Evolution of Cellular Diversity in Primary Motor Cortex of Human, Marmoset Monkey, and Mouse
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

The Mineralocorticoid Receptor Leads to Increased Expression of EGFR
www.nature.com/scientificreports OPEN The mineralocorticoid receptor leads to increased expression of EGFR and T‑type calcium channels that support HL‑1 cell hypertrophy Katharina Stroedecke1,2, Sandra Meinel1,2, Fritz Markwardt1, Udo Kloeckner1, Nicole Straetz1, Katja Quarch1, Barbara Schreier1, Michael Kopf1, Michael Gekle1 & Claudia Grossmann1* The EGF receptor (EGFR) has been extensively studied in tumor biology and recently a role in cardiovascular pathophysiology was suggested. The mineralocorticoid receptor (MR) is an important efector of the renin–angiotensin–aldosterone‑system and elicits pathophysiological efects in the cardiovascular system; however, the underlying molecular mechanisms are unclear. Our aim was to investigate the importance of EGFR for MR‑mediated cardiovascular pathophysiology because MR is known to induce EGFR expression. We identifed a SNP within the EGFR promoter that modulates MR‑induced EGFR expression. In RNA‑sequencing and qPCR experiments in heart tissue of EGFR KO and WT mice, changes in EGFR abundance led to diferential expression of cardiac ion channels, especially of the T‑type calcium channel CACNA1H. Accordingly, CACNA1H expression was increased in WT mice after in vivo MR activation by aldosterone but not in respective EGFR KO mice. Aldosterone‑ and EGF‑responsiveness of CACNA1H expression was confrmed in HL‑1 cells by Western blot and by measuring peak current density of T‑type calcium channels. Aldosterone‑induced CACNA1H protein expression could be abrogated by the EGFR inhibitor AG1478. Furthermore, inhibition of T‑type calcium channels with mibefradil or ML218 reduced diameter, volume and BNP levels in HL‑1 cells. In conclusion the MR regulates EGFR and CACNA1H expression, which has an efect on HL‑1 cell diameter, and the extent of this regulation seems to depend on the SNP‑216 (G/T) genotype. -

