Rabbit Anti-PAMCI/RASF9/FITC Conjugated Antibody
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Protein Identities in Evs Isolated from U87-MG GBM Cells As Determined by NG LC-MS/MS
Protein identities in EVs isolated from U87-MG GBM cells as determined by NG LC-MS/MS. No. Accession Description Σ Coverage Σ# Proteins Σ# Unique Peptides Σ# Peptides Σ# PSMs # AAs MW [kDa] calc. pI 1 A8MS94 Putative golgin subfamily A member 2-like protein 5 OS=Homo sapiens PE=5 SV=2 - [GG2L5_HUMAN] 100 1 1 7 88 110 12,03704523 5,681152344 2 P60660 Myosin light polypeptide 6 OS=Homo sapiens GN=MYL6 PE=1 SV=2 - [MYL6_HUMAN] 100 3 5 17 173 151 16,91913397 4,652832031 3 Q6ZYL4 General transcription factor IIH subunit 5 OS=Homo sapiens GN=GTF2H5 PE=1 SV=1 - [TF2H5_HUMAN] 98,59 1 1 4 13 71 8,048185945 4,652832031 4 P60709 Actin, cytoplasmic 1 OS=Homo sapiens GN=ACTB PE=1 SV=1 - [ACTB_HUMAN] 97,6 5 5 35 917 375 41,70973209 5,478027344 5 P13489 Ribonuclease inhibitor OS=Homo sapiens GN=RNH1 PE=1 SV=2 - [RINI_HUMAN] 96,75 1 12 37 173 461 49,94108966 4,817871094 6 P09382 Galectin-1 OS=Homo sapiens GN=LGALS1 PE=1 SV=2 - [LEG1_HUMAN] 96,3 1 7 14 283 135 14,70620005 5,503417969 7 P60174 Triosephosphate isomerase OS=Homo sapiens GN=TPI1 PE=1 SV=3 - [TPIS_HUMAN] 95,1 3 16 25 375 286 30,77169764 5,922363281 8 P04406 Glyceraldehyde-3-phosphate dehydrogenase OS=Homo sapiens GN=GAPDH PE=1 SV=3 - [G3P_HUMAN] 94,63 2 13 31 509 335 36,03039959 8,455566406 9 Q15185 Prostaglandin E synthase 3 OS=Homo sapiens GN=PTGES3 PE=1 SV=1 - [TEBP_HUMAN] 93,13 1 5 12 74 160 18,68541938 4,538574219 10 P09417 Dihydropteridine reductase OS=Homo sapiens GN=QDPR PE=1 SV=2 - [DHPR_HUMAN] 93,03 1 1 17 69 244 25,77302971 7,371582031 11 P01911 HLA class II histocompatibility antigen, -
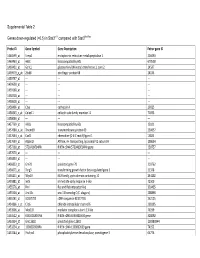
0.5) in Stat3∆/∆ Compared with Stat3flox/Flox
Supplemental Table 2 Genes down-regulated (<0.5) in Stat3∆/∆ compared with Stat3flox/flox Probe ID Gene Symbol Gene Description Entrez gene ID 1460599_at Ermp1 endoplasmic reticulum metallopeptidase 1 226090 1460463_at H60c histocompatibility 60c 670558 1460431_at Gcnt1 glucosaminyl (N-acetyl) transferase 1, core 2 14537 1459979_x_at Zfp68 zinc finger protein 68 24135 1459747_at --- --- --- 1459608_at --- --- --- 1459168_at --- --- --- 1458718_at --- --- --- 1458618_at --- --- --- 1458466_at Ctsa cathepsin A 19025 1458345_s_at Colec11 collectin sub-family member 11 71693 1458046_at --- --- --- 1457769_at H60a histocompatibility 60a 15101 1457680_a_at Tmem69 transmembrane protein 69 230657 1457644_s_at Cxcl1 chemokine (C-X-C motif) ligand 1 14825 1457639_at Atp6v1h ATPase, H+ transporting, lysosomal V1 subunit H 108664 1457260_at 5730409E04Rik RIKEN cDNA 5730409E04Rik gene 230757 1457070_at --- --- --- 1456893_at --- --- --- 1456823_at Gm70 predicted gene 70 210762 1456671_at Tbrg3 transforming growth factor beta regulated gene 3 21378 1456211_at Nlrp10 NLR family, pyrin domain containing 10 244202 1455881_at Ier5l immediate early response 5-like 72500 1455576_at Rinl Ras and Rab interactor-like 320435 1455304_at Unc13c unc-13 homolog C (C. elegans) 208898 1455241_at BC037703 cDNA sequence BC037703 242125 1454866_s_at Clic6 chloride intracellular channel 6 209195 1453906_at Med13l mediator complex subunit 13-like 76199 1453522_at 6530401N04Rik RIKEN cDNA 6530401N04 gene 328092 1453354_at Gm11602 predicted gene 11602 100380944 1453234_at -

Molecular Cancer Biomed Central
