Identification of the Mrna-Associated Top3β- TDRD3-FMRP (TTF)
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

2020 Program Book
PROGRAM BOOK Note that TAGC was cancelled and held online with a different schedule and program. This document serves as a record of the original program designed for the in-person meeting. April 22–26, 2020 Gaylord National Resort & Convention Center Metro Washington, DC TABLE OF CONTENTS About the GSA ........................................................................................................................................................ 3 Conference Organizers ...........................................................................................................................................4 General Information ...............................................................................................................................................7 Mobile App ....................................................................................................................................................7 Registration, Badges, and Pre-ordered T-shirts .............................................................................................7 Oral Presenters: Speaker Ready Room - Camellia 4.......................................................................................7 Poster Sessions and Exhibits - Prince George’s Exhibition Hall ......................................................................7 GSA Central - Booth 520 ................................................................................................................................8 Internet Access ..............................................................................................................................................8 -

Arginine Methylation Facilitates the Recruitment of TOP3B to Chromatin to Prevent R Loop Accumulation
Molecular Cell Article Arginine Methylation Facilitates the Recruitment of TOP3B to Chromatin to Prevent R Loop Accumulation Yanzhong Yang,1 Kevin M. McBride,1 Sean Hensley,1 Yue Lu,1 Frederic Chedin,2 and Mark T. Bedford1,* 1The University of Texas MD Anderson Cancer Center, P.O. Box 389, Smithville, TX 78957, USA 2Department of Molecular & Cellular Biology, The University of California at Davis, Davis, CA 95616, USA *Correspondence: [email protected] http://dx.doi.org/10.1016/j.molcel.2014.01.011 SUMMARY nine methyltransferase 1), which deposit the H4R3me2a and H3R17me2a marks, respectively (Yang and Bedford, 2013). Tudor domain-containing protein 3 (TDRD3) is a Both of these marks are recognized by the Tudor domain of major methylarginine effector molecule that reads TDRD3 (Yang et al., 2010), a protein that is enriched at the pro- methyl-histone marks and facilitates gene transcrip- moters of highly transcribed genes and can likely also associate tion. However, the underlying mechanism by which with the C-terminal domain (CTD) of RNA Polymerase II (RNAP II) TDRD3 functions as a transcriptional coactivator is (Sims et al., 2011). TDRD3 has no enzymatic activity of its unknown. We identified topoisomerase IIIB (TOP3B) own, but here we show that it is tightly complexed with DNA topoisomerase IIIb (TOP3B), an interaction that bestows, in as a component of the TDRD3 complex. TDRD3 part, coactivator activity on TDRD3. TOP3B is a member of serves as a molecular bridge between TOP3B and the 1A subfamily of DNA topoisomerases and, as such, targets arginine-methylated histones. The TDRD3-TOP3B underwound or negatively supercoiled DNA (Wang, 2002). -

MAFB Determines Human Macrophage Anti-Inflammatory
MAFB Determines Human Macrophage Anti-Inflammatory Polarization: Relevance for the Pathogenic Mechanisms Operating in Multicentric Carpotarsal Osteolysis This information is current as of October 4, 2021. Víctor D. Cuevas, Laura Anta, Rafael Samaniego, Emmanuel Orta-Zavalza, Juan Vladimir de la Rosa, Geneviève Baujat, Ángeles Domínguez-Soto, Paloma Sánchez-Mateos, María M. Escribese, Antonio Castrillo, Valérie Cormier-Daire, Miguel A. Vega and Ángel L. Corbí Downloaded from J Immunol 2017; 198:2070-2081; Prepublished online 16 January 2017; doi: 10.4049/jimmunol.1601667 http://www.jimmunol.org/content/198/5/2070 http://www.jimmunol.org/ Supplementary http://www.jimmunol.org/content/suppl/2017/01/15/jimmunol.160166 Material 7.DCSupplemental References This article cites 69 articles, 22 of which you can access for free at: http://www.jimmunol.org/content/198/5/2070.full#ref-list-1 by guest on October 4, 2021 Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2017 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. -