Bppart and Bpmax: RNA-RNA Interaction Partition Function and Structure Prediction for the Base Pair Counting Model
BPPart and BPMax: RNA-RNA Interaction Partition Function and Structure Prediction for the Base Pair Counting Model Ali Ebrahimpour-Boroojeny, Sanjay Rajopadhye, and Hamidreza Chitsaz ∗ Department of Computer Science, Colorado State University Abstract A few elite classes of RNA-RNA interaction (RRI), with complex roles in cellular functions such as miRNA-target and lncRNAs in human health, have already been studied. Accordingly, RRI bioinfor- matics tools tailored for those elite classes have been proposed in the last decade. Interestingly, there are somewhat unnoticed mRNA-mRNA interactions in the literature with potentially drastic biological roles. Hence, there is a need for high-throughput generic RRI bioinformatics tools. We revisit our RRI partition function algorithm, piRNA, which happens to be the most comprehensive and computationally-intensive thermodynamic model for RRI. We propose simpler models that are shown to retain the vast majority of the thermodynamic information that piRNA captures. We simplify the energy model and instead consider only weighted base pair counting to obtain BPPart for Base-pair Partition function and BPMax for Base-pair Maximization which are 225 and 1350 faster ◦ × × than piRNA, with a correlation of 0.855 and 0.836 with piRNA at 37 C on 50,500 experimentally charac- terized RRIs. This correlation increases to 0.920 and 0.904, respectively, at 180◦C. − Finally, we apply our algorithm BPPart to discover two disease-related RNAs, SNORD3D and TRAF3, and hypothesize their potential roles in Parkinson's disease and Cerebral Autosomal Dominant Arteri- opathy with Subcortical Infarcts and Leukoencephalopathy (CADASIL). 1 Introduction Since mid 1990s with the advent of RNA interference discovery, RNA-RNA interaction (RRI) has moved to the spotlight in modern, post-genome biology. -
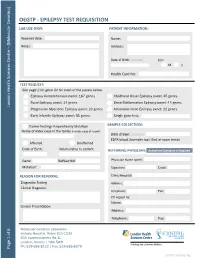
Oegtp - Epilepsy Test Requisition Lab Use Only: Patient Information
OEGTP - EPILEPSY TEST REQUISITION LAB USE ONLY: PATIENT INFORMATION: Received date: Name: Notes: Address: Date of Birth: YY/MM/DD Sex: M F Health Card No: TEST REQUEST: See page 2 for gene list for each of the panels below Epilepsy Comprehensive panel: 167 genes Childhood Onset Epilepsy panel: 45 genes Focal Epilepsy panel: 14 genes Brain Malformation Epilepsy panel: 44 genes London Health Sciences Centre – (Molecular Genetics) London Health Sciences Centre Progressive Myoclonic Epilepsy panel: 20 genes Actionable Gene Epilepsy panel: 22 genes Early Infantile Epilepsy panel: 51 genes Single gene test: Carrier Testing/ KnownFamily Mutation SAMPLE COLLECTION: Name of index case in the family (include copy of report) Date drawn: YY/MM/DD EDTA blood (lavender top) (5ml at room temp) Affected Unaffected Date of Birth: Relationship to patient: REFERRING PHYSICIAN: Authorized Signature is Required Gene: RefSeq:NM Physician Name (print): Mutation: Signature: Email: REASON FOR REFERRAL: Clinic/Hospital: Diagnostic Testing Address: Clinical Diagnosis: Telephone: Fax: CC report to: Name: Clinical Presentation: Address: Telephone: Fax: Molecular Genetics Laboratory Victoria Hospital, Room B10-123A 800 Commissioners Rd. E. London, Ontario | N6A 5W9 Pathology and Laboratory Medicine Ph: 519-685-8122 | Fax: 519-685-8279 Page 1 of 6 Page OEGTP (2021/05/28) OEGTP - EPILEPSY TEST PANELS Patient Identifier: COMPREHENSIVE EPILEPSY PANEL: 167 Genes ACTB, ACTG1, ADSL, AKT3, ALDH7A1, AMT, AP3B2, ARFGEF2, ARHGEF9, ARV1, ARX, ASAH1, ASNS, ATP1A3, ATP6V0A2, ATP7A, -

Rescue of Motor Coordination by Purkinje Cell-Targeted Restoration of Kv3.3 Channels in Kcnc3-Null Mice Requires Kcnc1
The Journal of Neuroscience, December 16, 2009 • 29(50):15735–15744 • 15735 Cellular/Molecular Rescue of Motor Coordination by Purkinje Cell-Targeted Restoration of Kv3.3 Channels in Kcnc3-Null Mice Requires Kcnc1 Edward C. Hurlock, Mitali Bose, Ganon Pierce, and Rolf H. Joho Department of Neuroscience, The University of Texas Southwestern Medical Center, Dallas, Texas 75390-9111 The role of cerebellar Kv3.1 and Kv3.3 channels in motor coordination was examined with an emphasis on the deep cerebellar nuclei (DCN). Kv3 channel subunits encoded by Kcnc genes are distinguished by rapid activation and deactivation kinetics that support high-frequency, narrow action potential firing. Previously we reported that increased lateral deviation while ambulating and slips while traversing a narrow beam of ataxic Kcnc3-null mice were corrected by restoration of Kv3.3 channels specifically to Purkinje cells, whereas Kcnc3-mutant mice additionally lacking one Kcnc1 allele were partially rescued. Here, we report mice lacking all Kcnc1 and Kcnc3 alleles exhibit no such rescue. For Purkinje cell output to reach the rest of the brain it must be conveyed by neurons of the DCN or vestibular nuclei. As Kcnc1, but not Kcnc3, alleles are lost, mutant mice exhibit increasing gait ataxia accompanied by spike broadening and deceleration in DCN neurons, suggesting the facet of coordination rescued by Purkinje-cell-restricted Kv3.3 restoration in mice lacking just Kcnc3 is hypermetria, while gait ataxia emerges when additionally Kcnc1 alleles are lost. Thus, fast repolarization in Purkinje cells appears important for normal movement velocity, whereas DCN neurons are a prime candidate locus where fast repolarization is necessary for normal gait patterning. -

