Hi-C Detects Novel Structural Variants in HL-60 and HL-60/S4 Cell Lines
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Alzheimer's Disease Protective P522R Variant of PLCG2
bioRxiv preprint doi: https://doi.org/10.1101/2020.04.27.059600; this version posted April 28, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. The Alzheimer’s disease protective P522R variant of PLCG2, consistently enhances stimulus-dependent PLCγ2 activation, depleting substrate and altering cell function. Emily Maguire #1, Georgina E. Menzies#1, Thomas Phillips#1, Michael Sasner2, Harriet M. Williams2, Magdalena A. Czubala3, Neil Evans1, Emma L Cope4, Rebecca Sims5, Gareth R. Howell2, Emyr Lloyd-Evans4, Julie Williams†1,5, Nicholas D. Allen†4 and Philip R. Taylor†*1,3. 1 UK Dementia Research Institute at Cardiff, Hadyn Ellis Building, Maindy Road, Cardiff, CF24 4HQ, Wales, UK. 2 The Jackson Laboratory, Bar Harbor, Maine 04660, USA. 3 Systems Immunity University Research Institute, Tenovus Building, Heath Park, Cardiff CF 14 4XN, Wales, UK. 4 School of Biosciences, Cardiff University, Museum Avenue, Cardiff, CF10 3AX. 5 MRC Centre for Neuropsychiatric Genetics & Genomics, Hadyn Ellis Building, Maindy Road, Cardiff, CF24 4HQ, Wales, UK. #†These authors contributed equally *To whom correspondence should be addressed (lead contact): Prof Philip R. Taylor; Tel: +44(0)2920687328; Email: [email protected]. Abstract: Recent genome-wide association studies of Alzheimer’s disease (AD) have identified variants implicating immune pathways in disease development. A rare coding variant of PLCG2, which encodes PLCγ2, shows a significant protective effect for AD (rs72824905, P522R, P=5.38x10-10, Odds Ratio = 0.68). -

Plasma Lipidome Is Dysregulated in Alzheimer's Disease and Is
Liu et al. Translational Psychiatry (2021) 11:344 https://doi.org/10.1038/s41398-021-01362-2 Translational Psychiatry ARTICLE Open Access Plasma lipidome is dysregulated in Alzheimer’s disease and is associated with disease risk genes Yue Liu1,2, Anbupalam Thalamuthu1, Karen A. Mather1,3, John Crawford1, Marina Ulanova1, Matthew Wai Kin Wong1, Russell Pickford4, Perminder S. Sachdev 1,5 and Nady Braidy 1,6 Abstract Lipidomics research could provide insights of pathobiological mechanisms in Alzheimer’s disease. This study explores a battery of plasma lipids that can differentiate Alzheimer’s disease (AD) patients from healthy controls and determines whether lipid profiles correlate with genetic risk for AD. AD plasma samples were collected from the Sydney Memory and Ageing Study (MAS) Sydney, Australia (aged range 75–97 years; 51.2% male). Untargeted lipidomics analysis was performed by liquid chromatography coupled–mass spectrometry (LC–MS/MS). We found that several lipid species from nine lipid classes, particularly sphingomyelins (SMs), cholesterol esters (ChEs), phosphatidylcholines (PCs), phosphatidylethanolamines (PIs), phosphatidylinositols (PIs), and triglycerides (TGs) are dysregulated in AD patients and may help discriminate them from healthy controls. However, when the lipid species were grouped together into lipid subgroups, only the DG group was significantly higher in AD. ChEs, SMs, and TGs resulted in good classification accuracy using the Glmnet algorithm (elastic net penalization for the generalized linear model [glm]) with more than 80% AUC. In general, group lipids and the lipid subclasses LPC and PE had less classification accuracy compared to the other subclasses. We also found significant increases in SMs, PIs, and the LPE/PE ratio in human U251 astroglioma cell lines exposed to pathophysiological concentrations of oligomeric Aβ42. -