Transcriptional Profile of Human Anti-Inflamatory Macrophages Under Homeostatic, Activating and Pathological Conditions
UNIVERSIDAD COMPLUTENSE DE MADRID FACULTAD DE CIENCIAS QUÍMICAS Departamento de Bioquímica y Biología Molecular I TESIS DOCTORAL Transcriptional profile of human anti-inflamatory macrophages under homeostatic, activating and pathological conditions Perfil transcripcional de macrófagos antiinflamatorios humanos en condiciones de homeostasis, activación y patológicas MEMORIA PARA OPTAR AL GRADO DE DOCTOR PRESENTADA POR Víctor Delgado Cuevas Directores María Marta Escribese Alonso Ángel Luís Corbí López Madrid, 2017 © Víctor Delgado Cuevas, 2016 Universidad Complutense de Madrid Facultad de Ciencias Químicas Dpto. de Bioquímica y Biología Molecular I TRANSCRIPTIONAL PROFILE OF HUMAN ANTI-INFLAMMATORY MACROPHAGES UNDER HOMEOSTATIC, ACTIVATING AND PATHOLOGICAL CONDITIONS Perfil transcripcional de macrófagos antiinflamatorios humanos en condiciones de homeostasis, activación y patológicas. Víctor Delgado Cuevas Tesis Doctoral Madrid 2016 Universidad Complutense de Madrid Facultad de Ciencias Químicas Dpto. de Bioquímica y Biología Molecular I TRANSCRIPTIONAL PROFILE OF HUMAN ANTI-INFLAMMATORY MACROPHAGES UNDER HOMEOSTATIC, ACTIVATING AND PATHOLOGICAL CONDITIONS Perfil transcripcional de macrófagos antiinflamatorios humanos en condiciones de homeostasis, activación y patológicas. Este trabajo ha sido realizado por Víctor Delgado Cuevas para optar al grado de Doctor en el Centro de Investigaciones Biológicas de Madrid (CSIC), bajo la dirección de la Dra. María Marta Escribese Alonso y el Dr. Ángel Luís Corbí López Fdo. Dra. María Marta Escribese -
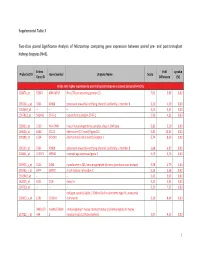
Supplemental Table 3 Two-Class Paired Significance Analysis of Microarrays Comparing Gene Expression Between Paired
Supplemental Table 3 Two‐class paired Significance Analysis of Microarrays comparing gene expression between paired pre‐ and post‐transplant kidneys biopsies (N=8). Entrez Fold q‐value Probe Set ID Gene Symbol Unigene Name Score Gene ID Difference (%) Probe sets higher expressed in post‐transplant biopsies in paired analysis (N=1871) 218870_at 55843 ARHGAP15 Rho GTPase activating protein 15 7,01 3,99 0,00 205304_s_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 6,30 4,50 0,00 1563649_at ‐‐ ‐‐ ‐‐ 6,24 3,51 0,00 1567913_at 541466 CT45‐1 cancer/testis antigen CT45‐1 5,90 4,21 0,00 203932_at 3109 HLA‐DMB major histocompatibility complex, class II, DM beta 5,83 3,20 0,00 204606_at 6366 CCL21 chemokine (C‐C motif) ligand 21 5,82 10,42 0,00 205898_at 1524 CX3CR1 chemokine (C‐X3‐C motif) receptor 1 5,74 8,50 0,00 205303_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 5,68 6,87 0,00 226841_at 219972 MPEG1 macrophage expressed gene 1 5,59 3,76 0,00 203923_s_at 1536 CYBB cytochrome b‐245, beta polypeptide (chronic granulomatous disease) 5,58 4,70 0,00 210135_s_at 6474 SHOX2 short stature homeobox 2 5,53 5,58 0,00 1562642_at ‐‐ ‐‐ ‐‐ 5,42 5,03 0,00 242605_at 1634 DCN decorin 5,23 3,92 0,00 228750_at ‐‐ ‐‐ ‐‐ 5,21 7,22 0,00 collagen, type III, alpha 1 (Ehlers‐Danlos syndrome type IV, autosomal 201852_x_at 1281 COL3A1 dominant) 5,10 8,46 0,00 3493///3 IGHA1///IGHA immunoglobulin heavy constant alpha 1///immunoglobulin heavy 217022_s_at 494 2 constant alpha 2 (A2m marker) 5,07 9,53 0,00 1 202311_s_at -

