Gene Expression Profiles in the Brain
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Anti-Inflammatory Effects of V-3 Polyunsaturated Fatty Acids And
ORIGINAL ARTICLE Anti-inflammatory Effects of v-3 Polyunsaturated Fatty Acids and Soluble Epoxide Hydrolase Inhibitors in Angiotensin-II–Dependent Hypertension Arzu Ulu, MSc,* Todd R. Harris, PhD,* Christophe Morisseau, PhD,* Christina Miyabe, BS,* Hiromi Inoue, DVM,† Gertrud Schuster, PhD,‡ Hua Dong, PhD,* Ana-Maria Iosif, PhD,§ Jun-Yan Liu, PhD,* Robert H. Weiss, MD,†¶ Nipavan Chiamvimonvat, MD,k¶ John D. Imig, PhD,** and Bruce D. Hammock, PhD* renal markers of inflammation (ie, prostaglandins and MCP-1), Abstract: The mechanisms underlying the anti-inflammatory and downregulated an epithelial sodium channel, and upregulated angioten- antihypertensive effects of long-chain v-3 polyunsaturated fatty acids sin-converting enzyme-2 message and significantly modulated cyclo- (v-3 PUFAs) are still unclear. The epoxides of an v-6 fatty acid, oxygenase and lipoxygenase metabolic pathways. Overall, our findings arachidonic acid epoxyeicosatrienoic acids also exhibit antihyperten- suggest that epoxides of the v-3 PUFAs contribute to lowering systolic sive and anti-inflammatory effects. Thus, we hypothesized that the blood pressure and attenuating inflammation in part by reduced pros- major v-3 PUFAs, including eicosapentaenoic acid (EPA) and doco- taglandinsandMCP-1andbyupregulation of angiotensin-converting sahexaenoic acid (DHA), may lower the blood pressure and attenuate enzyme-2 in angiotensin-II–dependent hypertension. renal markers of inflammation through their epoxide metabolites. v Here, we supplemented mice with an v-3 rich diet for 3 weeks in Key Words: -3 polyunsaturated fatty acids, EPA, DHA, soluble – a murine model of angiotensin-II–dependent hypertension. Also, epoxide hydrolase inhibitors, angiotensin-II dependent hypertension because EPA and DHA epoxides are metabolized by soluble epoxide (J Cardiovasc PharmacolÔ 2013;62:285–297) hydrolase (sEH), we tested the combination of an sEH inhibitor and the v-3 rich diet. -

Isyte: Integrated Systems Tool for Eye Gene Discovery
Lens iSyTE: Integrated Systems Tool for Eye Gene Discovery Salil A. Lachke,1,2,3,4 Joshua W. K. Ho,1,4,5 Gregory V. Kryukov,1,4,6 Daniel J. O’Connell,1 Anton Aboukhalil,1,7 Martha L. Bulyk,1,8,9 Peter J. Park,1,5,10 and Richard L. Maas1 PURPOSE. To facilitate the identification of genes associated ther investigation. Extension of this approach to other ocular with cataract and other ocular defects, the authors developed tissue components will facilitate eye disease gene discovery. and validated a computational tool termed iSyTE (integrated (Invest Ophthalmol Vis Sci. 2012;53:1617–1627) DOI: Systems Tool for Eye gene discovery; http://bioinformatics. 10.1167/iovs.11-8839 udel.edu/Research/iSyTE). iSyTE uses a mouse embryonic lens gene expression data set as a bioinformatics filter to select candidate genes from human or mouse genomic regions impli- ven with the advent of high-throughput sequencing, the cated in disease and to prioritize them for further mutational Ediscovery of genes associated with congenital birth defects and functional analyses. such as eye defects remains a challenge. We sought to develop METHODS. Microarray gene expression profiles were obtained a straightforward experimental approach that could facilitate for microdissected embryonic mouse lens at three key devel- the identification of candidate genes for developmental disor- opmental time points in the transition from the embryonic day ders, and, as proof-of-principle, we chose defects involving the (E)10.5 stage of lens placode invagination to E12.5 lens primary ocular lens. Opacification of the lens results in cataract, a leading cause of blindness that affects 77 million persons and fiber cell differentiation. -