Molecular Cancer BioMed Central Research Open Access The novel RASSF6 and RASSF10 candidate tumour suppressor genes are frequently epigenetically inactivated in childhood leukaemias Luke B Hesson1, Thomas L Dunwell1, Wendy N Cooper1, Daniel Catchpoole2, Anna T Brini3, Raffaella Chiaramonte4, Mike Griffiths5, Andrew D Chalmers6, Eamonn R Maher1,5 and Farida Latif*1 Address: 1Department of Medical and Molecular Genetics, Department of Reproductive and Child Health, Institute of Biomedical Research, Medical School, University of Birmingham, Edgbaston, B15 2TT, UK, 2The Children's Hospital at Westmead, Locked Bay 4001, Westmead, NSW, 2145, Australia, 3Department of Medical Pharmacology, Faculty of Medicine, Università degli Studi di Milano, Italy, 4Department of Medicine, Surgery and Dentistry, Università degli Studi di Milano, via Di Rudinì 8, 20142 Milan, Italy, 5West Midlands Regional Genetics Service, Birmingham Women's Hospital, Edgbaston, Birmingham, B15 2TG, UK and 6Centre for Regenerative Medicine, Department of Biology and Biochemistry, University of Bath, Bath BA2 7AY, UK Email: Luke B Hesson - [email protected]; Thomas L Dunwell - [email protected]; Wendy N Cooper - [email protected]; Daniel Catchpoole - [email protected]; Anna T Brini - [email protected]; Raffaella Chiaramonte - [email protected]; Mike Griffiths - [email protected]; Andrew D Chalmers - [email protected]; Eamonn R Maher - [email protected]; Farida Latif* - [email protected] * Corresponding author Published: 1 July 2009 Received: 20 April 2009 Accepted: 1 July 2009 Molecular Cancer 2009, 8:42 doi:10.1186/1476-4598-8-42 This article is available from: http://www.molecular-cancer.com/content/8/1/42 © 2009 Hesson et al; licensee BioMed Central Ltd. -

Inhibition of Mitochondrial Complex II in Neuronal Cells Triggers Unique
www.nature.com/scientificreports OPEN Inhibition of mitochondrial complex II in neuronal cells triggers unique pathways culminating in autophagy with implications for neurodegeneration Sathyanarayanan Ranganayaki1, Neema Jamshidi2, Mohamad Aiyaz3, Santhosh‑Kumar Rashmi4, Narayanappa Gayathri4, Pulleri Kandi Harsha5, Balasundaram Padmanabhan6 & Muchukunte Mukunda Srinivas Bharath7* Mitochondrial dysfunction and neurodegeneration underlie movement disorders such as Parkinson’s disease, Huntington’s disease and Manganism among others. As a corollary, inhibition of mitochondrial complex I (CI) and complex II (CII) by toxins 1‑methyl‑4‑phenylpyridinium (MPP+) and 3‑nitropropionic acid (3‑NPA) respectively, induced degenerative changes noted in such neurodegenerative diseases. We aimed to unravel the down‑stream pathways associated with CII inhibition and compared with CI inhibition and the Manganese (Mn) neurotoxicity. Genome‑wide transcriptomics of N27 neuronal cells exposed to 3‑NPA, compared with MPP+ and Mn revealed varied transcriptomic profle. Along with mitochondrial and synaptic pathways, Autophagy was the predominant pathway diferentially regulated in the 3‑NPA model with implications for neuronal survival. This pathway was unique to 3‑NPA, as substantiated by in silico modelling of the three toxins. Morphological and biochemical validation of autophagy markers in the cell model of 3‑NPA revealed incomplete autophagy mediated by mechanistic Target of Rapamycin Complex 2 (mTORC2) pathway. Interestingly, Brain Derived Neurotrophic Factor -

Identification of Key Pathways and Genes in Dementia Via Integrated Bioinformatics Analysis
bioRxiv preprint doi: https://doi.org/10.1101/2021.04.18.440371; this version posted July 19, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Identification of Key Pathways and Genes in Dementia via Integrated Bioinformatics Analysis Basavaraj Vastrad1, Chanabasayya Vastrad*2 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karnataka, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India bioRxiv preprint doi: https://doi.org/10.1101/2021.04.18.440371; this version posted July 19, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract To provide a better understanding of dementia at the molecular level, this study aimed to identify the genes and key pathways associated with dementia by using integrated bioinformatics analysis. Based on the expression profiling by high throughput sequencing dataset GSE153960 derived from the Gene Expression Omnibus (GEO), the differentially expressed genes (DEGs) between patients with dementia and healthy controls were identified. With DEGs, we performed a series of functional enrichment analyses. Then, a protein–protein interaction (PPI) network, modules, miRNA-hub gene regulatory network and TF-hub gene regulatory network was constructed, analyzed and visualized, with which the hub genes miRNAs and TFs nodes were screened out. Finally, validation of hub genes was performed by using receiver operating characteristic curve (ROC) analysis. -

