DDB1 Is Essential for Genomic Stability in Developing Epidermis
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

DDB1 Targets Chk1 to the Cul4 E3 Ligase Complex in Normal Cycling Cells and in Cells Experiencing Replication Stress
Published OnlineFirst March 10, 2009; DOI: 10.1158/0008-5472.CAN-08-3382 Research Article DDB1 Targets Chk1 to the Cul4 E3 Ligase Complex in Normal Cycling Cells and in Cells Experiencing Replication Stress Van Leung-Pineda,1 Jiwon Huh,1 and Helen Piwnica-Worms1,2,3 Departments of 1Cell Biology and Physiology and 2Internal Medicine and 3Howard Hughes Medical Institute, Washington University School of Medicine, St. Louis, Missouri Abstract protein FANCE (8, 10–12). Chk1 carries out its functions both in the The Chk1 protein kinase preserves genome integrity in normal nucleus and at the centrosome (13). Drugs that block Chk1 kinase proliferating cells and in cells experiencing replicative and activity or enhance its proteolysis by interfering with binding to genotoxic stress. Chk1 is currently being targeted in antican- heat shock protein 90 (HSP90) are currently being tested as cer regimens. Here, we identify damaged DNA-binding protein anticancer agents (14–17). 1 (DDB1) as a novel Chk1-interacting protein. DDB1 is part of Chk1 is regulated by reversible phosphorylation and by an E3 ligase complex that includes the cullin proteins Cul4A ubiquitin-mediated proteolysis. Under periods of replicative stress, and Cul4B. We report that Cul4A/DDB1 negatively regulates the ATRIP/ATR module binds to single-stranded DNA and, Chk1 stability in vivo. Chk1 associates with Cul4A/DDB1 together with Rad17 and the 9-1-1 complex, activates Chk1 in a Claspin-dependent manner (18–22). ATR directly phosphorylates during an unperturbed cell division cycle and both Chk1 317 345 phosphorylation and replication stress enhanced these inter- Chk1 on two COOH-terminal residues: Ser (S317) and Ser actions. -

The Differential Expression of Core Genes in Nucleotide Excision Repair Pathway Indicates Colorectal Carcinogenesis and Prognosis
Hindawi BioMed Research International Volume 2018, Article ID 9651320, 10 pages https://doi.org/10.1155/2018/9651320 Research Article The Differential Expression of Core Genes in Nucleotide Excision Repair Pathway Indicates Colorectal Carcinogenesis and Prognosis Jingwei Liu, Hao Li, Liping Sun, Xue Feng, Zhenning Wang , Yuan Yuan , and Chengzhong Xing Tumor Etiology and Screening Department, Cancer Institute and General Surgery, Te First Hospital of China Medical University and Key Laboratory of Cancer Etiology and Prevention, Liaoning Provincial Education Department, China Medical University, Shenyang 110001, China Correspondence should be addressed to Yuan Yuan; [email protected] and Chengzhong Xing; [email protected] Received 19 October 2017; Revised 12 December 2017; Accepted 14 December 2017; Published 15 January 2018 Academic Editor: Paul W. Doetsch Copyright © 2018 Jingwei Liu et al. Tis is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Background. Nucleotide excision repair (NER) plays a critical role in maintaining genome integrity. Tis study aimed to investigate theexpressionofNERgenesandtheirassociationswithcolorectalcancer(CRC)development.Method. Expressions of NER genes in CRC and normal tissues were analysed by ONCOMINE. Te Cancer Genome Atlas (TCGA) data were downloaded to explore relationship of NER expression with clinicopathological parameters and survival of CRC. Results. ERCC1, ERCC2, ERCC5, and DDB2 were upregulated while ERCC4 was downregulated in CRC. For colon cancer, high ERCC3 expression was related to better T stage; ERCC5 expression indicated deeper T stage and distant metastasis; DDB2 expression suggested earlier TNM stage. For rectal cancer, ERCC2 expression correlated with favourable T stage; XPA expression predicted worse TNM stage. -

