Assessment of Genetic Diversity in Picea Orientalis (L.) Link
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Two New Chrysomyxa Rust Species on the Endemic Plant, Picea Asperata in Western China, and Expanded Description of C
Phytotaxa 292 (3): 218–230 ISSN 1179-3155 (print edition) http://www.mapress.com/j/pt/ PHYTOTAXA Copyright © 2017 Magnolia Press Article ISSN 1179-3163 (online edition) https://doi.org/10.11646/phytotaxa.292.3.2 Two new Chrysomyxa rust species on the endemic plant, Picea asperata in western China, and expanded description of C. succinea JING CAO1, CHENG-MING TIAN1, YING-MEI LIANG2 & CHONG-JUAN YOU1* 1The Key Laboratory for Silviculture and Conservation of Ministry of Education, Beijing Forestry University, Beijing 100083, China 2Museum of Beijing Forestry University, Beijing 100083, China *Corresponding author: [email protected] Abstract Two new rust species, Chrysomyxa diebuensis and C. zhuoniensis, on Picea asperata are recognized by morphological characters and DNA sequence data. A detailed description, illustrations, and discussion concerning morphologically similar and phylogenetically closely related species are provided for each species. From light and scanning electron microscopy observations C. diebuensis is characterized by the nailhead to peltate aeciospores, with separated stilt-like base. C. zhuoni- ensis differs from other known Chrysomyxa species in the annulate aeciospores with distinct longitudinal smooth cap at ends of spores, as well as with a broken, fissured edge. Analysis based on internal transcribed spacer region (ITS) partial gene sequences reveals that the two species cluster as a highly supported group in the phylogenetic trees. Correlations between the morphological and phylogenetic features are discussed. Illustrations and a detailed description are also provided for the aecia of C. succinea in China for the first time. Keywords: aeciospores, molecular phylogeny, spruce needle rust, taxonomy Introduction Picea asperata Mast.is native to western China, widely distributed in Qinghai, Gansu, Shaanxi and western Sichuan. -

Pinus Sabiniana) in Oregon Frank Callahan PO Box 5531, Central Point, OR 97502
Discovering Gray Pine (Pinus sabiniana) in Oregon Frank Callahan PO Box 5531, Central Point, OR 97502 “Th e tree is remarkable for its airy, widespread tropical appearance, which suggest a region of palms rather than cool pine woods. Th e sunbeams sift through even the leafi est trees with scarcely any interruption, and the weary, heated traveler fi nds little protection in their shade.” –John Muir (1894) ntil fairly recently, gray pine was believed to be restricted to UCalifornia, where John Muir encountered it. But the fi rst report of it in Oregon dates back to 1831, when David Douglas wrote to the Linnaean Society of his rediscovery of Pinus sabiniana in California. In his letter from San Juan Bautista, Douglas claimed to have collected this pine in 1826 in Oregon while looking for sugar pine (Pinus lambertiana) between the Columbia and Umpqua rivers (Griffin 1962). Unfortunately, Douglas lost most of his fi eld notes and specimens when his canoe overturned in the Santiam River (Harvey 1947). Lacking notes and specimens, he was reluctant to report his original discovery of the new pine in Oregon until he found it again in California (Griffin 1962). Despite the delay in reporting it, Douglas clearly indicated that he had seen this pine before he found it in California, and the Umpqua region has suitable habitat for gray pine. John Strong Newberry1 (1857), naturalist on the 1855 Pacifi c Railroad Survey, described an Oregon distribution for Pinus sabiniana: “It was found by our party in the valleys of the coast ranges as far north as Fort Lane in Oregon.” Fort Lane was on the eastern fl ank of Blackwell Hill (between Central Point and Gold Hill in Jackson County), so his description may also include the Th e lone gray pine at Tolo, near the old Fort Lane site, displays the characteristic architecture of multiple Applegate Valley. -