University of Bath Research Portal
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by University of Bath Research Portal Citation for published version: Sherwood, V, Recino, A, Jeffries, A, Ward, A & Chalmers, AD 2010, 'The N-terminal RASSF family: a new group of Ras-association-domain-containing proteins, with emerging links to cancer formation', Biochemical Journal, vol. 425, no. 2, pp. 303-311. https://doi.org/10.1042/BJ20091318 DOI: 10.1042/BJ20091318 Publication date: 2010 Link to publication The final version of record is available at http://www.biochemj.org/bj/default.htm University of Bath General rights Copyright and moral rights for the publications made accessible in the public portal are retained by the authors and/or other copyright owners and it is a condition of accessing publications that users recognise and abide by the legal requirements associated with these rights. Take down policy If you believe that this document breaches copyright please contact us providing details, and we will remove access to the work immediately and investigate your claim. Download date: 12. May. 2019 The N-terminal RASSF family; A new group of Ras association domain containing proteins, with emerging links to cancer formation. Victoria Sherwood *†, Asha Recino*, Alex Jeffries*, Andrew Ward*, Andrew D Chalmers*1. *, Centre for Regenerative Medicine, Department of Biology and Biochemistry, University of Bath, Bath, BA2 7AY, UK. †, Present address, Cell and Experimental Pathology, Lund University, Malmö University Hospital, S-205 02 MALMÖ, Sweden. 1, to whom correspondence should be addressed (e mail [email protected]). Running title: The N-terminal RASSF family and cancer. -

Supporting Table 1. Genes Differentially Expressed in Heparg Cells by the Coculture Condition
Supporting Table 1. Genes Differentially Expressed in HepaRG Cells by the Coculture Condition List of Up-Regulated Genes Gene symbol Description GB acc EntrezID p-value FDR HepaRG HepaRG/LX2 Fold-change CYP1A1 cytochrome P450, family 1, subfamily A, polypeptide 1 NM_000499 1543 0.0000021 0.00591 0.54 8.55 15.87 ENO2 enolase 2 NM_001975 2026 0.0003024 0.0243 0.049 0.32 6.67 C15orf48 chromosome 15 open reading frame 48 NM_032413 84419 0.0000131 0.00851 1.28 6.97 5.56 WISP2 WNT1 inducible signaling pathway protein 2 NM_003881 8839 0.0000395 0.0113 1.94 10.56 5.56 COL13A1 collagen, type XIII, alpha 1 NM_080801 1305 0.0000713 0.0135 0.58 2.55 4.35 C4orf47 chromosome 4 open reading frame 47 NM_001114357 441054 0.000041 0.0113 1.23 5.01 4.00 ZP1 zona pellucida glycoprotein 1 NM_207341 22917 0.0000433 0.0114 0.2 0.8 4.00 DENND2A DENN/MADD domain containing 2A NM_015689 27147 0.0000763 0.0138 0.19 0.73 3.85 SAA2 serum amyloid A2 NM_001127380 6289 0.0002017 0.0204 0.91 3.45 3.85 CP ceruloplasmin NM_000096 1356 0.0000028 0.00591 1.07 3.97 3.70 TXNIP thioredoxin interacting protein NM_006472 10628 0.0001031 0.0155 0.1 0.37 3.57 TEK TEK tyrosine kinase, endothelial NM_000459 7010 0.0000543 0.0119 0.14 0.48 3.45 C3P1 complement component 3 precursor pseudogene NR_027300 388503 0.0003026 0.0243 0.63 2.06 3.33 IL1RL1 interleukin 1 receptor-like 1 NM_016232 9173 0.0000345 0.0111 1.15 3.65 3.13 NDRG1 N-myc downstream regulated 1 NM_006096 10397 0.0001186 0.0164 1.3 4.07 3.13 IL6 interleukin 6 NM_000600 3569 0.0001052 0.0157 0.45 1.36 3.03 S100A8 S100 calcium -

A Novel Role of RASSF9 in Maintaining Epidermal Homeostasis