Protein Interaction Network of Alternatively Spliced Isoforms from Brain Links Genetic Risk Factors for Autism
ARTICLE Received 24 Aug 2013 | Accepted 14 Mar 2014 | Published 11 Apr 2014 DOI: 10.1038/ncomms4650 OPEN Protein interaction network of alternatively spliced isoforms from brain links genetic risk factors for autism Roser Corominas1,*, Xinping Yang2,3,*, Guan Ning Lin1,*, Shuli Kang1,*, Yun Shen2,3, Lila Ghamsari2,3,w, Martin Broly2,3, Maria Rodriguez2,3, Stanley Tam2,3, Shelly A. Trigg2,3,w, Changyu Fan2,3, Song Yi2,3, Murat Tasan4, Irma Lemmens5, Xingyan Kuang6, Nan Zhao6, Dheeraj Malhotra7, Jacob J. Michaelson7,w, Vladimir Vacic8, Michael A. Calderwood2,3, Frederick P. Roth2,3,4, Jan Tavernier5, Steve Horvath9, Kourosh Salehi-Ashtiani2,3,w, Dmitry Korkin6, Jonathan Sebat7, David E. Hill2,3, Tong Hao2,3, Marc Vidal2,3 & Lilia M. Iakoucheva1 Increased risk for autism spectrum disorders (ASD) is attributed to hundreds of genetic loci. The convergence of ASD variants have been investigated using various approaches, including protein interactions extracted from the published literature. However, these datasets are frequently incomplete, carry biases and are limited to interactions of a single splicing isoform, which may not be expressed in the disease-relevant tissue. Here we introduce a new interactome mapping approach by experimentally identifying interactions between brain-expressed alternatively spliced variants of ASD risk factors. The Autism Spliceform Interaction Network reveals that almost half of the detected interactions and about 30% of the newly identified interacting partners represent contribution from splicing variants, emphasizing the importance of isoform networks. Isoform interactions greatly contribute to establishing direct physical connections between proteins from the de novo autism CNVs. Our findings demonstrate the critical role of spliceform networks for translating genetic knowledge into a better understanding of human diseases. -

Parallel Molecular Evolution in Pathways, Genes, and Sites in High-Elevation Hummingbirds Revealed by Comparative Transcriptomics
GBE Parallel Molecular Evolution in Pathways, Genes, and Sites in High-Elevation Hummingbirds Revealed by Comparative Transcriptomics Marisa C.W. Lim1,*, Christopher C. Witt2, Catherine H. Graham1,3,andLilianaM.Davalos 1,4 1Department of Ecology and Evolution, Stony Brook University 2 Museum of Southwestern Biology and Department of Biology, University of New Mexico Downloaded from https://academic.oup.com/gbe/article-abstract/11/6/1552/5494706 by guest on 08 June 2019 3Swiss Federal Research Institute (WSL), Birmensdorf, Switzerland 4Consortium for Inter-Disciplinary Environmental Research, Stony Brook University *Corresponding author: E-mail: [email protected]. Accepted: May 12, 2019 Data deposition: The raw read data have been deposited in the NCBI Sequence Read Archive under BioProject: PRJNA543673, BioSample: SAMN11774663-SAMN11774674, SRA Study: SRP198856. All scripts used for analyses are available on Dryad: doi:10.5061/dryad.v961mb4. Abstract High-elevation organisms experience shared environmental challenges that include low oxygen availability, cold temperatures, and intense ultraviolet radiation. Consequently, repeated evolution of the same genetic mechanisms may occur across high-elevation taxa. To test this prediction, we investigated the extent to which the same biochemical pathways, genes, or sites were subject to parallel molecular evolution for 12 Andean hummingbird species (family: Trochilidae) representing several independent transitions to high elevation across the phylogeny. Across high-elevation species, we discovered parallel evolution for several pathways and genes with evidence of positive selection. In particular, positively selected genes were frequently part of cellular respiration, metabolism, or cell death pathways. To further examine the role of elevation in our analyses, we compared results for low- and high-elevation species and tested different thresholds for defining elevation categories. -