Genetic Diversity and Conservation of Picea Chihuahuana Martínez: a Review
Vol. 13(28), pp. 2786-2795, 9 July, 2014 DOI: 10.5897/AJB2014.13645 Article Number: CADB48845877 ISSN 1684-5315 African Journal of Biotechnology Copyright © 2014 Author(s) retain the copyright of this article http://www.academicjournals.org/AJB Review Genetic diversity and conservation of Picea chihuahuana Martínez: A review Quiñones-Pérez, Carmen Zulema1, Sáenz-Romero, Cuauhtémoc2 and Wehenkel, Christian1* 1Institute of Forestry and Wood Industry, Universidad Juárez del Estado de Durango, Durango, México. 2Institute of Agricultural and Forestry Research, Universidad Michoacana de San Nicolás de Hidalgo, Michoacán, México. Received 20 January, 2014; Accepted 16 June, 2014 The conservation of genetic diversity in tree populations is an essential component of sustainable forest management. Picea chihuahuana Martínez is an endemic conifer species in Mexico and is considered to be endangered. P. chihuahuana covers a total area of no more than 300 ha at the Sierra Madre Occidental, a mountain range that harbor a high diversity of tree species. There are 40 populations of the species that have been identified in the region, and it cannot be found elsewhere. These populations form clusters within gallery forests and are usually associated with eight other tree genera. The P. chihuahuana community is mostly well preserved. Owing to its remarkable characteristics and high conservation value, P. chihuahuana has been the subject of several studies aimed at learning more about the genetic structure, ecology and potential effects of climate change. However, the overall applicability of such studies is to confirm a dataset to develop management tools to help decision makers and to implement preservation and conservation strategies using genetic diversity. -

RHS the Garden Magazine Index 2020
GardenThe INDEX 2020 Volume 145, Parts 1–12 Index 2020 January 2020 February 2020 March 2020 April 2020 May 2020 June 2020 1 2 3 4 5 6 Coloured numbers campestre ‘William ‘Voodoo’ 9: 78 ‘Kaleidoscope’ lauterbachiana Plas Brondanw, North in bold before the page Caldwell’ 3: 32, 32 ‘Zwartkop’ 7: 22, 22; 11: 46, 46 1: 56, 57 Wales 12: 38–42, 38–42 number(s) denote the x freemanii Autumn 8: 54, 54 ‘Lavender Lady’ 6: 12, macrorrhizos 11: 33, 33 Andrews, Susyn, on: part number (month). Blaze (‘Jeffersred’) Aeschynanthus 3: 138 12; 11: 46–47, 47 micholitziana 2: 78 hollies, AGM cultivars Each part is paginated 10: 14, 14–15 Aesculus ‘Macho Mocha’ Aloe Safari Sunrise (‘X5’) 12: 31, 31 separately. griseum 1: 49; 2: 14, 14– hippocastanum 11: 46, 47 6: 12, 12 Anemone: 15; 11: 34, 35; 12: 10, 10; ‘Hampton Court ‘Mayan Queen’ 11: 46 Aloysia: ‘Frilly Knickers’ 9: 7, 7 Numbers in italics 12: 83 Gold’ 3: 89, 89 ‘Pineapple Express’ citrodora (lemon Wild Swan denote an image. micrantham 10: 80 ‘Wisselink’ 3: 89, 89 11: 47 verbena) 6: 87, 87, 88; (‘Macane001’) 5: 74, palmatum 4: 74–75; x neglecta ‘Silver Fox’ 11: 47 to infuse gin 4: 82, 83 74, 76 Where a plant has a 12: 65, 65 ‘Erythroblastos’ Aglaonema (Chinese gratissima angelica root to infuse Trade Designation ‘Garnet’ 10: 27, 27 3: 88, 88 evergreen): 1: 57; 7: 34, (whitebrush or gin 4: 82, 82 (also known as a selling platanoides Agapanthus: 5: 82, 83 34; 12: 32, 32 spearmint verbena) Angelonia Serena Series name) it is typeset in ‘Walderseei’ 3: 87, 87 ‘Blue Dot 9: 109 ‘King of Siam’ 1: 56, 57 6: 86, 88 8: 16, 17 a different font to pseudoplatanus ‘Bressingham Blue’ pictum ‘Tricolor’ Alstroemeria: angel’s trumpet (see distinguish it from the ‘Brilliantissimum’ 9: 109 1: 44, 45 Indian Summer Brugmansia) cultivar name (shown 3: 86, 86–87 ‘Cally Blue 9: 109 Agrostis nebulosa (‘Tesronto’) 8: 16, 16 Angwin, Kirsty, on: in ‘Single Quotes’). -

