Interferon Alpha 6 (IFNA6) (NM 021002) Human Recombinant Protein Product Data
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Recombinant Human IFN-Α6 Mammalian Cell-Expressed Human Interferon Alpha 6 (Alpha K) with HSA Cat
Recombinant human IFN-α6 Mammalian cell-expressed human interferon alpha 6 (alpha K) with HSA Cat. code: rcyc-hifna6 https://www.invivogen.com/human-ifna6 For research use only, not for diagnostic or therapeutic use Version 18I12-NJ PRODUCT INFORMATION BACKGROUND Contents Type I interferons (IFN) include the IFN-α family, IFN-β, IFN-ε, IFN-κ • 1 µg of lyophilized recombinant human IFN-α6 and IFN-ω. IFN-αs are important anti-viral cytokines that also play Note: This product is sterile filtered prior to lyophilization. anti-proliferative and immuno-modulatory functions. The human IFN-α • 1.5 ml endotoxin-free water family comprises 13 genes encoding 12 proteins, IFN-α13 being identical to IFN-α1. All IFN-αs bind to a common heterodimer receptor IFNAR1/ Storage and stability IFNAR2. The ternary complex signals through the Janus kinase (JAK) • Recombinant human IFN-α6 is shipped at room temperature. Upon and signal transducer and activator of transcription (STAT) signaling receipt it should be immediately stored at -20 °C. Lyophilized product is stable for 6 months at -20°C. pathway, inducing the formation of the ISGF3 transcriptional complex • Upon resuspension, prepare aliquots of recombinant human IFN-α6 and (STAT1/STAT2/IRF9). ISGF3 binds to IFN-stimulated response elements 2 store at -80 °C for 4 months. (ISRE) in the promoters of numerous IFN-stimulated genes (ISGs) . Note: Avoid repeated freeze-thaw cycles. Quality control IFN-α IFN-α • Purity: ≥ 95% (SDS-PAGE) Low affinity High affinity • Endotoxin: ≤ 1 EU/µg • The biological activity has been confirmed using HEK-Blue™ IFN-α/β IFNAR1 IFNAR2 cells (see validation data sheet available on our website). -

Supplementary Material DNA Methylation in Inflammatory Pathways Modifies the Association Between BMI and Adult-Onset Non- Atopic
Supplementary Material DNA Methylation in Inflammatory Pathways Modifies the Association between BMI and Adult-Onset Non- Atopic Asthma Ayoung Jeong 1,2, Medea Imboden 1,2, Akram Ghantous 3, Alexei Novoloaca 3, Anne-Elie Carsin 4,5,6, Manolis Kogevinas 4,5,6, Christian Schindler 1,2, Gianfranco Lovison 7, Zdenko Herceg 3, Cyrille Cuenin 3, Roel Vermeulen 8, Deborah Jarvis 9, André F. S. Amaral 9, Florian Kronenberg 10, Paolo Vineis 11,12 and Nicole Probst-Hensch 1,2,* 1 Swiss Tropical and Public Health Institute, 4051 Basel, Switzerland; [email protected] (A.J.); [email protected] (M.I.); [email protected] (C.S.) 2 Department of Public Health, University of Basel, 4001 Basel, Switzerland 3 International Agency for Research on Cancer, 69372 Lyon, France; [email protected] (A.G.); [email protected] (A.N.); [email protected] (Z.H.); [email protected] (C.C.) 4 ISGlobal, Barcelona Institute for Global Health, 08003 Barcelona, Spain; [email protected] (A.-E.C.); [email protected] (M.K.) 5 Universitat Pompeu Fabra (UPF), 08002 Barcelona, Spain 6 CIBER Epidemiología y Salud Pública (CIBERESP), 08005 Barcelona, Spain 7 Department of Economics, Business and Statistics, University of Palermo, 90128 Palermo, Italy; [email protected] 8 Environmental Epidemiology Division, Utrecht University, Institute for Risk Assessment Sciences, 3584CM Utrecht, Netherlands; [email protected] 9 Population Health and Occupational Disease, National Heart and Lung Institute, Imperial College, SW3 6LR London, UK; [email protected] (D.J.); [email protected] (A.F.S.A.) 10 Division of Genetic Epidemiology, Medical University of Innsbruck, 6020 Innsbruck, Austria; [email protected] 11 MRC-PHE Centre for Environment and Health, School of Public Health, Imperial College London, W2 1PG London, UK; [email protected] 12 Italian Institute for Genomic Medicine (IIGM), 10126 Turin, Italy * Correspondence: [email protected]; Tel.: +41-61-284-8378 Int. -