Association Between Chymase Gene Polymorphisms and Atrial Fibrillation
Zhou et al. BMC Cardiovascular Disorders (2019) 19:321 https://doi.org/10.1186/s12872-019-01300-7 RESEARCH ARTICLE Open Access Association between chymase gene polymorphisms and atrial fibrillation in Chinese Han population Dongchen Zhou, Yuewei Chen, Jiaxin Wu, Jiabo Shen, Yushan Shang, Liangrong Zheng and Xudong Xie* Abstract Background: Chymase is the major angiotensin II (Ang II)-forming enzyme in cardiovascular tissue, with an important role in atrial remodeling. This study aimed to examine the association between chymase 1 gene (CMA1) polymorphisms and atrial fibrillation (AF) in a Chinese Han population. Methods: This case-control study enrolled 126 patients with lone AF and 120 age- and sex-matched healthy controls, all from a Chinese Han population. Five CMA1 polymorphisms were genotyped. Results: The CMA1 polymorphism rs1800875 (G-1903A) was associated with AF. The frequency of the GG genotype was significantly higher in AF patients compared with controls (p = 0.009). Haplotype analysis further demonstrated an increased risk of AF associated with the rs1800875-G haplotype (Hap8 TGTTG, odds ratio (OR) = 1.668, 95% CI 1.132–2.458, p = 0.009), and a decreased risk for the rs1800875-A haplotype (Hap5 TATTG, OR = 0.178, 95% CI 0.042– 0.749, p = 0.008). Conclusions: CMA1 polymorphisms may be associated with AF, and the rs1800875 GG genotype might be a susceptibility factor for AF in the Chinese Han population. Keywords: Atrial fibrillation, CMA1, Chymase, Single nucleotide polymorphism, Angiotensin II Background ‘lone AF’, occur in the absence of identifiable underlying Atrial fibrillation (AF) is the most common type of sus- cardiovascular or other comorbid diseases [4]. -

Inhibiting the Soluble Epoxide Hydrolase Increases the Epfas and ERK1/2 Expression in the Hippocampus of Licl-Pilocarpine Post-Status Epilepticus Rat Model
Inhibiting the soluble epoxide hydrolase increases the EpFAs and ERK1/2 expression in the hippocampus of LiCl-pilocarpine post-status epilepticus rat model Weifeng Peng Zhongshan Hospital Fudan University https://orcid.org/0000-0002-6225-1409 Yijun Shen Zhongshan Hospital Fudan University Qiang Wang Zhongshan Hospital Fudan University Jing Ding ( [email protected] ) Zhongshan Hospital Fudan University Xin Wang ( [email protected] ) Zhongshan Hospital Fudan University Research Article Keywords: epilepsy, soluble epoxide hydrolase, epoxygenated fatty acids (EpFAs), extracellular signal- activated protein kinase 1/2 (ERK1/2) Posted Date: April 15th, 2021 DOI: https://doi.org/10.21203/rs.3.rs-385335/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/20 Abstract Purpose This study aimed to investigate the enzyme activity of soluble epoxide hydrolase (sEH) and quantify metabolic substrates i.e. epoxygenated fatty acids (EpFAs) and products of sEH in the hippocampus after administering TPPU [1-triuoromethoxyphenyl-3-(1-propionylpiperidin-4-yl)urea], the inhibitor of sEH, and further explored whether the extracellular signal-activated protein kinase 1/2 (ERK1/2) was involved in the anti-seizure effect of TPPU in the lithium chloride (LiCl)-pilocarpine induced post-status epilepticus (SE) rat model. Methods The rats were intraperitoneally (I.P.) injected with LiCl and pilocarpine to induce SE and then spontaneous recurrent seizures (SRS) were observed. Rats were randomly assigned into SRS + 0.1 TPPU group (intragastrically administering 0.1 mg/kg/d TPPU), SRS + PEG 400 group (administering the vehicle instead), and Control group. -