(UV-DDB) Dimerization and Its Roles in Chromatinized DNA Repair
Damaged DNA induced UV-damaged DNA-binding protein (UV-DDB) dimerization and its roles in chromatinized DNA repair Joanne I. Yeha,b,1, Arthur S. Levinec,d, Shoucheng Dua, Unmesh Chintea, Harshad Ghodkee, Hong Wangd,e, Haibin Shia, Ching L. Hsiehc,d, James F. Conwaya, Bennett Van Houtend,e, and Vesna Rapić-Otrinc,d aDepartments of Structural Biology, bBioengineering, cMicrobiology and Molecular Genetics, ePharmacology and Chemical Biology, and dUniversity of Pittsburgh Cancer Institute, University of Pittsburgh School of Medicine, Pittsburgh, PA 15260 AUTHOR SUMMARY Exposure to UV radiation (DDB1-CUL4A DDB2) with can damage DNA that if chromatin modification and left unrepaired can cause the subsequent steps in the mutations leading to skin repair pathway. aging and skin cancer. In Here we report the crystal humans, the nucleotide structure of the full-length excision repair (NER) * human DDB2 bound to proteins function to damaged DNA in a complex recognize and repair with human DDB1 (Fig. P1). UV-damaged DNA. Defects While a large portion of the in DNA repair caused by N-terminal region of the mutations of these repair zebrafish DDB2 in the proteins have been linked earlier structure could not to several genetic diseases, be modeled, we have characterized by cancer resolved the 3D structure of predisposition (xeroderma the N-terminal domain of pigmentosum, XP) or DDB2. Our structure reveals premature aging (Cockayne secondary interactions syndrome), illustrating the between the N-terminal Fig. P1. Composite model of a dimeric DDB1-CUL4ADDB2 ubiquitin functional significance of DDB2 domain of DDB2 and a ligase-nucleosome complex. A model of a dimeric DDB1-CUL4A in a neighboring repair proteins to genomic complex with a nucleosome core particle, generated according to the relative integrity. -

Epigenetic Regulation of DNA Repair Genes and Implications for Tumor Therapy ⁎ ⁎ Markus Christmann , Bernd Kaina
Mutation Research-Reviews in Mutation Research xxx (xxxx) xxx–xxx Contents lists available at ScienceDirect Mutation Research-Reviews in Mutation Research journal homepage: www.elsevier.com/locate/mutrev Review Epigenetic regulation of DNA repair genes and implications for tumor therapy ⁎ ⁎ Markus Christmann , Bernd Kaina Department of Toxicology, University of Mainz, Obere Zahlbacher Str. 67, D-55131 Mainz, Germany ARTICLE INFO ABSTRACT Keywords: DNA repair represents the first barrier against genotoxic stress causing metabolic changes, inflammation and DNA repair cancer. Besides its role in preventing cancer, DNA repair needs also to be considered during cancer treatment Genotoxic stress with radiation and DNA damaging drugs as it impacts therapy outcome. The DNA repair capacity is mainly Epigenetic silencing governed by the expression level of repair genes. Alterations in the expression of repair genes can occur due to tumor formation mutations in their coding or promoter region, changes in the expression of transcription factors activating or Cancer therapy repressing these genes, and/or epigenetic factors changing histone modifications and CpG promoter methylation MGMT Promoter methylation or demethylation levels. In this review we provide an overview on the epigenetic regulation of DNA repair genes. GADD45 We summarize the mechanisms underlying CpG methylation and demethylation, with de novo methyl- TET transferases and DNA repair involved in gain and loss of CpG methylation, respectively. We discuss the role of p53 components of the DNA damage response, p53, PARP-1 and GADD45a on the regulation of the DNA (cytosine-5)- methyltransferase DNMT1, the key enzyme responsible for gene silencing. We stress the relevance of epigenetic silencing of DNA repair genes for tumor formation and tumor therapy. -