And IRAK1 Knock-In Mice Production Defined by the Study of IRAK2 Two
Two Phases of Inflammatory Mediator Production Defined by the Study of IRAK2 and IRAK1 Knock-in Mice This information is current as Eduardo Pauls, Sambit K. Nanda, Hilary Smith, Rachel of September 29, 2021. Toth, J. Simon C. Arthur and Philip Cohen J Immunol 2013; 191:2717-2730; Prepublished online 5 August 2013; doi: 10.4049/jimmunol.1203268 http://www.jimmunol.org/content/191/5/2717 Downloaded from Supplementary http://www.jimmunol.org/content/suppl/2013/08/06/jimmunol.120326 Material 8.DC1 http://www.jimmunol.org/ References This article cites 55 articles, 33 of which you can access for free at: http://www.jimmunol.org/content/191/5/2717.full#ref-list-1 Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision by guest on September 29, 2021 • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts Errata An erratum has been published regarding this article. Please see next page or: /content/191/10/5317.full.pdf The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. -

Prevalent Homozygous Deletions of Type I Interferon and Defensin Genes in Human Cancers Associate with Immunotherapy Resistance
Author Manuscript Published OnlineFirst on April 4, 2018; DOI: 10.1158/1078-0432.CCR-17-3008 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. Prevalent Homozygous Deletions of Type I Interferon and Defensin Genes in Human Cancers Associate with Immunotherapy Resistance Zhenqing Ye1; Haidong Dong2,3; Ying Li1; Tao Ma1,4; Haojie Huang4; Hon-Sing Leong4; Jeanette Eckel-Passow1; Jean-Pierre A. Kocher1; Han Liang5; LiguoWang1,4,* 1Division of Biomedical Statistics and Informatics, Department of Health Sciences, Mayo Clinic, Rochester, Minnesota 55905, United States of America 2Department of Immunology, College of Medicine, Mayo Clinic, Rochester, Minnesota 55905, United States of America 3Department of Urology, Mayo Clinic, Rochester, Minnesota 55905, United States of America 4Department of Biochemistry and Molecular Biology, Mayo Clinic, Rochester, Minnesota 55905, United States of America 5Department of Bioinformatics and Computational Biology, The University of Texas MD Anderson Cancer Center, Houston, Texas 77030, United States of America Running title: Pervasive Deletion of Interferon/Defensin in Human Cancers Keywords: Homozygous deletion, Type-I interferon, Defensin, immunotherapy resistance, cancer * Correspondence to: Liguo Wang, Ph.D. Associate Professor Division of Biomedical Statistics and Informatics, Mayo Clinic 200 1st St SW Rochester, MN 55905, USA Phone: +1-507-284-8728 Fax: +1-507-284-0745 Email: [email protected] The authors declare no potential conflicts of interest. 1 Downloaded from clincancerres.aacrjournals.org on September 29, 2021. © 2018 American Association for Cancer Research. Author Manuscript Published OnlineFirst on April 4, 2018; DOI: 10.1158/1078-0432.CCR-17-3008 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. -

( 12 ) United States Patent