Antibody Response Cell Antigen Receptor Signaling And
Lysophosphatidic Acid Receptor 5 Inhibits B Cell Antigen Receptor Signaling and Antibody Response This information is current as Jiancheng Hu, Shannon K. Oda, Kristin Shotts, Erin E. of September 24, 2021. Donovan, Pamela Strauch, Lindsey M. Pujanauski, Francisco Victorino, Amin Al-Shami, Yuko Fujiwara, Gabor Tigyi, Tamas Oravecz, Roberta Pelanda and Raul M. Torres J Immunol 2014; 193:85-95; Prepublished online 2 June 2014; Downloaded from doi: 10.4049/jimmunol.1300429 http://www.jimmunol.org/content/193/1/85 Supplementary http://www.jimmunol.org/content/suppl/2014/05/31/jimmunol.130042 http://www.jimmunol.org/ Material 9.DCSupplemental References This article cites 63 articles, 17 of which you can access for free at: http://www.jimmunol.org/content/193/1/85.full#ref-list-1 Why The JI? Submit online. by guest on September 24, 2021 • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2014 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Lysophosphatidic Acid Receptor 5 Inhibits B Cell Antigen Receptor Signaling and Antibody Response Jiancheng Hu,*,1,2 Shannon K. -

Survival-Associated Metabolic Genes in Colon and Rectal Cancers
Survival-associated Metabolic Genes in Colon and Rectal Cancers Yanfen Cui ( [email protected] ) Tianjin Cancer Institute: Tianjin Tumor Hospital https://orcid.org/0000-0001-7760-7503 Baoai Han tianjin tumor hospital He Zhang tianjin tumor hospital Zhiyong Wang tianjin tumor hospital Hui Liu tianjin tumor hospital Fei Zhang tianjin tumor hospital Ruifang Niu tianjin tumor hospital Research Keywords: colon cancer, rectal cancer, prognosis, metabolism Posted Date: December 4th, 2020 DOI: https://doi.org/10.21203/rs.3.rs-117478/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/42 Abstract Background Uncontrolled proliferation is the most prominent biological feature of tumors. To rapidly proliferate and maximize the use of available nutrients, tumor cells regulate their metabolic behavior and the expression of metabolism-related genes (MRGs). In this study, we aimed to construct prognosis models for colon and rectal cancers, using MRGs to indicate the prognoses of patients. Methods We rst acquired the gene expression proles of colon and rectal cancers from the TCGA and GEO database, and utilized univariate Cox analysis, lasso regression, and multivariable cox analysis to identify MRGs for risk models. Then GSEA and KEGG functional enrichment analysis were utilized to identify the metabolism pathway of MRGs in the risk models and analyzed these genes comprehensively using GSCALite. Results Eight genes (CPT1C, PLCB2, PLA2G2D, GAMT, ENPP2, PIP4K2B, GPX3, and GSR) in the colon cancer risk model and six genes (TDO2, PKLR, GAMT, EARS2, ACO1, and WAS) in the rectal cancer risk model were identied successfully. Multivariate Cox analysis indicated that the models predicted overall survival accurately and independently for patients with colon or rectal cancer. -

S41436-020-01011-X.Pdf