Current and Potential Spatial Distribution of Six Endangered Pine Species of Mexico: Towards a Conservation Strategy
Article Current and Potential Spatial Distribution of Six Endangered Pine Species of Mexico: Towards a Conservation Strategy Martin Enrique Romero-Sanchez * , Ramiro Perez-Miranda, Antonio Gonzalez-Hernandez, Mario Valerio Velasco-Garcia , Efraín Velasco-Bautista and Andrés Flores National Institute on Forestry, Agriculture and Livestock Research, Progreso 5, Barrio de Santa Catarina, Coyoacan, 04010 Mexico City, Mexico; [email protected] (R.P.-M.); [email protected] (A.G.-H.); [email protected] (M.V.V.-G.); [email protected] (E.V.-B.); fl[email protected] (A.F.) * Correspondence: [email protected]; Tel.: +52-553-626-8698 Received: 24 October 2018; Accepted: 6 December 2018; Published: 12 December 2018 Abstract: Mexico is home to the highest species diversity of pines: 46 species out of 113 reported around the world. Within the great diversity of pines in Mexico, Pinus culminicola Andresen et Beaman, P. jaliscana Perez de la Rosa, P. maximartinenzii Rzed., P. nelsonii Shaw, P. pinceana Gordon, and P. rzedowskii Madrigal et M. Caball. are six catalogued as threatened or endangered due to their restricted distribution and low population density. Therefore, they are of special interest for forest conservation purposes. In this paper, we aim to provide up-to-date information on the spatial distribution of these six pine species according to different historical registers coming from different herbaria distributed around the country by using spatial modeling. Therefore, we recovered historical observations of the natural distribution of each species and modelled suitable areas of distribution according to environmental requirements. Finally, we evaluated the distributions by contrasting changes of vegetation in the period 1991–2016. -

(Picea Orientalis (L.) Link) Stands in Turkey
Kastamonu Üni., Orman Fakültesi Dergisi, 2011, 11 (2):209 – 218 Kastamonu Univ., Journal of Forestry Faculty Examining Some Functional Characteristics in Subalpine Pure Oriental Spruce ( Picea orientalis (L.) Link) Stands in Turkey Ali Ömer ÜÇLER 1, *Zafer YÜCESAN 1, Ercan OKTAN 1, Ali DEMİRCİ 1, Hakkı YAVUZ 1, Altay Uğur GÜL 2 1Karadeniz Technical University Faculty of Forestry, Trabzon/TURKEY 2Celal Bayar University School of Tobacco Expertise, Manisa/TURKEY *Corresponding Author: [email protected] Geliş Tarihi:10.10.2011 Abstract This study was carried out in order to state the functional differences between the pure oriental spruce stands in subalpine zone. In the research, the pure oriental spruce stands in the subalpine zone were evaluated according to their wood production (F1), erosion control (F2), water production (F3), grass productivity-wild life habitat (F4), scenic beauty (F5) and avalanche prevention (F6) functions. Functional characteristics were rated as a scalar value between 1 (the worst) and 5 (the best) according to some observations, and exact differences were fixed between the functional characteristics of the treeline and the timberline stands. Determinations were realized in total of 46 sampling plots. 23 of the sampling plots were in treeline and 23 of them were in timberline. According to the results the F1, F2 and F6 values in the timberline were more effective than treeline stands. However F3, F4 and F5 values in the treeline were more effective than timber line stands. The average F1 value was 1.22 in the treeline and 3.04 in the timberline, F2 value was 3.0 in treeline and 3.61 in timberline, F3 value was 2.91 in the treeline and 2.43 in the timberline, F4 value was 3.35 in the treeline and 2.74 in the timberline, F5 value was 3.70 in treeline and 3.43 in timberline. -

Rutgers Home Gardeners School: Dwarf Conifer Presentation
Dwarf Conifers and Other Dwarf Ornamentals: Their Use in the Landscape Why Dwarf Conifers and Dwarf Ornamentals? Long lifespan in the garden Take up little space Management/Maintenance is done with both feet on the ground Many species live in pots 1 Attributes of dwarf conifers in containers They are easily transported from one part of the garden to another Evergreen forms provide much needed color/texture and form in the winter landscape. Some of the larger dwarfs end up bonsaiing themselves naturally in pots due to root restriction Once established, they require the moisture requirements of the rest of the garden Growing requirements: Light Full sun to half a day is required If only half a day you can choose between morning and afternoon. Note: With dwarf flowering ornamentals, they prefer more than half a day of sun. 2 Growing requirements: Water With permanent plantings in the garden, water twice a week for the first 4 to 6 weeks, after the initial time period, drop back to once a week. With pots and container gardening, water twice a week for the first 4 to 6 weeks and once a week thereafter through November 15 through the 30th. Note: containers will need to be checked for water needs throughout the winter. Growing requirements: Fertilization With dwarf conifers, they can go without fertilizer every other year. Application is early April: soil has to warm up so the fertilizer is used. Use an organic fertilizer like Hollytone. With all other dwarf ornamentals, feed in early April, Hollytone or Plant tone for example. -