Textual Description of Annaffy
Textual Description of annaffy Colin A. Smith June 20, 2021 Introduction annaffy is part of the Bioconductor project. It is designed to help interface between Affymetrix analysis results and web-based databases. It provides classes and functions for accessing those resources both interactively and through statically generated HTML pages. The core functionality of annaffy depends on annotation contained in Bioconductor data packages. The data packages are created by the SQLForge code inside of another package called AnnotationDbi. It gathers annotation data from many diverse sources and makes the information easily processed by R. Preconstructed packages for most Affymetrix chips are available on the Bioconductor web site. 1 Loading Annotation Data annaffy represents each type of annotation data as a different class. Currently imple- mented classes include: aafSymbol gene symbol aafDescription gene description/name aafFunction gene functional description aafChromosome genomic chromosome aafChromLoc location on the chromosome (in base pairs) aafGenBank GenBank accession number aafLocusLink LocusLink ids (almost never more than one) aafCytoband mapped cytoband location 1 aafUniGene UniGene cluster ids (almost never more than one) aafPubMed PubMed ids aafGO Gene Ontology identifiers, names, types, and evidence codes aafPathway KEGG pathway identifiers and names For each class, there is a constructor function with the same name. It takes as arguments a vector of Affymetrix probe ids as well as the chip name. The chip name corresponds to the name of the data package that contains the annotation. If the data package for the chip is not already loaded, the constructor will attempt to load it. The constructor returns a list of the corresponding objects populated with annotation data. -

Functional Characterization of the TTF Complex and Its Role in Neurodevelopmental Disorders
Functional characterization of the TTF complex and its role in neurodevelopmental disorders Doctoral Dissertation (Thesis) to obtain the degree of “Doktor der Naturwissenschaften” (Dr. rer. nat.) awarded by the Julius-Maximilians-Universität Würzburg submitted by Cornelia Brosi from Würzburg, Germany Würzburg 2017 Submitted to the Faculty of Chemistry and Pharmacy on December 21, 2017 Evaluators of the written Dissertation (Thesis) Supervisor – 1. Evaluator: Prof. Dr. Utz Fischer 2. Evaluator: Prof. Dr. Alexander Buchberger Examiners of the Public Defense 1. Examiner: Prof. Dr. Utz Fischer 2. Examiner: Prof. Dr. Alexander Buchberger 3. Examiner: PD Dr. Sibylle Jablonka Date of the Public Defense Doctoral Certificate awarded on Table of contents 1. Summary.......................................................................................................................................... 1 2. Zusammenfassung ........................................................................................................................... 3 3. Introduction ..................................................................................................................................... 5 3.1. mRNA processing - an important step in the eukaryotic gene expression .................................. 5 3.2. The mRNP code regulates the post-transcriptional gene expression .......................................... 7 3.3. mRNA-binding proteins ............................................................................................................. -
Sheet1 Page 1 Gene Symbol Gene Description Entrez Gene ID
Sheet1 RefSeq ID ProbeSets Gene Symbol Gene Description Entrez Gene ID Sequence annotation Seed matches location(s) Ago-2 binding specific enrichment (replicate 1) Ago-2 binding specific enrichment (replicate 2) OE lysate log2 fold change (replicate 1) OE lysate log2 fold change (replicate 2) Probability Pulled down in Karginov? NM_005646 202813_at TARBP1 Homo sapiens TAR (HIV-1) RNA binding protein 1 (TARBP1), mRNA. 6894 TR(1..5130)CDS(1..4866) 4868..4874,5006..5013 3.73 2.53 -1.54 -0.44 1 Yes NM_001665 203175_at RHOG Homo sapiens ras homolog gene family, member G (rho G) (RHOG), mRNA. 391 TR(1..1332)CDS(159..734) 810..817,782..788,790..796,873..879 3.56 2.78 -1.62 -1 1 Yes NM_002742 205880_at PRKD1 Homo sapiens protein kinase D1 (PRKD1), mRNA. 5587 TR(1..3679)CDS(182..2920) 3538..3544,3202..3208 4.15 1.83 -2.55 -0.42 1 Yes NM_003068 213139_at SNAI2 Homo sapiens snail homolog 2 (Drosophila) (SNAI2), mRNA. 6591 TR(1..2101)CDS(165..971) 1410..1417,1814..1820,1610..1616 3.5 2.79 -1.38 -0.31 1 Yes NM_006270 212647_at RRAS Homo sapiens related RAS viral (r-ras) oncogene homolog (RRAS), mRNA. 6237 TR(1..1013)CDS(46..702) 871..877 3.82 2.27 -1.54 -0.55 1 Yes NM_025188 219923_at,242056_at TRIM45 Homo sapiens tripartite motif-containing 45 (TRIM45), mRNA. 80263 TR(1..3584)CDS(589..2331) 3408..3414,2437..2444,3425..3431,2781..2787 3.87 1.89 -0.62 -0.09 1 Yes NM_024684 221600_s_at,221599_at C11orf67 Homo sapiens chromosome 11 open reading frame 67 (C11orf67), mRNA. -

