir angegebenen Hilfsmittel und Quellen verwendet habe. Des Weiteren erkläre ich, dass diese Arbeit weder in gleicher noch in ähnlicher Form in einem Prüfungsverfahren vorgelegen hat und ich noch keinen Promotionsversuch unternommen habe. Gerbrunn, 4. Mai 2007 Tatjana Schilling Table of contents i Table of contents 1 Summary ........................................................................................................................ 1 1.1 Summary.................................................................................................................... 1 1.2 Zusammenfassung..................................................................................................... 2 2 Introduction.................................................................................................................... 4 2.1 Osteoporosis and the fatty degeneration of the bone marrow..................................... 4 2.2 Adipose and bone -
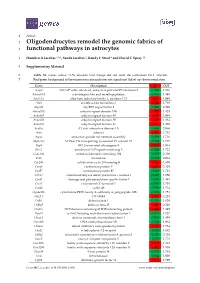
Oligodendrocytes Remodel the Genomic Fabrics of Functional Pathways in Astrocytes
1 Article 2 Oligodendrocytes remodel the genomic fabrics of 3 functional pathways in astrocytes 4 Dumitru A Iacobas 1,2,*, Sanda Iacobas 3, Randy F Stout 4 and David C Spray 2,5 5 Supplementary Material 6 Table S1. Genes whose >1.5x absolute fold-change did not meet the individual CUT criterion. 7 Red/green background of the expression ratio indicates not significant (false) up-/down-regulation. Gene Description X CUT Acap2 ArfGAP with coiled-coil, ankyrin repeat and PH domains 2 -1.540 1.816 Adamts18 a disintegrin-like and metallopeptidase -1.514 1.594 Akr1c12 aldo-keto reductase family 1, member C12 1.866 1.994 Alx3 aristaless-like homeobox 3 1.536 1.769 Alyref2 Aly/REF export factor 2 -1.880 2.208 Ankrd33b ankyrin repeat domain 33B 1.593 1.829 Ankrd45 ankyrin repeat domain 45 1.514 1.984 Ankrd50 ankyrin repeat domain 50 1.628 1.832 Ankrd61 ankyrin repeat domain 61 1.645 1.802 Arid1a AT rich interactive domain 1A -1.668 2.066 Artn artemin 1.524 1.732 Aspm abnormal spindle microtubule assembly -1.693 1.716 Atp6v1e1 ATPase, H+ transporting, lysosomal V1 subunit E1 -1.679 1.777 Bag4 BCL2-associated athanogene 4 1.723 1.914 Birc3 baculoviral IAP repeat-containing 3 -1.588 1.722 Ccdc104 coiled-coil domain containing 104 -1.819 2.130 Ccl2 chemokine -1.699 2.034 Cdc20b cell division cycle 20 homolog B 1.512 1.605 Cenpf centromere protein F 2.041 2.128 Cep97 centrosomal protein 97 -1.641 1.723 COX1 mitochondrially encoded cytochrome c oxidase I -1.607 1.650 Cpsf7 cleavage and polyadenylation specific factor 7 -1.635 1.891 Crct1 cysteine-rich -

Glial Cell Dysfunction in C9orf72-Related Amyotrophic Lateral Sclerosis and Frontotemporal Dementia
cells Review Glial Cell Dysfunction in C9orf72-Related Amyotrophic Lateral Sclerosis and Frontotemporal Dementia Mehdi Ghasemi * , Kiandokht Keyhanian and Catherine Douthwright Department of Neurology, University of Massachusetts Medical School, Worcester, MA 01655, USA; [email protected] (K.K.); [email protected] (C.D.) * Correspondence: [email protected]; Tel.: +1-774-441-7726; Fax: +1-508-856-4485 Abstract: Since the discovery of the chromosome 9 open reading frame 72 (C9orf72) repeat expansion mutation in 2011 as the most common genetic abnormality in amyotrophic lateral sclerosis (ALS, also known as Lou Gehrig’s disease) and frontotemporal dementia (FTD), progress in understanding the signaling pathways related to this mutation can only be described as intriguing. Two major theories have been suggested—(i) loss of function or haploinsufficiency and (ii) toxic gain of function from either C9orf72 repeat RNA or dipeptide repeat proteins (DPRs) generated from repeat-associated non-ATG (RAN) translation. Each theory has provided various signaling pathways that potentially participate in the disease progression. Dysregulation of the immune system, particularly glial cell dysfunction (mainly microglia and astrocytes), is demonstrated to play a pivotal role in both loss and gain of function theories of C9orf72 pathogenesis. In this review, we discuss the pathogenic roles of glial cells in C9orf72 ALS/FTD as evidenced by pre-clinical and clinical studies showing the presence of gliosis in C9orf72 ALS/FTD, pathologic hallmarks in glial cells, including TAR DNA-binding protein 43 (TDP-43) and p62 aggregates, and toxicity of C9orf72 glial cells. A better understanding of these pathways can provide new insights into the development of therapies targeting glial cell Citation: Ghasemi, M.; Keyhanian, abnormalities in C9orf72 ALS/FTD. -