Vegetation Responses to Interglacial Warming
Discussion Paper | Discussion Paper | Discussion Paper | Discussion Paper | Clim. Past Discuss., 9, 245–267, 2013 www.clim-past-discuss.net/9/245/2013/ Climate doi:10.5194/cpd-9-245-2013 of the Past CPD © Author(s) 2013. CC Attribution 3.0 License. Discussions 9, 245–267, 2013 This discussion paper is/has been under review for the journal Climate of the Past (CP). Vegetation Please refer to the corresponding final paper in CP if available. responses to interglacial warming A. V. Lozhkin and Vegetation responses to interglacial P. M. Anderson warming in the Arctic, examples from Lake El’gygytgyn, northeast Siberia Title Page Abstract Introduction 1 2 A. V. Lozhkin and P. M. Anderson Conclusions References 1 Northeast Interdisciplinary Scientific Research Institute, Far East Branch, Russian Academy Tables Figures of Sciences, 16 Portovaya Street, Magadan, 685000, Russia 2Earth & Space Sciences and Quaternary Research Center, University of Washington, Seattle, 98195-1310, USA J I Received: 28 August 2012 – Accepted: 3 September 2012 – Published: 15 January 2013 J I Correspondence to: P. M. Anderson ([email protected]) Back Close Published by Copernicus Publications on behalf of the European Geosciences Union. Full Screen / Esc Printer-friendly Version Interactive Discussion 245 Discussion Paper | Discussion Paper | Discussion Paper | Discussion Paper | Abstract CPD Palynological data from Lake El’gygytgyn reveal responses of plant communities to a range of climatic conditions that can help assess the possible impact of global warm- 9, 245–267, 2013 ing on arctoboreal ecosystems. Vegetation associated with climatic optima suggests 5 two types of interglacial responses: one is dominated by deciduous taxa (the post- Vegetation glacial thermal maximum (PGTM) and marine isotope stage (MIS5)) and the second responses to by evergreen conifers (MIS11, MIS31). -

Ecosystems and Diversity of the Sierra Madre Occidental
Ecosystems and Diversity of the Sierra Madre Occidental M. S. González-Elizondo, M. González-Elizondo, L. Ruacho González, I.L. Lopez Enriquez, F.I. Retana Rentería, and J.A. Tena Flores CIIDIR I.P.N. Unidad Durango, Mexico Abstract—The Sierra Madre Occidental (SMO) is the largest continuous ignimbrite plate on Earth. Despite its high biological and cultural diversity and enormous environmental and economical importance, it is yet not well known. We describe the vegetation and present a preliminary regionalization based on physio- graphic, climatic, and floristic criteria. A confluence of three main ecoregions in the area corresponds with three ecosystems: Temperate Sierras (Madrean), Semi-Arid Highlands (Madrean Xerophylous) and Tropical Dry Forests (Tropical). The Madrean region harbors five major vegetation types: pine forests, mixed conifer forests, pine-oak forests, oak forests and temperate mesophytic forests. The Madrean Xerophylous region has oak or pine-oak woodlands and evergreen juniper scrub with transitions toward the grassland and xerophylous scrub areas of the Mexican high plateau. The Tropical ecosystem, not discussed here, includes tropical deciduous forests and subtropical scrub. Besides fragmentation and deforestation resulting from anthropogenic activities, other dramatic changes are occurring in the SMO, including damage caused by bark beetles (Dendroctonus) in extensive areas, particularly in drought-stressed forests, as well as the expansion of chaparral and Dodoneaea scrub at the expense of temperate forest and woodlands. Comments on how these effects are being addressed are made. Introduction as megacenters of plant diversity: northern Sierra Madre Occidental and the Madrean Archipelago (Felger and others 1997) and the Upper The Sierra Madre Occidental (SMO) or Western Sierra Madre is the Mezquital River region (González-Elizondo 1997). -

Conifer Quarterly
Conifer Quarterly Vol. 26 No. 2 Spring 2009 Picea abies ‘Reflexa’ Photo by Dennis Groh The Conifer Quarterly is th e pub lication of the American Conifer Society Contents 7 How I spruced up Georgia by Tom Cox 10 A Fungus Made Us Do It by Bill Van Kosky 17 Brassica hirta by Phil Syphrit 19 Stonehedge, the Garden of Ken and Elena Jordan Text and Photos by Jack Ayers 26 Winter Burn on Conifers by Ellen Kelley 28 Evaluating Landscape Adaptability of Hemlock Species Picea abies 'Lanham's Beehive' ( spp.) in Western North Carolina Photo by Randall C. Smith, Courtesy of Iseli Nursery Tsuga by Anthony V. LeBude and Thomas G. Ranney 32 Collecting Northern Conifers by William Journeay 36 Conifer Destinations by Tom Cox 38 Conifers with Leaves by Maud Henne American Conifer Society Voices 2 President’s Message 4 Editor’s Memo 16 Letter to the Editor 41 ACS 2009 National Meeting 44 2009 Western Region Meeting 46 $7,000 Awarded to Four Southeast Regional Gardens Picea omorika ‘Pimoko’ Photo by Randall C. Smith, Courtesy of Iseli Nursery Vol. 26 No. 2 CONIFER QUARTERLY 3 Conifer AS I SEE IT FROM THE PRESIDENT ’S DESK “Everything is connected to everything At the recently concluded Winter way to recruit new members. else. There is one ecosphere for all living Board Meeting, the board focused on The charge to each region, then, is to Quarterly organisms and what affects one, affects membership. Our concern is not as much develop a plan to increase membership in all. ” with members who do not renew their region. -