Zurich Open Repository and Archive University of Zurich Main Library Strickhofstrasse 39 CH-8057 Zurich www.zora.uzh.ch Year: 2020 New insights into the clinical and molecular spectrum of the novel CYFIP2-related neurodevelopmental disorder and impairment of the WRC-mediated actin dynamics Begemann, Anaïs ; Sticht, Heinrich ; Begtrup, Amber ; Vitobello, Antonio ; Faivre, Laurence ; Asadollahi, Reza ; Zweier, Markus ; Steindl, Katharina ; Rauch, Anita Abstract: PURPOSE A few de novo missense variants in the cytoplasmic FMRP-interacting protein 2 (CYFIP2) gene have recently been described as a novel cause of severe intellectual disability, seizures, and hypotonia in 18 individuals, with p.Arg87 substitutions in the majority. METHODS We assembled data from 19 newly identified and all 18 previously published individuals with CYFIP2 variants. By structural modeling and investigation of WAVE-regulatory complex (WRC)-mediated actin polymerization in six patient fibroblast lines we assessed the impact of CYFIP2 variants on the WRC. RESULTS Sixteenof 19 individuals harbor two previously described and 11 novel (likely) disease-associated missense variants. We report p.Asp724 as second mutational hotspot (4/19 cases). Genotype-phenotype correlation con- firms a consistently severe phenotype in p.Arg87 patients but a more variable phenotype inp.Asp724 and other substitutions. Three individuals with milder phenotypes carry putative loss-of-function vari- ants, which remain of unclear pathogenicity. Structural modeling predicted missense variants to disturb interactions within the WRC or impair CYFIP2 stability. Consistent with its role in WRC-mediated actin polymerization we substantiate aberrant regulation of the actin cytoskeleton in patient fibroblasts. CONCLUSION Our study expands the clinical and molecular spectrum of CYFIP2-related neurodevel- opmental disorder and provides evidence for aberrant WRC-mediated actin dynamics as contributing cellular pathomechanism. -

Supplemental Information
Supplemental information Dissection of the genomic structure of the miR-183/96/182 gene. Previously, we showed that the miR-183/96/182 cluster is an intergenic miRNA cluster, located in a ~60-kb interval between the genes encoding nuclear respiratory factor-1 (Nrf1) and ubiquitin-conjugating enzyme E2H (Ube2h) on mouse chr6qA3.3 (1). To start to uncover the genomic structure of the miR- 183/96/182 gene, we first studied genomic features around miR-183/96/182 in the UCSC genome browser (http://genome.UCSC.edu/), and identified two CpG islands 3.4-6.5 kb 5’ of pre-miR-183, the most 5’ miRNA of the cluster (Fig. 1A; Fig. S1 and Seq. S1). A cDNA clone, AK044220, located at 3.2-4.6 kb 5’ to pre-miR-183, encompasses the second CpG island (Fig. 1A; Fig. S1). We hypothesized that this cDNA clone was derived from 5’ exon(s) of the primary transcript of the miR-183/96/182 gene, as CpG islands are often associated with promoters (2). Supporting this hypothesis, multiple expressed sequences detected by gene-trap clones, including clone D016D06 (3, 4), were co-localized with the cDNA clone AK044220 (Fig. 1A; Fig. S1). Clone D016D06, deposited by the German GeneTrap Consortium (GGTC) (http://tikus.gsf.de) (3, 4), was derived from insertion of a retroviral construct, rFlpROSAβgeo in 129S2 ES cells (Fig. 1A and C). The rFlpROSAβgeo construct carries a promoterless reporter gene, the β−geo cassette - an in-frame fusion of the β-galactosidase and neomycin resistance (Neor) gene (5), with a splicing acceptor (SA) immediately upstream, and a polyA signal downstream of the β−geo cassette (Fig. -

Description Cy5 LG Cy3 MI Ratio(Cy3/Cy5) C9orf135
