Pyrroloquinoline Quinone - Wikipedia
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Auxiliary Iron–Sulfur Cofactors in Radical SAM Enzymes☆
Biochimica et Biophysica Acta 1853 (2015) 1316–1334 Contents lists available at ScienceDirect Biochimica et Biophysica Acta journal homepage: www.elsevier.com/locate/bbamcr Review Auxiliary iron–sulfur cofactors in radical SAM enzymes☆ Nicholas D. Lanz a, Squire J. Booker a,b,⁎ a Department of Biochemistry and Molecular Biology, The Pennsylvania State University, University Park, PA 16802, United States b Department of Chemistry, The Pennsylvania State University, University Park, PA 16802, United States article info abstract Article history: A vast number of enzymes are now known to belong to a superfamily known as radical SAM, which all contain a Received 19 September 2014 [4Fe–4S] cluster ligated by three cysteine residues. The remaining, unligated, iron ion of the cluster binds in Received in revised form 15 December 2014 contact with the α-amino and α-carboxylate groups of S-adenosyl-L-methionine (SAM). This binding mode Accepted 6 January 2015 facilitates inner-sphere electron transfer from the reduced form of the cluster into the sulfur atom of SAM, Available online 15 January 2015 resulting in a reductive cleavage of SAM to methionine and a 5′-deoxyadenosyl radical. The 5′-deoxyadenosyl Keywords: radical then abstracts a target substrate hydrogen atom, initiating a wide variety of radical-based transforma- – Radical SAM tions. A subset of radical SAM enzymes contains one or more additional iron sulfur clusters that are required Iron–sulfur cluster for the reactions they catalyze. However, outside of a subset of sulfur insertion reactions, very little is known S-adenosylmethionine about the roles of these additional clusters. This review will highlight the most recent advances in the identifica- Cofactor maturation tion and characterization of radical SAM enzymes that harbor auxiliary iron–sulfur clusters. -

Reduction by the Methylreductase System in Methanobacterium Bryantii WILLIAM B
JOURNAL OF BACTERIOLOGY, Jan. 1987, p. 87-92 Vol. 169, No. 1 0021-9193/87/010087-06$02.00/0 Copyright © 1987, American Society for Microbiology Inhibition by Corrins of the ATP-Dependent Activation and CO2 Reduction by the Methylreductase System in Methanobacterium bryantii WILLIAM B. WHITMAN'* AND RALPH S. WOLFE2 Department of Microbiology, University of Georgia, Athens, Georgia 30602,1 and Department of Microbiology, University ofIllinois, Urbana, Illinois 618012 Received 1 August 1986/Accepted 28 September 1986 Corrins inhibited the ATP-dependent activation of the methylreductase system and the methyl coenzyme M-dependent reduction of CO2 in extracts of Methanobacterium bryantii resolved from low-molecular-weight factors. The concentrations of cobinamides and cobamides required for one-half of maximal inhibition of the ATP-depen4ent activation were between 1 and 5 ,M. Cobinamides were more inhibitory at lower concentra- tiops than cobamides. Deoxyadenosylcobalamin was not inhibitory at concentrations up to 25 ,uM. The inhibition of CO2 reduction was competitive with respect to CO2. The concentration of methylcobalamin required for one-half of maximal inhibition was 5 ,M. Other cobamideg inhibited at similar concentrations, but diaquacobinami4e inhibited at lower concentrations. With respect to their affinities and specificities for corrins, inhibition of both the ATP-dependent activation'and CO2 reduction closely resembled the corrin- dependent activation of the methylreductase described in similar extracts (W. B. Whitman and R. S. Wolfe, J. Bacteriol. 164:165-172, 1985). However, whether the multiple effects of corrins are due to action at a single site is unknown. The effect of corrins (cobamides and cobinamides) on in CO2 reduction. -
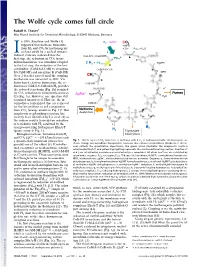
The Wolfe Cycle Comes Full Circle
The Wolfe cycle comes full circle Rudolf K. Thauer1 Max Planck Institute for Terrestrial Microbiology, D-35043 Marburg, Germany n 1988, Rouvière and Wolfe (1) H - ΔμNa+ 2 CO2 suggested that methane formation + MFR from H and CO by methanogenic + 2H+ *Fd + H O I 2 2 ox 2 archaea could be a cyclical process. j O = Indirect evidence indicated that the CoB-SH + CoM-SH fi *Fd 2- a rst step, the reduction of CO2 to for- red R mylmethanofuran, was somehow coupled + * H MPT 2 H2 Fdox 4 to the last step, the reduction of the het- h erodisulfide (CoM-S-S-CoB) to coenzyme CoM-S-S-CoB b MFR M (CoM-SH) and coenzyme B (CoB-SH). H Over 2 decades passed until the coupling C 4 10 mechanism was unraveled in 2011: Via g flavin-based electron bifurcation, the re- CoB-SH duction of CoM-S-S-CoB with H provides 2 H+ the reduced ferredoxin (Fig. 1h) required c + Purines for CO2 reduction to formylmethanofuran ΔμNa + H MPT 4 f H O (2) (Fig. 1a). However, one question still 2 remained unanswered: How are the in- termediates replenished that are removed CoM-SH for the biosynthesis of cell components H Methionine d from CO2 (orange arrows in Fig. 1)? This Acetyl-CoA e anaplerotic (replenishing) reaction has F420 F420H2 recently been identified by Lie et al. (3) as F420 F420H2 the sodium motive force-driven reduction H i of ferredoxin with H2 catalyzed by the i energy-converting hydrogenase EhaA-T H2 (green arrow in Fig. -