US010428349B2 (12 ) United States Patent ( 10 ) Patent No. : US 10 , 428 ,349 B2 DeRosa et al . (45 ) Date of Patent: Oct . 1 , 2019 ( 54 ) MULTIMERIC CODING NUCLEIC ACID C12N 2830 / 50 ; C12N 9 / 1018 ; A61K AND USES THEREOF 38 / 1816 ; A61K 38 /45 ; A61K 38/ 44 ; ( 71 ) Applicant: Translate Bio , Inc ., Lexington , MA A61K 38 / 177 ; A61K 48 /005 (US ) See application file for complete search history . (72 ) Inventors : Frank DeRosa , Lexington , MA (US ) ; Michael Heartlein , Lexington , MA (56 ) References Cited (US ) ; Daniel Crawford , Lexington , U . S . PATENT DOCUMENTS MA (US ) ; Shrirang Karve , Lexington , 5 , 705 , 385 A 1 / 1998 Bally et al. MA (US ) 5 ,976 , 567 A 11/ 1999 Wheeler ( 73 ) Assignee : Translate Bio , Inc ., Lexington , MA 5 , 981, 501 A 11/ 1999 Wheeler et al. 6 ,489 ,464 B1 12 /2002 Agrawal et al. (US ) 6 ,534 ,484 B13 / 2003 Wheeler et al. ( * ) Notice : Subject to any disclaimer , the term of this 6 , 815 ,432 B2 11/ 2004 Wheeler et al. patent is extended or adjusted under 35 7 , 422 , 902 B1 9 /2008 Wheeler et al . 7 , 745 ,651 B2 6 / 2010 Heyes et al . U . S . C . 154 ( b ) by 0 days. 7 , 799 , 565 B2 9 / 2010 MacLachlan et al. (21 ) Appl. No. : 16 / 280, 772 7 , 803 , 397 B2 9 / 2010 Heyes et al . 7 , 901, 708 B2 3 / 2011 MacLachlan et al. ( 22 ) Filed : Feb . 20 , 2019 8 , 101 ,741 B2 1 / 2012 MacLachlan et al . 8 , 188 , 263 B2 5 /2012 MacLachlan et al . (65 ) Prior Publication Data 8 , 236 , 943 B2 8 /2012 Lee et al . -
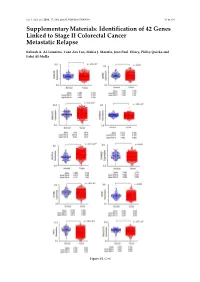
Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse
Int. J. Mol. Sci. 2016, 17, 598; doi:10.3390/ijms17040598 S1 of S16 Supplementary Materials: Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse Rabeah A. Al-Temaimi, Tuan Zea Tan, Makia J. Marafie, Jean Paul Thiery, Philip Quirke and Fahd Al-Mulla Figure S1. Cont. Int. J. Mol. Sci. 2016, 17, 598; doi:10.3390/ijms17040598 S2 of S16 Figure S1. Mean expression levels of fourteen genes of significant association with CRC DFS and OS that are differentially expressed in normal colon compared to CRC tissues. Each dot represents a sample. Table S1. Copy number aberrations associated with poor disease-free survival and metastasis in early stage II CRC as predicted by STAC and SPPS combined methodologies with resident gene symbols. CN stands for copy number, whereas CNV is copy number variation. Region Cytoband % of CNV Count of Region Event Gene Symbols Length Location Overlap Genes chr1:113,025,076–113,199,133 174,057 p13.2 CN Loss 0.0 2 AKR7A2P1, SLC16A1 chr1:141,465,960–141,822,265 356,305 q12–q21.1 CN Gain 95.9 1 SRGAP2B MIR5087, LOC10013000 0, FLJ39739, LOC10028679 3, PPIAL4G, PPIAL4A, NBPF14, chr1:144,911,564–146,242,907 1,331,343 q21.1 CN Gain 99.6 16 NBPF15, NBPF16, PPIAL4E, NBPF16, PPIAL4D, PPIAL4F, LOC645166, LOC388692, FCGR1C chr1:177,209,428–177,226,812 17,384 q25.3 CN Gain 0.0 0 chr1:197,652,888–197,676,831 23,943 q32.1 CN Gain 0.0 1 KIF21B chr1:201,015,278–201,033,308 18,030 q32.1 CN Gain 0.0 1 PLEKHA6 chr1:201,289,154–201,298,247 9093 q32.1 CN Gain 0.0 0 chr1:216,820,186–217,043,421 223,235 q41 CN -

Single Cell Derived Clonal Analysis of Human Glioblastoma Links