Supplementary Table S2. DNA microarray dataset of top 30 differentially expressed genes and housekeeping genes Up-regulated in SI cancer cells Symbol Description Cy5_LG Cy3_MI Ratio(Cy3/Cy5) C9orf135 Uncharacterized protein C9orf135 1.37 307.8 224.3 KIAA1245 Notch homolog 2 N-terminal like protein 1.82 406.8 223.6 APITD1 Centromere protein S (CENP-S) 2.56 560.3 218.7 PPIL6 Peptidyl-prolyl cis-trans isomerase-like 6 1.85 343.0 185.7 MYCBP2 Probable E3 ubiquitin-protein ligase MYCBP2 2.01 332.1 165.6 ANGPTL4 Angiopoietin-related protein 4 precursor 10.26 1500.1 146.3 C10orf79 Novel protein (Fragment) 2.86 415.5 145.5 NP_653323.1 KPL2 protein isoform 2 1.82 257.5 141.6 ZNF345 Zinc finger protein 345 1.58 215.7 136.8 SIX1 Homeobox protein SIX1 1.59 216.1 136.0 KLHL7 Kelch-like protein 7 1.91 254.4 133.3 TBX1 T-box transcription factor TBX1 1.57 209.0 133.1 PAG1 Phosphoprotein associated with glycosphingolipid-enriched microdomains 1 2.19 290.9 133.0 NOL12 Nucleolar protein 12 1.61 203.7 126.5 ZNF606 Zinc finger protein 606 2.12 267.7 126.0 NFKBIE NF-kappa-B inhibitor epsilon 5.28 658.3 124.7 ZMYND10 Zinc finger MYND domain-containing protein 10 6.85 835.5 121.9 hCG_23177 - 14.94 1758.9 117.7 KIF3A Kinesin-like protein KIF3A 1.94 224.9 116.1 Q9C0K3_HUMAN Actin-related protein Arp11 3.38 368.0 108.9 NP_056263.1 DPCD protein 2.61 270.5 103.7 GBP1 Interferon-induced guanylate-binding protein 1 1.46 149.1 102.3 NP_660151.2 NAD(P) dependent steroid dehydrogenase-like 1.35 137.4 101.5 NP_689672.2 CDNA FLJ90761 fis, clone THYRO1000099 3.68 372.5 -
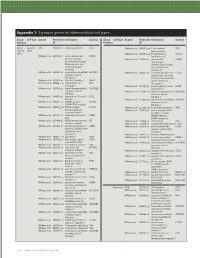
Technical Note, Appendix: an Analysis of Blood Processing Methods to Prepare Samples for Genechip® Expression Profiling (Pdf, 1
Appendix 1: Signature genes for different blood cell types. Blood Cell Type Source Probe Set Description Symbol Blood Cell Type Source Probe Set Description Symbol Fraction ID Fraction ID Mono- Lympho- GSK 203547_at CD4 antigen (p55) CD4 Whitney et al. 209813_x_at T cell receptor TRG nuclear cytes gamma locus cells Whitney et al. 209995_s_at T-cell leukemia/ TCL1A Whitney et al. 203104_at colony stimulating CSF1R lymphoma 1A factor 1 receptor, Whitney et al. 210164_at granzyme B GZMB formerly McDonough (granzyme 2, feline sarcoma viral cytotoxic T-lymphocyte- (v-fms) oncogene associated serine homolog esterase 1) Whitney et al. 203290_at major histocompatibility HLA-DQA1 Whitney et al. 210321_at similar to granzyme B CTLA1 complex, class II, (granzyme 2, cytotoxic DQ alpha 1 T-lymphocyte-associated Whitney et al. 203413_at NEL-like 2 (chicken) NELL2 serine esterase 1) Whitney et al. 203828_s_at natural killer cell NK4 (H. sapiens) transcript 4 Whitney et al. 212827_at immunoglobulin heavy IGHM Whitney et al. 203932_at major histocompatibility HLA-DMB constant mu complex, class II, Whitney et al. 212998_x_at major histocompatibility HLA-DQB1 DM beta complex, class II, Whitney et al. 204655_at chemokine (C-C motif) CCL5 DQ beta 1 ligand 5 Whitney et al. 212999_x_at major histocompatibility HLA-DQB Whitney et al. 204661_at CDW52 antigen CDW52 complex, class II, (CAMPATH-1 antigen) DQ beta 1 Whitney et al. 205049_s_at CD79A antigen CD79A Whitney et al. 213193_x_at T cell receptor beta locus TRB (immunoglobulin- Whitney et al. 213425_at Homo sapiens cDNA associated alpha) FLJ11441 fis, clone Whitney et al. 205291_at interleukin 2 receptor, IL2RB HEMBA1001323, beta mRNA sequence Whitney et al. -

Time Resolved Quantitative Phosphoproteomics Reveals Distinct Patterns of SHP2