Research Article Complex and Multidimensional Lipid Raft Alterations in a Murine Model of Alzheimer’S Disease
SAGE-Hindawi Access to Research International Journal of Alzheimer’s Disease Volume 2010, Article ID 604792, 56 pages doi:10.4061/2010/604792 Research Article Complex and Multidimensional Lipid Raft Alterations in a Murine Model of Alzheimer’s Disease Wayne Chadwick, 1 Randall Brenneman,1, 2 Bronwen Martin,3 and Stuart Maudsley1 1 Receptor Pharmacology Unit, National Institute on Aging, National Institutes of Health, 251 Bayview Boulevard, Suite 100, Baltimore, MD 21224, USA 2 Miller School of Medicine, University of Miami, Miami, FL 33124, USA 3 Metabolism Unit, National Institute on Aging, National Institutes of Health, 251 Bayview Boulevard, Suite 100, Baltimore, MD 21224, USA Correspondence should be addressed to Stuart Maudsley, [email protected] Received 17 May 2010; Accepted 27 July 2010 Academic Editor: Gemma Casadesus Copyright © 2010 Wayne Chadwick et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Various animal models of Alzheimer’s disease (AD) have been created to assist our appreciation of AD pathophysiology, as well as aid development of novel therapeutic strategies. Despite the discovery of mutated proteins that predict the development of AD, there are likely to be many other proteins also involved in this disorder. Complex physiological processes are mediated by coherent interactions of clusters of functionally related proteins. Synaptic dysfunction is one of the hallmarks of AD. Synaptic proteins are organized into multiprotein complexes in high-density membrane structures, known as lipid rafts. These microdomains enable coherent clustering of synergistic signaling proteins. -

Constitutively Active Calcineurin Induces Cardiac Endoplasmic Reticulum Stress and Protects Against Apoptosis That Is Mediated by Α-Crystallin-B
Constitutively active calcineurin induces cardiac endoplasmic reticulum stress and protects against apoptosis that is mediated by α-crystallin-B Nicolas Bousettea,b, Shaan Chugha,b, Vincent Fongc,d, Ruth Isserlinc,d, Kyoung-Han Kima, Allen Volchuke, Peter H. Backxa,b, Peter Liuf, Thomas Kislingerg, David H. MacLennanc,d,1, Andrew Emilic,d, and Anthony O. Gramolinia,b,1 aDepartment of Physiology, University of Toronto, Toronto, ON, Canada M5G 1A8; bHeart and Stroke/Richard Lewar Centre of Excellence, University of Toronto, Toronto, ON, Canada M5S 3E2; cThe Banting and Best Department of Medical Research, University of Toronto, Toronto, ON, Canada M5G 1L6; dThe Charles H. Best Institute and Donnelly Center for Cellular and Biomolecular Research, University of Toronto, Toronto, ON, Canada M5S 3E1; eDivision of Cellular and Molecular Biology, Toronto General Research Institute and gDepartment of Medical Biophysics, University Health Network, Toronto, ON, Canada M5G 1L7; and fDepartment of Medicine, University Health Network, Toronto, ON, Canada M5G 2C4 Contributed by David H. MacLennan, September 14, 2010 (sent for review June 30, 2010) Cardiac-specific overexpression of a constitutively active form of calcineurin expression (6), thus our objective was to identify calcineurin A (CNA) leads directly to cardiac hypertrophy in the alterations in protein expression that accompany the pathophysi- CNA mouse model. Because cardiac hypertrophy is a prominent ological mechanisms associated with this form of cardiac disease. characteristic of many cardiomyopathies, we deduced that delineat- CNA mice manifest with a severe hypertrophic phenotype as ing the proteomic profile of ventricular tissue from this model might early as 2 wk after birth; they demonstrate significantly elevated identify novel, widely applicable therapeutic targets. -

Co-Targeting Myelin Inhibitors and Cspgs Markedly Enhances
RESEARCH ARTICLE Co-targeting myelin inhibitors and CSPGs markedly enhances regeneration of GDNF-stimulated, but not conditioning- lesioned, sensory axons into the spinal cord Jinbin Zhai1,2, Hyukmin Kim1,2, Seung Baek Han1,2, Meredith Manire1,2, Rachel Yoo1,2, Shuhuan Pang1,2, George M Smith1,2, Young-Jin Son1,2* 1Shriners Hospitals Pediatric Research Center and Center for Neural Repair and Rehabilitation, Lewis Katz School of Medicine, Temple University, Philadelphia, United States; 2Center for Neural Repair and Rehabilitation, Lewis Katz School of Medicine, Temple University, Philadelphia, United States Abstract A major barrier to intraspinal regeneration after dorsal root (DR) injury is the DR entry zone (DREZ), the CNS/PNS interface. DR axons stop regenerating at the DREZ, even if regenerative capacity is increased by a nerve conditioning lesion. This potent blockade has long been attributed to myelin-associated inhibitors and (CSPGs), but incomplete lesions and conflicting reports have prevented conclusive agreement. Here, we evaluated DR regeneration in mice using novel strategies to facilitate complete lesions and analyses, selective tracing of proprioceptive and mechanoreceptive axons, and the first simultaneous targeting of Nogo/Reticulon-4, MAG, OMgp, CSPGs, and GDNF. Co-eliminating myelin inhibitors and CSPGs elicited regeneration of only a few conditioning-lesioned DR axons across the DREZ. Their absence, however, markedly and synergistically enhanced regeneration of GDNF-stimulated axons, highlighting the importance of *For correspondence: sufficiently elevating intrinsic growth capacity. We also conclude that myelin inhibitors and CSPGs [email protected] are not the primary mechanism stopping axons at the DREZ. Competing interests: The authors declare that no competing interests exist.