DNA Methylation of Telomere-Related Genes and Cancer Risk
Author Manuscript Published OnlineFirst on June 12, 2018; DOI: 10.1158/1940-6207.CAPR-17-0413 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. DNA Methylation of Telomere-Related Genes and Cancer Risk Brian T Joyce1, Yinan Zheng1, Drew Nannini1, Zhou Zhang1, Lei Liu2, Tao Gao1, Masha Kocherginsky3, Robert Murphy4, Hushan Yang5, Chad J. Achenbach6, Lewis R. Roberts7, Mirjam Hoxha8, Jincheng Shen9, Pantel Vokonas10,11, Joel Schwartz12, Andrea Baccarelli13, Lifang Hou1 1. Center for Population Epigenetics, Robert H. Lurie Comprehensive Cancer Center and Department of Preventive Medicine, Northwestern University Feinberg School of Medicine, 680 N. Lake Shore Dr., Suite 1400, Chicago, IL, 60611, USA 2. Division of Biostatistics, Washington University in St. Louis, USA 3. Department of Preventive Medicine, Northwestern University Feinberg School of Medicine, USA 4. Center for Global Health, Feinberg School of Medicine, Northwestern University, USA 5. Division of Population Science, Department of Medical Oncology, Sidney Kimmel Cancer Center, Thomas Jefferson University, USA 6. Department of Medicine, Northwestern University Feinberg School of Medicine, USA 7. Division of Gastroenterology and Hepatology, Department of Medicine, Mayo Clinic, USA 8. Molecular Epidemiology and Environmental Epigenetics Laboratory, Department of Clinical Sciences and Community Health, Università degli Studi di Milano, Italy 9. Department of Population Health Sciences, University of Utah School of Medicine, USA 10. VA Normative Aging Study, VA Boston Healthcare System, USA 11. Department of Medicine, Boston University School of Medicine, USA 12. Department of Environmental Health, Harvard School of Public Health, USA 13. Department of Environmental Health Science, Mailman School of Public Health, Columbia University, USA Journal: Cancer Prevention Research Word Count (limit 5000): 3909 Number of Tables/Figures (limit 6): 6 1 Downloaded from cancerpreventionresearch.aacrjournals.org on September 24, 2021. -
Neurodegeneration in Accelerated Aging
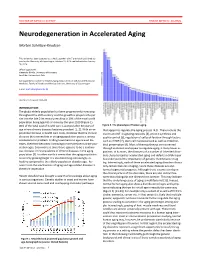
DOCTOR OF MEDICAL SCIENCE DANISH MEDICAL JOURNAL Neurodegeneration in Accelerated Aging Morten Scheibye-Knudsen This review has been accepted as a thesis together with 7 previously published pa- pers by the University of Copenhagen, October 16, 2014 and defended on January 14, 2016 Official opponents: Alexander Bürkle, University of Konstanz Lars Eide, University of Oslo Correspondence: Center for Healthy Aging, Department of Cellular and Molecular Medicine, Faculty of Health and Medical Sciences, University of Copenhagen E-mail: [email protected] Dan Med J 2016;63(11):B5308 INTRODUCTION The global elderly population has been progressively increasing throughout the 20th century and this growth is projected to per- sist into the late 21st century resulting in 20% of the total world population being aged 65 or more by the year 2100 (Figure 1). 80% of the total cost of health care is accrued after 40 years of Figure 2. The phenotype of human aging. age where chronic diseases become prevalent [1, 2]. With an ex- that appear to regulate the aging process [4,5]. These include the ponential increase in health care costs, it follows that the chronic insulin and IGF-1 signaling cascades [4], protein synthesis and diseases that accumulate in an aging population poses a serious quality control [6], regulation of cell proliferation through factors socioeconomic problem. Finding treatments to age related dis- such as mTOR [7], stem cell maintenance 8 as well as mitochon- eases, therefore becomes increasingly more pertinent as the pop- drial preservation [9]. Most of these pathways are conserved ulation ages. Even more so since there appears to be a continu- through evolution and appear to regulate aging in many lower or- ous increase in the prevalence of chronic diseases in the aging ganisms. -