TDRD3 (D3O2G) Rabbit Mab A
Revision 1 C 0 2 - t TDRD3 (D3O2G) Rabbit mAb a e r o t S Orders: 877-616-CELL (2355) [email protected] Support: 877-678-TECH (8324) 2 4 Web: [email protected] 9 www.cellsignal.com 5 # 3 Trask Lane Danvers Massachusetts 01923 USA For Research Use Only. Not For Use In Diagnostic Procedures. Applications: Reactivity: Sensitivity: MW (kDa): Source/Isotype: UniProt ID: Entrez-Gene Id: WB, IP H M R Mk Endogenous 83 Rabbit IgG Q9H7E2 81550 Product Usage Information Application Dilution Western Blotting 1:1000 Immunoprecipitation 1:50 Storage Supplied in 10 mM sodium HEPES (pH 7.5), 150 mM NaCl, 100 µg/ml BSA, 50% glycerol and less than 0.02% sodium azide. Store at –20°C. Do not aliquot the antibody. Specificity / Sensitivity TDRD3 (D3O2G) Rabbit mAb recognizes endogenous levels of total TDRD3 protein. Species Reactivity: Human, Mouse, Rat, Monkey Source / Purification Monoclonal antibody is produced by immunizing animals with a synthetic peptide corresponding to residues near the carboxy terminus of human TDRD3 protein. Background Tudor domain-containing protein 3 (TDRD3) contains a tudor domain through which it binds to asymmetric di-methyl histone H3 (Arg17) and asymmetric di-methyl histone H4 (Arg3). Both of these histone marks are associated with transcription activation (1,2). TDRD3 is targeted to estrogen responsive genes where it performs as a coactivator (1). TDRD3 acts as a scaffolding protein to recruit Topoisomerase III B (TOP3B) to target genes to reduce transcription-generated R loops by relaxing negatively supercoiled DNA thereby reducing genomic instability (3). In addition to the nucleus, TDRD3 also resides in the cytoplasm where it associates with actively translating polyribosomes and accumulates into stress granules upon exposure to stress, implicating its role in post- transcriptional regulation of RNA (2,4). -

Tdrd3 Regulates the Progression of Meiosis II Through Translational
bioRxiv preprint doi: https://doi.org/10.1101/2021.02.17.431574; this version posted February 17, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Tdrd3 regulates the progression of meiosis II through translational 2 control of Emi2 mRNA in mouse oocytes 3 4 Natsumi Takei1, Keisuke Sato1, Yuki Takada1, Rajan Iyyappan2, Andrej Susor2, Takehiro 5 Yamamoto3, and Tomoya Kotani1,4,* 6 7 1Biosystems Science Course, Graduate School of Life Science, Hokkaido University, 8 Sapporo 060-0810, Japan 9 2Laboratory of Biochemistry and Molecular Biology of Germ Cells, Institute of Animal 10 Physiology and Genetics, CAS, Rumburska 89, 277 21 Libechov, Czech Republic. 11 3Department of Biochemistry, Keio University School of Medicine, Tokyo 160-8582, 12 Japan 13 4Department of Biological Sciences, Faculty of Science, Hokkaido University, Sapporo 14 060-0810, Japan 15 16 17 *Corresponding author. 18 Tomoya Kotani, 19 Department of Biological Sciences, Faculty of Science, Hokkaido University, 20 North 10 West 8, Sapporo, Hokkaido 060-0810, Japan 21 Tel.: +81-11-706-4455 22 Fax.: +81-11-706-4455 23 E-mail: [email protected] 24 25 26 Running title: Tdrd3 regulates oocyte meiosis 27 Key words: oocyte, meiosis, Emi2, cyclin B1, RNA granule, translational control 28 1 bioRxiv preprint doi: https://doi.org/10.1101/2021.02.17.431574; this version posted February 17, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. -