SUPPLEMENTARY INFORMATION: Single cell derived clonal analysis of human glioblastoma links functional and genomic heterogeneity ! Mona Meyer*, Jüri Reimand*, Xiaoyang Lan, Renee Head, Xueming Zhu, Michelle Kushida, Jane Bayani, Jessica C. Pressey, Anath Lionel, Ian D. Clarke, Michael Cusimano, Jeremy Squire, Stephen Scherer, Mark Bernstein, Melanie A. Woodin, Gary D. Bader**, and Peter B. Dirks**! ! * These authors contributed equally to this work.! ** Correspondence: [email protected] or [email protected]! ! Supplementary information - Meyer, Reimand et al. Supplementary methods" 4" Patient samples and fluorescence activated cell sorting (FACS)! 4! Differentiation! 4! Immunocytochemistry and EdU Imaging! 4! Proliferation! 5! Western blotting ! 5! Temozolomide treatment! 5! NCI drug library screen! 6! Orthotopic injections! 6! Immunohistochemistry on tumor sections! 6! Promoter methylation of MGMT! 6! Fluorescence in situ Hybridization (FISH)! 7! SNP6 microarray analysis and genome segmentation! 7! Calling copy number alterations! 8! Mapping altered genome segments to genes! 8! Recurrently altered genes with clonal variability! 9! Global analyses of copy number alterations! 9! Phylogenetic analysis of copy number alterations! 10! Microarray analysis! 10! Gene expression differences of TMZ resistant and sensitive clones of GBM-482! 10! Reverse transcription-PCR analyses! 11! Tumor subtype analysis of TMZ-sensitive and resistant clones! 11! Pathway analysis of gene expression in the TMZ-sensitive clone of GBM-482! 11! Supplementary figures and tables" 13" "2 Supplementary information - Meyer, Reimand et al. Table S1: Individual clones from all patient tumors are tumorigenic. ! 14! Fig. S1: clonal tumorigenicity.! 15! Fig. S2: clonal heterogeneity of EGFR and PTEN expression.! 20! Fig. S3: clonal heterogeneity of proliferation.! 21! Fig. -

Specialized Interferon Ligand Action in COVID19
medRxiv preprint doi: https://doi.org/10.1101/2021.07.29.21261325; this version posted August 1, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. It is made available under a CC-BY 4.0 International license . Specialized interferon ligand action in COVID19 Matthew D. Galbraith1,2, Kohl T. Kinning1, Kelly D. Sullivan1,3, Paula Araya1, Keith P. Smith1, Ross E. Granrath1, Jessica R. Shaw1, Ryan Baxter4, Kimberly R. Jordan4, Seth Russell5, Monika Dzieciatkowska6, Julie A. Reisz6, Fabia Gamboni6, Francesca CenDali6, Tusharkanti Ghosh7, AnDrew A. Monte8, Tellen D. Bennett9, Kirk C. Hansen6, Elena W.Y. Hsieh4,10, Angelo D’AlessanDro6, and Joaquin M. Espinosa1,2* Affiliations: 1Linda Crnic Institute for Down SynDrome, University of ColoraDo Anschutz MeDical Campus; Aurora, CO, USA. 2Department of Pharmacology, University of ColoraDo Anschutz MeDical Campus; Aurora, CO, USA. 3Department of PeDiatrics, Section of Developmental Biology, University of ColoraDo Anschutz MeDical Campus; Aurora, CO, USA. 4Department of Immunology anD Microbiology, University of ColoraDo Anschutz MeDical Campus; Aurora, CO, USA. 5Data Science to Patient Value, University of Colorado Anschutz Medical Campus; Aurora, CO, USA. 6Department of Biochemistry anD Molecular Genetics, University of ColoraDo Anschutz MeDical Campus; Aurora, CO, USA. 7Department of Biostatistics anD Informatics, ColoraDo School of Public Health; Aurora, CO, USA. 8Department of Emergency MeDicine, University of ColoraDo Anschutz MeDical Campus; Aurora, CO, USA. 9Department of PeDiatrics, Sections of Informatics anD Data Science anD Critical Care MeDicine, University of ColoraDo Anschutz Medical Campus; Aurora, CO, USA. -

Interferonsource.Com What's Inside New Products Research Tools Protocols & Tips Product Citations What's Your IFN-Α