Regulation of the Intranuclear Distribution of the Cockayne Syndrome Proteins Received: 26 July 2017 Teruaki Iyama, Mustafa N
www.nature.com/scientificreports OPEN Regulation of the Intranuclear Distribution of the Cockayne Syndrome Proteins Received: 26 July 2017 Teruaki Iyama, Mustafa N. Okur, Tyler Golato, Daniel R. McNeill, Huiming Lu , Accepted: 1 November 2018 Royce Hamilton, Aishwarya Raja, Vilhelm A. Bohr & David M. Wilson III Published: xx xx xxxx Cockayne syndrome (CS) is an inherited disorder that involves photosensitivity, developmental defects, progressive degeneration and characteristics of premature aging. Evidence indicates primarily nuclear roles for the major CS proteins, CSA and CSB, specifcally in DNA repair and RNA transcription. We reveal herein a complex regulation of CSB targeting that involves three major consensus signals: NLS1 (aa467-481), which directs nuclear and nucleolar localization in cooperation with NoLS1 (aa302-341), and NLS2 (aa1038-1055), which seemingly optimizes nuclear enrichment. CSB localization to the nucleolus was also found to be important for full UVC resistance. CSA, which does not contain any obvious targeting sequences, was adversely afected (i.e. presumably destabilized) by any form of truncation. No inter-coordination between the subnuclear localization of CSA and CSB was observed, implying that this aspect does not underlie the clinical features of CS. The E3 ubiquitin ligase binding partner of CSA, DDB1, played an important role in CSA stability (as well as DDB2), and facilitated CSA association with chromatin following UV irradiation; yet did not afect CSB chromatin binding. We also observed that initial recruitment of CSB to DNA interstrand crosslinks is similar in the nucleoplasm and nucleolus, although fnal accumulation is greater in the former. Whereas assembly of CSB at sites of DNA damage in the nucleolus was not afected by RNA polymerase I inhibition, stable retention at these sites of presumed repair was abrogated. -

Nuclear and Cytoplasmic WDR-23 Isoforms Mediate Differential Effects
www.nature.com/scientificreports OPEN Nuclear and cytoplasmic WDR- 23 isoforms mediate diferential efects on GEN-1 and SKN-1 Received: 13 May 2019 Accepted: 1 August 2019 substrates Published: xx xx xxxx Brett N. Spatola1,2, Jacqueline Y. Lo1,2,3, Bin Wang4 & Sean P. Curran1,2,5 Maintaining a healthy cellular environment requires the constant control of proteostasis. E3 ubiquitin ligase complexes facilitate the post-translational addition of ubiquitin, which based on the quantity and specifc lysine linkages, results in diferent outcomes. Our studies reveal the CUL4-DDB1 substrate receptor, WDR23, as both a positive and a negative regulator in cellular stress responses. These opposing roles are mediated by two distinct isoforms: WDR-23A in the cytoplasm and WDR-23B in the nucleus. C. elegans expressing only WDR-23A display activation of SKN-1 and enhanced survival to oxidative stress, whereas animals with restricted WDR-23B expression do not. Additionally, we identify GEN-1, a Holliday junction resolvase, as an evolutionarily conserved WDR-23 substrate and fnd that the nuclear and cytoplasmic isoforms of WDR-23 diferentially afect double-strand break repair. Our results suggest that through diferential ubiquitination, nuclear WDR-23B inhibits the activity of substrates, most likely by promoting protein turnover, while cytoplasmic WDR-23A performs a proteasome- independent role. Together, our results establish a cooperative role between two spatially distinct isoforms of WDR-23 in ensuring proper regulation of WDR-23 substrates. Proteostasis plays an integral part in ensuring organismal survival, and numerous diseases can arise when the balance of the creation, maintenance, and degradation of proteins is dysregulated1,2. -