Dynamic Clonal Progression in Xenografts of Acute Lymphoblastic Leukemia with Intrachromosomal Amplification of Chromosome 21
Sinclair PB, Blair HH, Ryan SL, Buechler L, Cheng J, Clayton J, Hanna R, Hollern S, Hawking Z, Bashton M, Schwab CJ, Jones L, Russell LJ, Marr H, Carey P, Halsey C, Heidenreich O, Moorman AV, Harrison CJ. Dynamic clonal progression in xenografts of acute lymphoblastic leukemia with intrachromosomal amplification of chromosome 21. Haematologica 2018, https://doi.org/10.3324/haematol.2017.172304 Copyright: © 2018, Ferrata Storti Foundation. Authors will grant copyright of their article to the Ferrata Storti Foundation. No formal permission will be required to reproduce parts (tables or illustrations) of published papers, provided the source is quoted appropriately and reproduction has no commercial intent. Reproductions with commercial intent will require written permission and payment of royalties. Please contact the office for requests: [email protected] DOI link to article: https://doi.org/10.3324/haematol.2017.172304 Date deposited: 20/03/2018 This work is licensed under a Creative Commons Attribution-NonCommercial 3.0 Unported License Newcastle University ePrints - eprint.ncl.ac.uk Published Ahead of Print on February 15, 2018, as doi:10.3324/haematol.2017.172304. Copyright 2018 Ferrata Storti Foundation. Dynamic clonal progression in xenografts of acute lymphoblastic leukemia with intrachromosomal amplification of chromosome 21 by Paul B. Sinclair, Helen H. Blair, Sarra L. Ryan, Lars Buechler, Joanna Cheng, Jake Clayton, Rebecca Hanna, Shaun Hollern, Zoe Hawking, Matthew Bashton, Claire J. Schwab, Lisa Jones, Lisa J. Russell, Helen Marr, Peter Carey, Christina Halsey, Olaf Heidenreich, Anthony V. Moorman, and Christine J. Harrison Haematologica 2018 [Epub ahead of print] Citation: Paul B. Sinclair, Helen H. -

CD29 Identifies IFN-Γ–Producing Human CD8+ T Cells With
+ CD29 identifies IFN-γ–producing human CD8 T cells with an increased cytotoxic potential Benoît P. Nicoleta,b, Aurélie Guislaina,b, Floris P. J. van Alphenc, Raquel Gomez-Eerlandd, Ton N. M. Schumacherd, Maartje van den Biggelaarc,e, and Monika C. Wolkersa,b,1 aDepartment of Hematopoiesis, Sanquin Research, 1066 CX Amsterdam, The Netherlands; bLandsteiner Laboratory, Oncode Institute, Amsterdam University Medical Center, University of Amsterdam, 1105 AZ Amsterdam, The Netherlands; cDepartment of Research Facilities, Sanquin Research, 1066 CX Amsterdam, The Netherlands; dDivision of Molecular Oncology and Immunology, Oncode Institute, The Netherlands Cancer Institute, 1066 CX Amsterdam, The Netherlands; and eDepartment of Molecular and Cellular Haemostasis, Sanquin Research, 1066 CX Amsterdam, The Netherlands Edited by Anjana Rao, La Jolla Institute for Allergy and Immunology, La Jolla, CA, and approved February 12, 2020 (received for review August 12, 2019) Cytotoxic CD8+ T cells can effectively kill target cells by producing therefore developed a protocol that allowed for efficient iso- cytokines, chemokines, and granzymes. Expression of these effector lation of RNA and protein from fluorescence-activated cell molecules is however highly divergent, and tools that identify and sorting (FACS)-sorted fixed T cells after intracellular cytokine + preselect CD8 T cells with a cytotoxic expression profile are lacking. staining. With this top-down approach, we performed an un- + Human CD8 T cells can be divided into IFN-γ– and IL-2–producing biased RNA-sequencing (RNA-seq) and mass spectrometry cells. Unbiased transcriptomics and proteomics analysis on cytokine- γ– – + + (MS) analyses on IFN- and IL-2 producing primary human producing fixed CD8 T cells revealed that IL-2 cells produce helper + + + CD8 Tcells. -