Regulation of the Bovine Oviductal Fluid Proteome
REPRODUCTIONRESEARCH Regulation of the bovine oviductal fluid proteome Julie Lamy1, Valérie Labas1,2, Grégoire Harichaux1,2, Guillaume Tsikis1, Pascal Mermillod1 and Marie Saint-Dizier1,3 1Physiologie de la Reproduction et des Comportements (PRC), INRA, CNRS, IFCE, Université de Tours, Nouzilly, France, 2INRA, Plateforme d’Analyse Intégrative des Biomolécules (PAIB), Nouzilly, France and 3Université François Rabelais de Tours, UFR Sciences et Techniques, Tours, France Correspondence should be addressed to M Saint-Dizier; Email: [email protected] Abstract Our objective was to investigate the regulation of the proteome in the bovine oviductal fluid according to the stage of the oestrous cycle, to the side relative to ovulation and to local concentrations of steroid hormones. Luminal fluid samples from both oviducts were collected at four stages of the oestrous cycle: pre-ovulatory (Pre-ov), post-ovulatory (Post-ov), and mid- and late luteal phases from adult cyclic cows (18–25 cows/stage). The proteomes were assessed by nanoLC–MS/MS and quantified by label-free method. Totally, 482 proteins were identified including a limited number of proteins specific to one stage or one side. Proportions of differentially abundant proteins fluctuated from 10 to 24% between sides at one stage and from 4 to 20% among stages in a given side of ovulation. In oviductal fluids ipsilateral to ovulation, Annexin A1 was the most abundant protein at Pre-ov compared with Post-ov while numerous heat shock proteins were more abundant at Post-ov compared with Pre-ov. Among differentially abundant proteins, seven tended to be correlated with intra-oviductal concentrations of progesterone. -

CMA1 Rabbit Polyclonal Antibody – TA342809 | Origene
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for TA342809 CMA1 Rabbit Polyclonal Antibody Product data: Product Type: Primary Antibodies Applications: WB Recommended Dilution: WB Reactivity: Human Host: Rabbit Isotype: IgG Clonality: Polyclonal Immunogen: The immunogen for anti-CMA1 antibody: synthetic peptide directed towards the C terminal of human CMA1. Synthetic peptide located within the following region: EVKLRLMDPQACSHFRDFDHNLQLCVGNPRKTKSAFKGDSGGPLLCAGVA Formulation: Liquid. Purified antibody supplied in 1x PBS buffer with 0.09% (w/v) sodium azide and 2% sucrose. Note that this product is shipped as lyophilized powder to China customers. Conjugation: Unconjugated Storage: Store at -20°C as received. Stability: Stable for 12 months from date of receipt. Predicted Protein Size: 27 kDa Gene Name: chymase 1 Database Link: NP_001827 Entrez Gene 1215 Human P23946 Background: CMA1 is a chymotryptic serine proteinase that belongs to the peptidase family S1. It is expressed in mast cells and thought to function in the degradation of the extracellular matrix, the regulation of submucosal gland secretion, and the generation of vasoactive peptides. In the heart and blood vessels, this protein, rather than angiotensin converting enzyme, is largely responsible for converting angiotensin I to the vasoactive peptide angiotensin II. Angiotensin II has been implicated in blood pressure control and in the pathogenesis of hypertension, cardiac hypertrophy, and heart failure. Thus, this gene product is a target for cardiovascular disease therapies. This product is to be used for laboratory only. Not for diagnostic or therapeutic use. -