Defective DNA Repair and Chromatin Organization in Patients with Quiescent Systemic Lupus Erythematosus Vassilis L
Souliotis et al. Arthritis Research & Therapy (2016) 18:182 DOI 10.1186/s13075-016-1081-3 RESEARCH ARTICLE Open Access Defective DNA repair and chromatin organization in patients with quiescent systemic lupus erythematosus Vassilis L. Souliotis1,2*, Konstantinos Vougas3, Vassilis G. Gorgoulis3,4 and Petros P. Sfikakis2 Abstract Background: Excessive autoantibody production characterizing systemic lupus erythematosus (SLE) occurs irrespective of the disease’s clinical status and is linked to increased lymphocyte apoptosis. Herein, we tested the hypothesis that defective DNA damage repair contributes to increased apoptosis in SLE. Methods: We evaluated nucleotide excision repair at the N-ras locus, DNA double-strand breaks repair and apoptosis rates in peripheral blood mononuclear cells from anti-dsDNA autoantibody-positive patients (six with quiescent disease and six with proliferative nephritis) and matched healthy controls following ex vivo treatment with melphalan. Chromatin organization and expression levels of DNA repair- and apoptosis-associated genes were also studied in quiescent SLE. Results: Defective nucleotide excision repair and DNA double-strand breaks repair were found in SLE, with lupus nephritis patients showing higher DNA damage levels than those with quiescent disease. Melphalan-induced apoptosis rates were higher in SLE than control cells and correlated inversely with DNA repair efficiency. Chromatin at the N-ras locus was more condensed in SLE than controls, while treatment with the histone deacetylase inhibitor vorinostat resulted in hyperacetylation of histone H4, chromatin decondensation, amelioration of DNA repair efficiency and decreased apoptosis. Accordingly, genes involved in DNA damage repair and signaling pathways, such as DDB1, ERCC2, XPA, XPC, MRE11A, RAD50, PARP1, MLH1, MLH3, and ATM were significantly underexpressed in SLE versus controls, whereas PPP1R15A, BARD1 and BBC3 genes implicated in apoptosis were significantly overexpressed. -

Nucleotide Excision Repair Gene Expression After Cisplatin Treatment in Melanoma
Published OnlineFirst August 31, 2010; DOI: 10.1158/0008-5472.CAN-10-0161 Molecular and Cellular Pathobiology Cancer Research Nucleotide Excision Repair Gene Expression after Cisplatin Treatment in Melanoma Nikola A. Bowden1,2,3, Katie A. Ashton1,2,3, Kelly A. Avery-Kiejda4, Xu Dong Zhang4, Peter Hersey4, and Rodney J. Scott1,2,3,5 Abstract Two of the hallmark features of melanoma are its development as a result of chronic UV radiation exposure and the limited efficacy of cisplatin in the disease treatment. Both of these DNA-damaging agents result in large helix-distorting DNA damage that is recognized and repaired by nucleotide excision repair (NER). The aim of this study was to examine the expression of NER gene transcripts, p53, and p21 in melanoma cell lines treated with cisplatin compared with melanocytes. Basal expression of all genes was greater in the melanoma cell lines compared with melanocytes. Global genome repair (GGR) transcripts showed significantly decreased relative expression (RE) in melanoma cell lines 24 hours after cisplatin treatment. The basal RE of p53 was significantly higher in the melanoma cell lines compared with the melanocytes. However, induction of p53 was only significant in the melanocytes at 6 and 24 hours after cisplatin treatment. Inhibition of p53 expression significantly decreased the expression of all the GGR transcripts in melanocytes at 6 and 24 hours after cis- platin treatment. Although the RE levels were lower with p53 inhibition, the induction of the GGR genes was very similar to that in the control melanocytes and increased significantly across the time points. The findings from this study revealed reduced GGR transcript levels in melanoma cells 24 hours after cisplatin treatment. -