S41467-020-18249-3.Pdf
ARTICLE https://doi.org/10.1038/s41467-020-18249-3 OPEN Pharmacologically reversible zonation-dependent endothelial cell transcriptomic changes with neurodegenerative disease associations in the aged brain Lei Zhao1,2,17, Zhongqi Li 1,2,17, Joaquim S. L. Vong2,3,17, Xinyi Chen1,2, Hei-Ming Lai1,2,4,5,6, Leo Y. C. Yan1,2, Junzhe Huang1,2, Samuel K. H. Sy1,2,7, Xiaoyu Tian 8, Yu Huang 8, Ho Yin Edwin Chan5,9, Hon-Cheong So6,8, ✉ ✉ Wai-Lung Ng 10, Yamei Tang11, Wei-Jye Lin12,13, Vincent C. T. Mok1,5,6,14,15 &HoKo 1,2,4,5,6,8,14,16 1234567890():,; The molecular signatures of cells in the brain have been revealed in unprecedented detail, yet the ageing-associated genome-wide expression changes that may contribute to neurovas- cular dysfunction in neurodegenerative diseases remain elusive. Here, we report zonation- dependent transcriptomic changes in aged mouse brain endothelial cells (ECs), which pro- minently implicate altered immune/cytokine signaling in ECs of all vascular segments, and functional changes impacting the blood–brain barrier (BBB) and glucose/energy metabolism especially in capillary ECs (capECs). An overrepresentation of Alzheimer disease (AD) GWAS genes is evident among the human orthologs of the differentially expressed genes of aged capECs, while comparative analysis revealed a subset of concordantly downregulated, functionally important genes in human AD brains. Treatment with exenatide, a glucagon-like peptide-1 receptor agonist, strongly reverses aged mouse brain EC transcriptomic changes and BBB leakage, with associated attenuation of microglial priming. We thus revealed tran- scriptomic alterations underlying brain EC ageing that are complex yet pharmacologically reversible. -

Inhibition of Glutamate Decarboxylase (GAD) by Ethyl Ketopentenoate (EKP)
www.nature.com/scientificreports OPEN Inhibition of glutamate decarboxylase (GAD) by ethyl ketopentenoate (EKP) induces Received: 24 March 2017 Accepted: 9 June 2017 treatment-resistant epileptic Published online: 3 August 2017 seizures in zebrafsh Yifan Zhang 1, Michiel Vanmeert2, Aleksandra Siekierska1, Annelii Ny1, Jubi John3,6, Geert Callewaert 4, Eveline Lescrinier2, Wim Dehaen3, Peter A. M. de Witte1 & Rafal M. Kaminski 5 Epilepsy is a chronic brain disorder characterized by recurrent seizures due to abnormal, excessive and synchronous neuronal activities in the brain. It afects approximately 65 million people worldwide, one third of which are still estimated to sufer from refractory seizures. Glutamic acid decarboxylase (GAD) that converts glutamate into GABA is a key enzyme in the dynamic regulation of neural network excitability. Importantly, clinical evidence shows that lowered GAD activity is associated with several forms of epilepsy which are often treatment resistant. In the present study, we synthetized and explored the possibility of using ethyl ketopentenoate (EKP), a lipid-permeable GAD-inhibitor, to induce refractory seizures in zebrafsh larvae. Our results demonstrate that EKP evoked robust convulsive locomotor activities, excessive epileptiform discharges and upregulated c-fos expression in zebrafsh. Moreover, transgenic animals in which neuronal cells express apoaequorin, a Ca2+-sensitive bioluminescent photoprotein, displayed large luminescence signals indicating strong EKP-induced neuronal activation. Molecular docking data indicated that this proconvulsant activity resulted from the direct inhibition of both gad67 and gad65. Limited protective efcacy of tested anti-seizure drugs (ASDs) demonstrated a high level of treatment resistance of EKP-induced seizures. We conclude that the EKP zebrafsh model can serve as a high-throughput platform for novel ASDs discovery. -