A Novel Role of RASSF9 in Maintaining Epidermal Homeostasis Chiou-Mei Lee1,2*, Polung Yang3, Lih-Chyang Chen2, Chia-Chun Chen3, Shinn-Chih Wu4, Hsiao-Yun Cheng3, Yu-Sun Chang2,3 1 Department of Medical Research and Development, Chang Gung Memorial Hospital at Lin-Kou, Taoyuan, Taiwan, 2 Graduate Institutes of Biomedical Sciences, Chang Gung University, Taoyuan, Taiwan, 3 Chang Gung Molecular Medicine Research Center, Chang Gung University, Taoyuan, Taiwan, 4 Department of Animal Science and Technology, National Taiwan University, Taipei, Taiwan Abstract The physiological role of RASSF9, a member of the Ras-association domain family (RASSF), is currently unclear. Here, we report a mouse line in which an Epstein-Barr virus Latent Membrane Protein 1 (LMP1) transgene insertion has created a 7.2- kb chromosomal deletion, which abolished RASSF9 gene expression. The RASSF9-null mice exhibited interesting phenotypes that resembled human ageing, including growth retardation, short lifespan, less subcutaneous adipose layer and alopecia. In the wild-type mice, RASSF9 is predominantly expressed in the epidermal keratinocytes of skin, as determined by quantitative reverse-transcription PCR, immunofluorescence and in situ hybridization. In contrast, RASSF92/2 mice presented a dramatic change in epithelial organization of skin with increased proliferation and aberrant differentiation as detected by bromodeoxyuridine incorporation assays and immunofluorescence analyses. Furthermore, characteristic functions of RASSF92/2 versus wild type (WT) mouse primary keratinocytes showed significant proliferation linked to a reduction of p21Cip1 expression under growth or early differentiation conditions. Additionally, in RASSF92/2 keratinocytes there was a drastic down-modulation of terminal differentiation markers, which could be rescued by infection with a recombinant adenovirus, Adv/HA-RASSF9. -
In BRCA1 and BRCA2 Breast Cancers, Chromosome Breaks Occur Near Herpes Tumor Virus Sequences
Preprints (www.preprints.org) | NOT PEER-REVIEWED | Posted: 20 May 2021 doi:10.20944/preprints202105.0490.v1 Article In BRCA1 and BRCA2 breast cancers, chromosome breaks occur near herpes tumor virus sequences Bernard Friedenson 1* 1 Dept. of Biochemistry and Molecular Genetics, College of Medicine University of Illinois Chicago; [email protected] * Correspondence: [email protected]; Abstract: Inherited mutations in BRCA1 and BRCA2 genes increase risks for breast, ovarian, and other cancers. Both genes encode proteins for accurately repairing chromosome breaks. If mutations inactivate this function, chromosome fragments may not be restored correctly. Resulting chromosome rearrangements can become critical breast cancer drivers. Because I had data from thousands of cancer structural alterations that matched viral infections, I wondered whether infections contribute to chromosome breaks and rearrangements in hereditary breast cancers. There are currently no interventions to prevent chromosome breaks because they are thought to be unavoidable. However, if chromosome breaks come from infections, they can be treated or prevented. I used bioinformatic analyses to test publicly available breast cancer sequence data around chromosome breaks for DNA similarity to all known viruses. Human DNA flanking breakpoints usually had the strongest matches to Epstein-Barr virus (EBV) tumor variants HKHD40 and HKNPC60. Many breakpoints were near sites that anchor EBV genomes, human EBV tumor-like sequences, EBV-associated epigenetic marks, and fragile sites. On chromosome 2, sequences near EBV genome anchor sites accounted for 90% of breakpoints (p<0.0001). On chromosome 4, 51/52 inter-chromosomal breakpoints were close to EBV-like sequences. Five EBV genome anchor sites were near breast cancer breakpoints at precisely defined, disparate gene or LINE locations. -