The Dark Side of UV-Induced DNA Lesion Repair
G C A T T A C G G C A T genes Review The Dark Side of UV-Induced DNA Lesion Repair Wojciech Strzałka 1, Piotr Zgłobicki 1, Ewa Kowalska 1, Aneta Ba˙zant 1, Dariusz Dziga 2 and Agnieszka Katarzyna Bana´s 1,* 1 Department of Plant Biotechnology, Faculty of Biochemistry, Biophysics and Biotechnology, Jagiellonian University, Gronostajowa 7, 30-387 Krakow, Poland; [email protected] (W.S.); [email protected] (P.Z.); [email protected] (E.K.); [email protected] (A.B.) 2 Department of Microbiology, Faculty of Biochemistry, Biophysics and Biotechnology, Jagiellonian University, Gronostajowa 7, 30-387 Krakow, Poland; [email protected] * Correspondence: [email protected]; Tel.: +48-12-664-6410 Received: 27 October 2020; Accepted: 29 November 2020; Published: 2 December 2020 Abstract: In their life cycle, plants are exposed to various unfavorable environmental factors including ultraviolet (UV) radiation emitted by the Sun. UV-A and UV-B, which are partially absorbed by the ozone layer, reach the surface of the Earth causing harmful effects among the others on plant genetic material. The energy of UV light is sufficient to induce mutations in DNA. Some examples of DNA damage induced by UV are pyrimidine dimers, oxidized nucleotides as well as single and double-strand breaks. When exposed to light, plants can repair major UV-induced DNA lesions, i.e., pyrimidine dimers using photoreactivation. However, this highly efficient light-dependent DNA repair system is ineffective in dim light or at night. Moreover, it is helpless when it comes to the repair of DNA lesions other than pyrimidine dimers. -

UV-DDB) Dimerization and Its Roles in Chromatinized DNA Repair
Damaged DNA induced UV-damaged DNA-binding PNAS PLUS protein (UV-DDB) dimerization and its roles in chromatinized DNA repair Joanne I. Yeha,b,1, Arthur S. Levinec,d, Shoucheng Dua, Unmesh Chintea, Harshad Ghodkee, Hong Wangd,e, Haibin Shia, Ching L. Hsiehc,d, James F. Conwaya, Bennett Van Houtend,e, and Vesna Rapić-Otrinc,d aDepartments of Structural Biology, bBioengineering, cMicrobiology and Molecular Genetics, ePharmacology and Chemical Biology, and dUniversity of Pittsburgh Cancer Institute, University of Pittsburgh School of Medicine, Pittsburgh, PA 15260 Edited by Lorena S. Beese, Duke University School of Medicine, Durham, NC, and approved January 17, 2012 (received for review June 24, 2011) UV light-induced photoproducts are recognized and removed by The ubiquitination pathway has recently been shown to play an the nucleotide-excision repair (NER) pathway. In humans, the important regulatory function in the initiation of NER (13, 14). UV-damaged DNA-binding protein (UV-DDB) is part of a ubiquitin The DDB1 protein is part of the substrate-recruiting module for E3 ligase complex (DDB1-CUL4ADDB2) that initiates NER by recog- two closely related types of E3 ligases, the cullins CUL4A and nizing damaged chromatin with concomitant ubiquitination of CUL4B, which target proteins for ubiquitination (15, 16). The core histones at the lesion. We report the X-ray crystal structure DDB1-CUL4A complex belongs to a superfamily of cullin-RING of the human UV-DDB in a complex with damaged DNA and show ligases (CRL) (17–19), which participate in various aspects of the that the N-terminal domain of DDB2 makes critical contacts with UV-damage response for maintaining genome stability (20–22).