Interferon Matters A quarterly newsletter by PBL InterferonSource interferonsource.com May 2007, Issue 2 u What’s Inside What’s Your IFN-a Type? New Products Are all Type I IFNs created equal? Of Mice and Men page 4 & 5 Type I Interferons were the first cytokines to be The biological significance of the expression of discovered. They are a family of homologous cytokines so many different IFN-a subtypes has yet to be Rhesus/Cynomolgus originally identified for their antiviral activity but have determined. However, reports suggest that they show IFN-a2 Subtype also been shown to have both anti-proliferative and quantitatively distinct anti-viral, anti-proliferative, immunomodulatory activities. In humans and mice and killer cell-stimulatory activities. It is therefore Mouse IFN-a4 Subtype they are encoded by an intronless multigene family of importance for researchers to characterize and clustered on human chromosome 9 and murine study these IFN-a subtypes and their possible post- chromosome 4 respectively. translational modifications. Research Tools Also in both species there are multiple Interferon- The common models for IFN-a subtypes study are page 6 & 7 Alpha (IFN-a) genes and one Interferon-Beta (IFN-b) human and mouse. In 1998, Lawson et al showed that Human IFN-a gene. Other Type I Interferon genes described include there are in vivo differences in the antiviral activity of the Interferon-Omega (IFN-w) gene, murine limitin the mouse IFN-a subtypes.1 Using an experimental Multi-Subtype ELISA Kit gene, as well as the human/murine Interferon-Epilson, mouse model of IFN transgene expression in vivo, TM Interferon-Kappa, and Interferon-Tau (IFN-e, IFN-k, and the authors showed that mouse IFN-a1 has a greater iLite Human IFN-a Kit IFN-t). -

Variation in the Type I Interferon Gene Cluster on 9P21 Influences Susceptibility to Asthma and Atopy
Genes and Immunity (2006) 7, 169–178 & 2006 Nature Publishing Group All rights reserved 1466-4879/06 $30.00 www.nature.com/gene ORIGINAL ARTICLE Variation in the type I interferon gene cluster on 9p21 influences susceptibility to asthma and atopy A Chan1,2, DL Newman1,3, AM Shon, DH Schneider, S Kuldanek and C Ober Department of Human Genetics, The University of Chicago, Chicago, IL, USA A genome-wide screen for asthma and atopy susceptibility alleles conducted in the Hutterites, a founder population of European descent, reported evidence of linkage with a short tandem repeat polymorphism (STRP) within the type I interferon (IFN) gene cluster on chromosome 9p21. The goal of this study was to identify variation within the IFN gene cluster that influences susceptibility to asthma and atopic phenotypes. We screened approximately 25 kb of sequence, including the flanking sequence of all 15 functional genes and the single coding exon in 12, in Hutterites representing different IFNA-STRP genotypes. We identified 78 polymorphisms, and genotyped 40 of these (in 14 genes) in a large Hutterite pedigree. Modest associations (0.003oPo0.05) with asthma, bronchial hyper-responsiveness (BHR), and atopy were observed with individual variants or genes, spanning the entire 400 kb region. However, pairwise combinations of haplotypes between genes showed highly significant associations with different phenotypes (Po10À5) that were localized to specific pairs of genes or regions of this cluster. These results suggest that variation in multiple genes in the type I IFN cluster on 9p22 contribute to asthma and atopy susceptibility, and that not all genes contribute equally to all phenotypes. -

Protein-Protein Interaction Analysis of Human