Regulation of the Retinoblastoma Tumor Suppressor by the Novel Ras Effector NORE1A. Thibaut François Barnoud University of Louisville
University of Louisville ThinkIR: The University of Louisville's Institutional Repository Electronic Theses and Dissertations 12-2015 Regulation of the retinoblastoma tumor suppressor by the novel Ras effector NORE1A. Thibaut François Barnoud University of Louisville Follow this and additional works at: https://ir.library.louisville.edu/etd Part of the Cancer Biology Commons Recommended Citation Barnoud, Thibaut François, "Regulation of the retinoblastoma tumor suppressor by the novel Ras effector NORE1A." (2015). Electronic Theses and Dissertations. Paper 2320. https://doi.org/10.18297/etd/2320 This Doctoral Dissertation is brought to you for free and open access by ThinkIR: The nivU ersity of Louisville's Institutional Repository. It has been accepted for inclusion in Electronic Theses and Dissertations by an authorized administrator of ThinkIR: The nivU ersity of Louisville's Institutional Repository. This title appears here courtesy of the author, who has retained all other copyrights. For more information, please contact [email protected]. REGULATION OF THE RETINOBLASTOMA TUMOR SUPPRESSOR BY THE NOVEL RAS EFFECTOR NORE1A By Thibaut François Barnoud B.A., University of Louisville, 2009 M.S., University of Louisville, 2013 A Dissertation Submitted to the Faculty of the School of Medicine of the University of Louisville in Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy in Biochemistry and Molecular Genetics Department of Biochemistry and Molecular Genetics University of Louisville Louisville, KY December, 2015 Copyright 2015 by Thibaut François Barnoud All Rights Reserved REGULATION OF THE RETINOBLASTOMA TUMOR SUPPRESSOR BY THE NOVEL RAS EFFECTOR NORE1A By Thibaut François Barnoud B.A., University of Louisville, 2009 M.S., University of Louisville, 2013 A Dissertation Approved on November 9, 2015 by the following Dissertation Committee: ________________________________________ Dr. -

Network Mining Approach to Cancer Biomarker Discovery
NETWORK MINING APPROACH TO CANCER BIOMARKER DISCOVERY THESIS Presented in Partial Fulfillment of the Requirements for the Degree Master of Science in the Graduate School of The Ohio State University By Praneeth Uppalapati, B.E. Graduate Program in Computer Science and Engineering The Ohio State University 2010 Thesis Committee: Dr. Kun Huang, Advisor Dr. Raghu Machiraju Copyright by Praneeth Uppalapati 2010 ABSTRACT With the rapid development of high throughput gene expression profiling technology, molecule profiling has become a powerful tool to characterize disease subtypes and discover gene signatures. Most existing gene signature discovery methods apply statistical methods to select genes whose expression values can differentiate different subject groups. However, a drawback of these approaches is that the selected genes are not functionally related and hence cannot reveal biological mechanism behind the difference in the patient groups. Gene co-expression network analysis can be used to mine functionally related sets of genes that can be marked as potential biomarkers through survival analysis. We present an efficient heuristic algorithm EigenCut that exploits the properties of gene co- expression networks to mine functionally related and dense modules of genes. We apply this method to brain tumor (Glioblastoma Multiforme) study to obtain functionally related clusters. If functional groups of genes with predictive power on patient prognosis can be identified, insights on the mechanisms related to metastasis in GBM can be obtained and better therapeutical plan can be developed. We predicted potential biomarkers by dividing the patients into two groups based on their expression profiles over the genes in the clusters and comparing their survival outcome through survival analysis.