bioRxiv preprint doi: https://doi.org/10.1101/598664; this version posted April 12, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Time resolved quantitative phosphoproteomics reveals distinct patterns of SHP2 dependence in EGFR signaling Vidyasiri Vemulapalli1,2, Lily Chylek3, Alison Erickson4, Jonathan LaRochelle1,2, Kartik Subramanian3, Morvarid Mohseni5, Matthew LaMarche5, Michael G. Acker5, Peter K. Sorger3, Steven P. Gygi4, and Stephen C. Blacklow1,2* 1Department of Cancer Biology, Dana-Farber Cancer Institute Boston, MA 02115, USA 2Department of Biological Chemistry & Molecular Pharmacology, Blavatnik Institute, Harvard Medical School, Boston, MA 02115, USA 3Laboratory of Systems Pharmacology, Harvard Medical School, Boston, MA 02115, USA 4Department of Cell Biology, Harvard Medical School, Boston, MA 02115, USA 5Novartis Institutes for Biomedical Research, Cambridge, MA, 02139, USA *To whom correspondence should be addressed: [email protected] bioRxiv preprint doi: https://doi.org/10.1101/598664; this version posted April 12, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Abstract SHP2 is a protein tyrosine phosphatase that normally potentiates intracellular signaling by growth factors, antigen receptors, and some cytokines; it is frequently mutated in childhood leukemias and other cancers. Here, we examine the role of SHP2 in the responses of breast cancer cells to EGF by monitoring phosphoproteome dynamics when SHP2 is allosterically inhibited by the small molecule SHP099. -

A-To-I RNA Editing Does Not Change with Age in the Healthy Male Rat Brain
Biogerontology (2013) 14:395–400 DOI 10.1007/s10522-013-9433-8 RESEARCH ARTICLE A-to-I RNA editing does not change with age in the healthy male rat brain Andrew P. Holmes • Shona H. Wood • Brian J. Merry • Joa˜o Pedro de Magalha˜es Received: 18 January 2013 / Accepted: 15 May 2013 / Published online: 26 May 2013 Ó The Author(s) 2013. This article is published with open access at Springerlink.com Abstract RNA editing is a post-transcriptional pro- Introduction cess, which results in base substitution modifications to RNA. It is an important process in generating Adenosine to inosine (A-to-I) RNA editing is a post- protein diversity through amino acid substitution and transcriptional process that alters the sequences of the modulation of splicing events. Previous studies RNA molecules. The adenosine deaminases ADAR have suggested a link between gene-specific reduc- and ADARB1 convert specific adenosine residues on tions in adenosine to inosine RNA editing and aging in RNA to inosine bases. During translation, sequencing, the human brain. Here we demonstrate that changes in and splicing, inosine is recognized as guanosine. RNA editing observed in humans with age are not Therefore, A-to-I RNA editing has important impli- observed during aging in healthy rats. Furthermore, we cations in altering specific amino acids, miRNA identify a conserved editing site in rats, in Cog3.We targeting, and in the modulation of alternative splicing propose that either age-related changes in RNA (Nishikura 2010). editing are specific to primates or humans, or that Targets of A-to-I RNA editing are often genes they are the manifestation of disease pathology. -

Spatially Clustering De Novo Variants in CYFIP2, Encoding the Cytoplasmic FMRP Interacting Protein 2, Cause Intellectual Disability and Seizures