Based on Network Pharmacology Tools to Investigate the Molecular Mechanism of Cordyceps Sinensis on the Treatment of Diabetic Nephropathy
Hindawi Journal of Diabetes Research Volume 2021, Article ID 8891093, 12 pages https://doi.org/10.1155/2021/8891093 Research Article Based on Network Pharmacology Tools to Investigate the Molecular Mechanism of Cordyceps sinensis on the Treatment of Diabetic Nephropathy Yan Li,1 Lei Wang,2 Bojun Xu,1 Liangbin Zhao,1 Li Li,1 Keyang Xu,3 Anqi Tang,1 Shasha Zhou,1 Lu Song,1 Xiao Zhang,1 and Huakui Zhan 1 1Hospital of Chengdu University of Traditional Chinese Medicine, Chengdu, 610072 Sichuan, China 2Key Laboratory of Chinese Internal Medicine of Ministry of Education and Dongzhimen Hospital, Beijing University of Chinese Medicine, Beijing 100700, China 3Zhejiang Chinese Medical University, Hangzhou, 310053 Zhejiang, China Correspondence should be addressed to Huakui Zhan; [email protected] Received 27 August 2020; Revised 17 January 2021; Accepted 24 January 2021; Published 8 February 2021 Academic Editor: Michaelangela Barbieri Copyright © 2021 Yan Li et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Background. Diabetic nephropathy (DN) is one of the most common complications of diabetes mellitus and is a major cause of end- stage kidney disease. Cordyceps sinensis (Cordyceps, Dong Chong Xia Cao) is a widely applied ingredient for treating patients with DN in China, while the molecular mechanisms remain unclear. This study is aimed at revealing the therapeutic mechanisms of Cordyceps in DN by undertaking a network pharmacology analysis. Materials and Methods. In this study, active ingredients and associated target proteins of Cordyceps sinensis were obtained via Traditional Chinese Medicine Systems Pharmacology Database (TCMSP) and Swiss Target Prediction platform, then reconfirmed by using PubChem databases. -
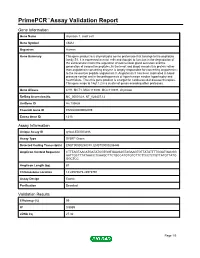
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name chymase 1, mast cell Gene Symbol CMA1 Organism Human Gene Summary This gene product is a chymotryptic serine proteinase that belongs to the peptidase family S1. It is expressed in mast cells and thought to function in the degradation of the extracellular matrix the regulation of submucosal gland secretion and the generation of vasoactive peptides. In the heart and blood vessels this protein rather than angiotensin converting enzyme is largely responsible for converting angiotensin I to the vasoactive peptide angiotensin II. Angiotensin II has been implicated in blood pressure control and in the pathogenesis of hypertension cardiac hypertrophy and heart failure. Thus this gene product is a target for cardiovascular disease therapies. This gene maps to 14q11.2 in a cluster of genes encoding other proteases. Gene Aliases CYH, MCT1, MGC119890, MGC119891, chymase RefSeq Accession No. NC_000014.8, NT_026437.12 UniGene ID Hs.135626 Ensembl Gene ID ENSG00000092009 Entrez Gene ID 1215 Assay Information Unique Assay ID qHsaCED0003486 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000250378, ENST00000206446 Amplicon Context Sequence CTTTAGTAACATGATATCGTGGTGAAGAGTAGAAGTGTTATATTTTGGATGACGG AATTGCTTTATAACCTCAAGCTTCTGCCATGTGTCTTCTTCCTCTGTTATGTTATG GGCTCC Amplicon Length (bp) 87 Chromosome Location 14:24975675-24975791 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 99 R2 0.9989 cDNA Cq 27.32 Page 1/5 PrimePCR™Assay Validation Report cDNA Tm (Celsius) 78 -

Gene Expression Profiling of Lymph Node Sub-Capsular Sinus Macrophages in Cancer