Whole-Genome and RNA Sequencing Reveal Variation and Transcriptomic Coordination in the Developing Human Prefrontal Cortex
UCSF UC San Francisco Previously Published Works Title Whole-Genome and RNA Sequencing Reveal Variation and Transcriptomic Coordination in the Developing Human Prefrontal Cortex. Permalink https://escholarship.org/uc/item/8j92s7z1 Journal Cell reports, 31(1) ISSN 2211-1247 Authors Werling, Donna M Pochareddy, Sirisha Choi, Jinmyung et al. Publication Date 2020-04-01 DOI 10.1016/j.celrep.2020.03.053 Peer reviewed eScholarship.org Powered by the California Digital Library University of California HHS Public Access Author manuscript Author ManuscriptAuthor Manuscript Author Cell Rep Manuscript Author . Author manuscript; Manuscript Author available in PMC 2020 June 15. Published in final edited form as: Cell Rep. 2020 April 07; 31(1): 107489. doi:10.1016/j.celrep.2020.03.053. Whole-Genome and RNA Sequencing Reveal Variation and Transcriptomic Coordination in the Developing Human Prefrontal Cortex Donna M. Werling1,2,32, Sirisha Pochareddy3,32, Jinmyung Choi3,32, Joon-Yong An1,4,5,32, Brooke Sheppard1, Minshi Peng6, Zhen Li3,7, Claudia Dastmalchi1, Gabriel Santpere3,8, André M.M. Sousa3, Andrew T.N Tebbenkamp3, Navjot Kaur3, Forrest O. Gulden3, Michael S. Breen9,10,11,12, Lindsay Liang1, Michael C. Gilson1, Xuefang Zhao13,14,15, Shan Dong1, Lambertus Klei16, A. Ercument Cicek17,18, Joseph D. Buxbaum9,10,11,19, Homa Adle- Biassette20, Jean-Leon Thomas21,22, Kimberly A. Aldinger23,24, Diana R. O’Day25, Ian A. Glass25, Noah A. Zaitlen26, Michael E. Talkowski13,14,15, Kathryn Roeder6,18, Matthew W. State1,27, Bernie Devlin16, Stephan J. Sanders1,27,33,*, -

Molecular Characterization of Selectively Vulnerable Neurons in Alzheimer's Disease
HHS Public Access Author manuscript Author ManuscriptAuthor Manuscript Author Nat Neurosci Manuscript Author . Author manuscript; Manuscript Author available in PMC 2021 July 11. Published in final edited form as: Nat Neurosci. 2021 February ; 24(2): 276–287. doi:10.1038/s41593-020-00764-7. Molecular characterization of selectively vulnerable neurons in Alzheimer’s Disease Kun Leng1,2,3,4,13, Emmy Li1,2,3,13, Rana Eser5, Antonia Piergies5, Rene Sit2, Michelle Tan2, Norma Neff2, Song Hua Li5, Roberta Diehl Rodriguez6, Claudia Kimie Suemoto7,8, Renata Elaine Paraizo Leite7, Alexander J. Ehrenberg5, Carlos A. Pasqualucci7, William W. Seeley5,9, Salvatore Spina5, Helmut Heinsen7,10, Lea T. Grinberg5,7,11,*, Martin Kampmann1,2,12,* 1Institute for Neurodegenerative Disease, University of California, San Francisco, San Francisco, CA, USA. 2Chan Zuckerberg Biohub, San Francisco, CA, USA. 3Biomedical Sciences Graduate Program, University of California, San Francisco, San Francisco, CA, USA. 4Medical Scientist Training Program, University of California, San Francisco, San Francisco, CA, USA. 5Memory and Aging Center, Weill Institute for Neurosciences, Department of Neurology, University of California, San Francisco, San Francisco, CA, USA. 6Department of Neurology, Universidade de São Paulo, Faculdade de Medicina, São Paulo, Brazil. Users may view, print, copy, and download text and data-mine the content in such documents, for the purposes of academic research, subject always to the full Conditions of use:http://www.nature.com/authors/editorial_policies/license.html#terms * [email protected]; [email protected]. AUTHOR CONTRIBUTIONS K.L., E.L., L.T.G. and M.K. conceptualized and led the overall project. K.L., L.T.G. -