In Vitro Plantlet Regeneration from Mature Zygotic Embryos of Pinus Pinea
In vitro plantlet regeneration from mature zygotic embryos of Pinus pinea L.: overcoming the rooting problems Carla Aparecida Ragonezi Gomes Lopes Thesis presented to obtain the PhD degree in Biology by the University of Évora Thesis presented to obtain the PhD degree Supervisors: Prof. Dr. Maria Amely Zavattieri in Biology Prof.by the Dr. UniversityMaria do Rosário of Évora Martins Prof. Dr. Ana Teresa Caldeira A s ÉVORA, MAY 2013 INSTITUTO DE INVESTIGAÇÃO E FORMAÇÃO AVANÇADA Contactos: Universidade de Évora Instituto de Investigação e Formação Avançada - IIFA Palácio do Vimioso | Largo Marquês de Marialva, Apart. 94 Funding 7002-554 Évora | Portugal Tel: (+351) 266 706 581 Fax: (+351) 266 744 677 email: [email protected] Ragonezi, C. (2012): In vitro plantlet regeneration from mature zygotic embryos of Pinus pinea L.: overcoming the rooting problems. Doctoral Dissertation presented at University of Évora, Portugal. Keywords: Adventitious rooting, biotization, ectomycorrhiza, in vitro mycorrhization, micropropagation, Pinus pinea L. This thesis was financed by the Portuguese Foundation for Science and Technology (FCT) SFRH / BD / 44119 / 2008 I hear and I forget I see and I remember I do and I understand Confucius (551 A.D. – 479 A.D.) Agradecimentos Como nenhuma grande realização da nossa vida é alcançada sozinha, esta parte da minha tese destina-se àquelas pessoas que fizeram com que este sonho longínquo e aparentemente intangível fosse certeza. Foram muitos anos, muito trabalho, muitas pessoas, distintos laboratórios, inúmeros equipamentos, muitas conversas, telefonemas e favores, por isso espero que aqueles que são citados tenham paciência de ler, e peço desculpas adiantadas para aqueles que me esqueci (não foi por mal, mas estava na fase crítica da entrega da tese). -
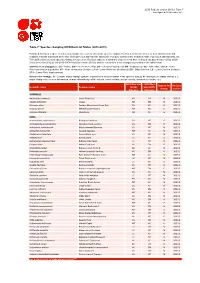
Table 7: Species Changing IUCN Red List Status (2012-2013)
IUCN Red List version 2013.2: Table 7 Last Updated: 25 November 2013 Table 7: Species changing IUCN Red List Status (2012-2013) Published listings of a species' status may change for a variety of reasons (genuine improvement or deterioration in status; new information being available that was not known at the time of the previous assessment; taxonomic changes; corrections to mistakes made in previous assessments, etc. To help Red List users interpret the changes between the Red List updates, a summary of species that have changed category between 2012 (IUCN Red List version 2012.2) and 2013 (IUCN Red List version 2013.2) and the reasons for these changes is provided in the table below. IUCN Red List Categories: EX - Extinct, EW - Extinct in the Wild, CR - Critically Endangered, EN - Endangered, VU - Vulnerable, LR/cd - Lower Risk/conservation dependent, NT - Near Threatened (includes LR/nt - Lower Risk/near threatened), DD - Data Deficient, LC - Least Concern (includes LR/lc - Lower Risk, least concern). Reasons for change: G - Genuine status change (genuine improvement or deterioration in the species' status); N - Non-genuine status change (i.e., status changes due to new information, improved knowledge of the criteria, incorrect data used previously, taxonomic revision, etc.) IUCN Red List IUCN Red Reason for Red List Scientific name Common name (2012) List (2013) change version Category Category MAMMALS Nycticebus javanicus Javan Slow Loris EN CR N 2013.2 Okapia johnstoni Okapi NT EN N 2013.2 Pteropus niger Greater Mascarene Flying