TDRD3 (C-20): Sc-84626
SAN TA C RUZ BI OTEC HNOL OG Y, INC . TDRD3 (C-20): sc-84626 BACKGROUND APPLICATIONS TDRD3 (tudor domain containing 3) is a 651 amino acid protein that localizes TDRD3 (C-20) is recommended for detection of TDRD3 of mouse, rat and to both the nucleus and the cytoplasm and contains one UBA domain and one human origin by Western Blotting (starting dilution 1:200, dilution range tudor domain. Expressed in lung, brain, heart, liver, placenta, kidney, pancreas 1:100-1:1000), immunoprecipitation [1-2 µg per 100-500 µg of total protein and skeletal muscle, TDRD3 exists as a component of mRNA stress granules (1 ml of cell lysate)], immunofluorescence (starting dilution 1:50, dilution and is thought to play a role in the translation of target mRNAs, as well as in range 1:50-1:500), immunohistochemistry (including paraffin-embedded the assembly and disassembly of stress granules. Multiple isoforms of TDRD3 sec tions) (starting dilution 1:50, dilution range 1:50-1:500) and solid phase exist due to alternative splicing events. The gene encoding TDRD3 maps to ELISA (starting dilution 1:30, dilution range 1:30-1:3000). human chromosome 13, which houses over 400 genes, such as BRCA2 and TDRD3 (C-20) is also recommended for detection of TDRD3 in additional RB1, and comprises nearly 4% of the human genome. Trisomy 13, also known species, including equine, canine, bovine, porcine and avian. as Patau syndrome, is deadly and the few who survive past one year suffer from permanent neurologic defects, difficulty eating and vulnerability to seri - Suitable for use as control antibody for TDRD3 siRNA (h): sc-76639, TDRD3 ous respiratory infections. -

Structural Basis of the Interaction Between Topoisomerase Iiiβ And
www.nature.com/scientificreports OPEN Structural basis of the interaction between Topoisomerase IIIβ and the TDRD3 auxiliary factor Received: 30 September 2016 Sakurako Goto-Ito1,2, Atsushi Yamagata1,2,3, Tomio S. Takahashi1,2, Yusuke Sato1,2,3 & Accepted: 03 January 2017 Shuya Fukai1,2,3 Published: 08 February 2017 Topoisomerase IIIβ (TOP3β) is a DNA/RNA topoisomerase that has been implicated in epigenetic or translational control of gene expression. In cells, TOP3β co-exists with its specific auxiliary factor, TDRD3. TDRD3 serves as a scaffold protein to recruit TOP3β to its DNA/RNA substrates accumulating in specific cellular sites such as methylated chromatins or neural stress granules. Here we report the crystal structures of the catalytic domain of TOP3β, the DUF1767–OB-fold domains of TDRD3 and their complex at 3.44 Å, 1.62 Å and 3.6 Å resolutions, respectively. The toroidal-shaped catalytic domain of TOP3β binds the OB-fold domain of TDRD3. The TDRD3 OB-fold domain harbors the insertion loop, which is protruding from the core structure. Both the insertion loop and core region interact with TOP3β. Our pull-down binding assays showed that hydrophobic characters of the core surface and the amino- and carboxy-terminal regions of the insertion loop are essential for the interaction. Furthermore, by comparison with the structure of the homologous Topoisomerase IIIα (TOP3α)–RMI1 complex, we identified Arg96, Val109, Phe139 and the short insertion loop of TDRD3 as the critical structural elements for the specific interaction with TOP3β to avoid the non-cognate interaction with TOP3α. Expression or inheritance of genetic information (i.e., transcription, replication or recombination) is associated with unwinding of DNA duplexes, which causes their topological distortions that impede these genetic processes.