European Journal of Human Genetics (2019) 27:747–759 https://doi.org/10.1038/s41431-018-0331-z ARTICLE Spatially clustering de novo variants in CYFIP2, encoding the cytoplasmic FMRP interacting protein 2, cause intellectual disability and seizures 1 1,2 3 3 2,4 5,6 Markus Zweier ● Anaïs Begemann ● Kirsty McWalter ● Megan T. Cho ● Lucia Abela ● Siddharth Banka ● 7 8 9,10 11 12 Bettina Behring ● Andrea Berger ● Chester W. Brown ● Maryline Carneiro ● Jiani Chen ● 13 14 13 Gregory M. Cooper ● Deciphering Developmental Disorders (DDD) Study ● Candice R. Finnila ● 3 15 16 1 5,6 17,18 Maria J. Guillen Sacoto ● Alex Henderson ● Ulrike Hüffmeier ● Pascal Joset ● Bronwyn Kerr ● Gaetan Lesca ● 19 5 20 3 9 Gloria S. Leszinski ● John Henry McDermott ● Meira R. Meltzer ● Kristin G. Monaghan ● Roya Mostafavi ● 21,22 2,4,23 24 12 21,22,25 Katrin Õunap ● Barbara Plecko ● Zöe Powis ● Gabriela Purcarin ● Tiia Reimand ● 19,26 20 27 12,28 29 Korbinian M. Riedhammer ● John M. Schreiber ● Deepa Sirsi ● Klaas J. Wierenga ● Monica H. Wojcik ● 1,30 1 31 1,2,32,33 Sorina M. Papuc ● Katharina Steindl ● Heinrich Sticht ● Anita Rauch Received: 30 May 2018 / Revised: 31 October 2018 / Accepted: 7 November 2018 / Published online: 21 January 2019 © European Society of Human Genetics 2019 1234567890();,: 1234567890();,: Abstract CYFIP2, encoding the evolutionary highly conserved cytoplasmic FMRP interacting protein 2, has previously been proposed as a candidate gene for intellectual disability and autism because of its important role linking FMRP-dependent transcription regulation and actin polymerization via the WAVE regulatory complex (WRC). Recently, de novo variants affecting the amino acid p.Arg87 of CYFIP2 were reported in four individuals with epileptic encephalopathy. -

Mutational Landscape and Clinical Outcome of Patients with De Novo Acute Myeloid Leukemia and Rearrangements Involving 11Q23/KMT2A
Mutational landscape and clinical outcome of patients with de novo acute myeloid leukemia and rearrangements involving 11q23/KMT2A Marius Billa,1,2, Krzysztof Mrózeka,1,2, Jessica Kohlschmidta,b, Ann-Kathrin Eisfelda,c, Christopher J. Walkera, Deedra Nicoleta,b, Dimitrios Papaioannoua, James S. Blachlya,c, Shelley Orwicka,c, Andrew J. Carrolld, Jonathan E. Kolitze, Bayard L. Powellf, Richard M. Stoneg, Albert de la Chapelleh,i,2, John C. Byrda,c, and Clara D. Bloomfielda,c aThe Ohio State University Comprehensive Cancer Center, Columbus, OH 43210; bAlliance for Clinical Trials in Oncology Statistics and Data Center, The Ohio State University Comprehensive Cancer Center, Columbus, OH 43210; cDivision of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, OH 43210; dDepartment of Genetics, University of Alabama at Birmingham, Birmingham, AL 35294; eNorthwell Health Cancer Institute, Zucker School of Medicine at Hofstra/Northwell, Lake Success, NY 11042; fDepartment of Internal Medicine, Section on Hematology & Oncology, Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, NC 27157; gDepartment of Medical Oncology, Dana-Farber/Partners Cancer Care, Boston, MA 02215; hHuman Cancer Genetics Program, Comprehensive Cancer Center, The Ohio State University, Columbus, OH 43210; and iDepartment of Cancer Biology and Genetics, Comprehensive Cancer Center, The Ohio State University, Columbus, OH 43210 Contributed by Albert de la Chapelle, August 28, 2020 (sent for review July 17, 2020; reviewed by Anne Hagemeijer and Stefan Klaus Bohlander) Balanced rearrangements involving the KMT2A gene, located at patterns that include high expression of HOXA genes and thereby 11q23, are among the most frequent chromosome aberrations in contribute to leukemogenesis (14–16).