Open Access Original Article Functional Interactions of IFNAR-2 Within Biological Networks Pak Armed Forces Med J 2020; 70 (1): 245-52 PROTEIN-PROTEIN INTERACTION ANALYSIS OF HUMAN INTERFERON ALPHA RECEPTOR 2 (IFNAR-2) PROTEIN USING STRING SERVER Gulshan Ara Trali, Ambreen Javed*, Alia Sadiq* Swat Medical College, Swat Pakistan, *HITEC-Institute of Medical Sciences, Taxila/National University of Medical Sciences (NUMS) Pakistan ABSTRACT Objective: To study the functional and molecular interactions of IFNAR-2 within biological networks Study Design: Computational analysis: STRING software Place and Duration of Study: Department of Biochemistry, HITEC-Institute of Medical Sciences, Taxila Cantt, Pakistan, from Dec 2017 to Jun 2018. Methodology: Protein sequence of IFNAR-2 protein was obtained from ‘National Centre for Biotechnology Information (NCBI)’ database and STRING analysis conducted by applying specific parameters including (1) Text mining (2) Experiments (3) Databases (4) Co-expression (5) Neighborhood (6) Gene fusion and (7) Co-occurrence for identifying protein-protein interactions and molecular associations. Maximum number of interactors was set at 20 and highest confidence level was set at 0.900. Results: Protein-protein interaction analysis translate that human IFNAR-2 protein has high level of interactions with a set of proteins of similar size, drawn from the genome. This set represents a partial biologically connected group of proteins. This information has a potential to set the basis for further experimental investigations in more integrated and biologically linked pathway-oriented perspective that results in more targeted outcomes. Conclusion: Functional and molecular enrichment through STRING analysis revealed that IFNAR-2 protein has strong associations and serves as a key player in antiviral response of immune system. -

Systems Approaches to Modeling Chronic Mucosal Inflammation
Hindawi Publishing Corporation BioMed Research International Volume 2013, Article ID 505864, 17 pages http://dx.doi.org/10.1155/2013/505864 Research Article Systems Approaches to Modeling Chronic Mucosal Inflammation Mridul Kalita,1 Bing Tian,2 Boning Gao,3 Sanjeev Choudhary,1,2,4 Thomas G. Wood,1,4,5 Joseph R. Carmical,5 Istvan Boldogh,1,6 Sankar Mitra,1,5 John D. Minna,3 and Allan R. Brasier1,2,4 1 Sealy Center for Molecular Medicine, The University of Texas Medical Branch, 301 University Boulevard, Galveston, TX 77555, USA 2 Department of Internal Medicine, The University of Texas Medical Branch, 301 University Boulevard, Galveston, TX 77555, USA 3 Hamon Center for Therapeutic Oncology Research, Department of Internal Medicine Pharmacology, University of Texas Southwestern Medical Center, Dallas, TX 75390, USA 4 Institute for Translational Sciences, The University of Texas Medical Branch, 301 University Boulevard, Galveston, TX 77555, USA 5 Departments of Biochemistry and Molecular Biology, The University of Texas Medical Branch, 301 University Boulevard, Galveston, TX 77555, USA 6 Microbiology and Immunology, The University of Texas Medical Branch, 301 University Boulevard, Galveston, TX 77555, USA Correspondence should be addressed to Allan R. Brasier; [email protected] Received 12 June 2013; Revised 8 August 2013; Accepted 9 August 2013 Academic Editor: Tao Huang Copyright © 2013 Mridul Kalita et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. The respiratory mucosa is a major coordinator of the inflammatory response in chronic airway diseases, including asthma and chronic obstructive pulmonary disease (COPD).