Effects on Production Traits of Haplotypes Among Casein Genes in Norwegian Goats and Evidence for a Site of Preferential Recombination
Copyright Ó 2006 by the Genetics Society of America DOI: 10.1534/genetics.106.058966 Effects on Production Traits of Haplotypes Among Casein Genes in Norwegian Goats and Evidence for a Site of Preferential Recombination Ben Hayes,1,2 Nina Hagesæther,1 Tormod A˚ dnøy, Grunde Pellerud, Paul R. Berg and Sigbjørn Lien Centre for Integrative Genetics and Department of Animal and Aquacultural Sciences, Norwegian University of Life Sciences, N-1432 A˚ s, Norway Manuscript received April 5, 2006 Accepted for publication July 2, 2006 ABSTRACT In goat milk the most abundant proteins are the casein genes, CSN1S1, CSN2, CSN1S2, and CSN3. Mutations have been identified within these genes affecting the level of gene expression, and effects on milk production traits have been reported. The aim of this study was to detect polymorphisms (SNPs) in the casein genes of Norwegian goats, resolve haplotype structures within the loci, and assess the effect of these haplotypes on milk production traits. Four hundred thirty-six Norwegian bucks were genotyped for 39 polymorphic sites across the four loci. The numbers of unique haplotypes present in each locus were 10, 6, 4, and 8 for CSN1S1, CSN2, CSN1S2, and CSN3, respectively. The effects of the CSN1S1 haplotypes on protein percentage and fat kilograms were significant, as were the effects of CSN3 haplotypes on fat percentage and protein percentage. A deletion in exon 12 of CSN1S1, unique to the Norwegian goat popu- lation, explained the effects of CSN1S1 haplotypes on fat kilograms, but not protein percentage. Inves- tigation of linkage disequilibrium between all possible pairs of SNPs revealed higher levels of linkage disequilbrium for SNP pairs within casein loci than for SNP pairs between casein loci, likely reflecting low levels of intragenic recombination. -

Novel Proteins Regulated by Mtor in Subependymal Giant Cell Astrocytomas of Patients with Tuberous Sclerosis Complex and New Therapeutic Implications
The American Journal of Pathology, Vol. 176, No. 4, April 2010 Copyright © American Society for Investigative Pathology DOI: 10.2353/ajpath.2010.090950 Molecular Pathogenesis of Genetic and Inherited Diseases Novel Proteins Regulated by mTOR in Subependymal Giant Cell Astrocytomas of Patients with Tuberous Sclerosis Complex and New Therapeutic Implications Magdalena Ewa Tyburczy,* Katarzyna Kotulska,† and demonstrated an effective modulation of SEGA Piotr Pokarowski,‡ Jakub Mieczkowski,* growth by pharmacological inhibition of both Joanna Kucharska,* Wieslawa Grajkowska,† mTOR and extracellular signal-regulated kinase sig- Maciej Roszkowski,§ Sergiusz Jozwiak,† naling pathways, which could represent a novel ther- and Bozena Kaminska* apeutic approach. (Am J Pathol 2010, 176:1878–1890; DOI: 10.2353/ajpath.2010.090950) From the Laboratory of Transcription Regulation,* the Nencki Institute of Experimental Biology, Warsaw; the Departments of Neurology and Epileptology,† and Neurosurgery,§ the Children’s Subependymal giant cell astrocytomas (SEGAs) are rare, Memorial Health Institute, Warsaw; and the Faculty of low-grade brain tumors (World Health Organization Grade I) Mathematics, Informatics, and Mechanics,‡ University of of a mixed glioneuronal lineage.1,2 They are observed in Warsaw, Poland 10% to 20% of patients with tuberous sclerosis complex (TSC) and are the major cause of morbidity in children and young adults with TSC.3 The disease affects about one in Subependymal giant cell astrocytomas (SEGAs) are 6000 people, is characterized by the formation of benign rare brain tumors associated with tuberous sclerosis tumors in multiple organs (mainly brain, heart, kidneys, skin, complex (TSC), a disease caused by mutations in TSC1 or lungs), and is often associated with epilepsy, mental or TSC2, resulting in enhancement of mammalian retardation, and autism.4,5 Tuberous sclerosis complex is target of rapamycin (mTOR) activity, dysregulation of caused by mutation in one of two tumor suppressor genes, cell growth, and tumorigenesis.