ORIGINAL RESEARCH published: 08 June 2021 doi: 10.3389/fimmu.2021.672123 Gene Expression Profiling of Lymph Node Sub-Capsular Sinus Macrophages in Cancer † † Danilo Pellin 1 , Natalie Claudio 2,3 , Zihan Guo 2,4, Tahereh Ziglari 2 and Ferdinando Pucci 2,3* 1 Gene Therapy Program, Dana-Farber/Boston Children’s Cancer and Blood Disorders Center, Harvard Medical School, Boston, MA, United States, 2 Department of Otolaryngology – Head and Neck Surgery, Oregon Health and Science University, Portland, OR, United States, 3 Department of Cell, Developmental & Cancer Biology, Oregon Health and Science University, Portland, OR, United States, 4 Program in Cancer Biology, Oregon Health and Science University, Portland, OR, United States Lymph nodes are key lymphoid organs collecting lymph fluid and migratory cells from the tissue area they survey. When cancerous cells arise within a tissue, the sentinel lymph node is the first immunological organ to mount an immune response. Sub-capsular sinus Edited by: Karine Rachel Prudent Breckpot, macrophages (SSMs) are specialized macrophages residing in the lymph nodes that play Vrije University Brussel, Belgium important roles as gatekeepers against particulate antigenic material. In the context of Reviewed by: cancer, SSMs capture tumor-derived extracellular vesicles (tEVs), a form of particulate Ioannis S. Pateras, National and Kapodistrian University of antigen released in high amounts by tumor cells. We and others have recently Athens, Greece demonstrated that SSMs possess anti-tumor activity because in their absence tumors Antonio Giovanni Solimando, progress faster. A comprehensive profiling of SSMs represents an important first step to University of Bari Aldo Moro, Italy Rohit Singh, identify the cellular and molecular mechanisms responsible for SSM anti-tumor activity. -

Single Cell Derived Clonal Analysis of Human Glioblastoma Links
SUPPLEMENTARY INFORMATION: Single cell derived clonal analysis of human glioblastoma links functional and genomic heterogeneity ! Mona Meyer*, Jüri Reimand*, Xiaoyang Lan, Renee Head, Xueming Zhu, Michelle Kushida, Jane Bayani, Jessica C. Pressey, Anath Lionel, Ian D. Clarke, Michael Cusimano, Jeremy Squire, Stephen Scherer, Mark Bernstein, Melanie A. Woodin, Gary D. Bader**, and Peter B. Dirks**! ! * These authors contributed equally to this work.! ** Correspondence: [email protected] or [email protected]! ! Supplementary information - Meyer, Reimand et al. Supplementary methods" 4" Patient samples and fluorescence activated cell sorting (FACS)! 4! Differentiation! 4! Immunocytochemistry and EdU Imaging! 4! Proliferation! 5! Western blotting ! 5! Temozolomide treatment! 5! NCI drug library screen! 6! Orthotopic injections! 6! Immunohistochemistry on tumor sections! 6! Promoter methylation of MGMT! 6! Fluorescence in situ Hybridization (FISH)! 7! SNP6 microarray analysis and genome segmentation! 7! Calling copy number alterations! 8! Mapping altered genome segments to genes! 8! Recurrently altered genes with clonal variability! 9! Global analyses of copy number alterations! 9! Phylogenetic analysis of copy number alterations! 10! Microarray analysis! 10! Gene expression differences of TMZ resistant and sensitive clones of GBM-482! 10! Reverse transcription-PCR analyses! 11! Tumor subtype analysis of TMZ-sensitive and resistant clones! 11! Pathway analysis of gene expression in the TMZ-sensitive clone of GBM-482! 11! Supplementary figures and tables" 13" "2 Supplementary information - Meyer, Reimand et al. Table S1: Individual clones from all patient tumors are tumorigenic. ! 14! Fig. S1: clonal tumorigenicity.! 15! Fig. S2: clonal heterogeneity of EGFR and PTEN expression.! 20! Fig. S3: clonal heterogeneity of proliferation.! 21! Fig.