Annotation Guidelines for Experimental Procedures
Annotation Guidelines for Experimental Procedures Developed By Mohammed Alliheedi Robert Mercer Version 1 April 14th, 2018 1- Introduction and background information What is rhetorical move? A rhetorical move can be defined as a text fragment that conveys a distinct communicative goal, in other words, a sentence that implies an author’s specific purpose to readers. What are the types of rhetorical moves? There are several types of rhetorical moves. However, we are interested in 4 rhetorical moves that are common in the method section of a scientific article that follows the Introduction Methods Results and Discussion (IMRaD) structure. 1- Description of a method: It is concerned with a sentence(s) that describes experimental events (e.g., “Beads with bound proteins were washed six times (for 10 min under rotation at 4°C) with pulldown buffer and proteins harvested in SDS-sample buffer, separated by SDS-PAGE, and analyzed by autoradiography.” (Ester & Uetz, 2008)). 2- Appeal to authority: It is concerned with a sentence(s) that discusses the use of standard methods, protocols, and procedures. There are two types of this move: - A reference to a well-established “standard” method (e.g., the use of a method like “PCR” or “electrophoresis”). - A reference to a method that was previously described in the literature (e.g., “Protein was determined using fluorescamine assay [41].” (Larsen, Frandesn and Treiman, 2001)). 3- Source of materials: It is concerned with a sentence(s) that lists the source of biological materials that are used in the experiment (e.g., “All microalgal strains used in this study are available at the Elizabeth Aidar Microalgae Culture Collection, Department of Marine Biology, Federal Fluminense University, Brazil.” (Larsen, Frandesn and Treiman, 2001)). -

Supplemental Methods
Supplemental Methods: Sample Collection Duplicate surface samples were collected from the Amazon River plume aboard the R/V Knorr in June 2010 (4 52.71’N, 51 21.59’W) during a period of high river discharge. The collection site (Station 10, 4° 52.71’N, 51° 21.59’W; S = 21.0; T = 29.6°C), located ~ 500 Km to the north of the Amazon River mouth, was characterized by the presence of coastal diatoms in the top 8 m of the water column. Sampling was conducted between 0700 and 0900 local time by gently impeller pumping (modified Rule 1800 submersible sump pump) surface water through 10 m of tygon tubing (3 cm) to the ship's deck where it then flowed through a 156 µm mesh into 20 L carboys. In the lab, cells were partitioned into two size fractions by sequential filtration (using a Masterflex peristaltic pump) of the pre-filtered seawater through a 2.0 µm pore-size, 142 mm diameter polycarbonate (PCTE) membrane filter (Sterlitech Corporation, Kent, CWA) and a 0.22 µm pore-size, 142 mm diameter Supor membrane filter (Pall, Port Washington, NY). Metagenomic and non-selective metatranscriptomic analyses were conducted on both pore-size filters; poly(A)-selected (eukaryote-dominated) metatranscriptomic analyses were conducted only on the larger pore-size filter (2.0 µm pore-size). All filters were immediately submerged in RNAlater (Applied Biosystems, Austin, TX) in sterile 50 mL conical tubes, incubated at room temperature overnight and then stored at -80oC until extraction. Filtration and stabilization of each sample was completed within 30 min of water collection. -

Hydrogenases of Methanogens
ANRV413-BI79-18 ARI 27 April 2010 21:0 Hydrogenases from Methanogenic Archaea, Nickel, a Novel Cofactor, and H2 Storage Rudolf K. Thauer, Anne-Kristin Kaster, Meike Goenrich, Michael Schick, Takeshi Hiromoto, and Seigo Shima Max Planck Institute for Terrestrial Microbiology, D-35043 Marburg, Germany; email: [email protected] Annu. Rev. Biochem. 2010. 79:507–36 Key Words First published online as a Review in Advance on H2 activation, energy-converting hydrogenase, complex I of the March 17, 2010 respiratory chain, chemiosmotic coupling, electron bifurcation, The Annual Review of Biochemistry is online at reversed electron transfer biochem.annualreviews.org This article’s doi: Abstract 10.1146/annurev.biochem.030508.152103 Most methanogenic archaea reduce CO2 with H2 to CH4. For the Copyright c 2010 by Annual Reviews. activation of H2, they use different [NiFe]-hydrogenases, namely All rights reserved energy-converting [NiFe]-hydrogenases, heterodisulfide reductase- 0066-4154/10/0707-0507$20.00 associated [NiFe]-hydrogenase or methanophenazine-reducing by University of Texas - Austin on 06/10/13. For personal use only. [NiFe]-hydrogenase, and F420-reducing [NiFe]-hydrogenase. The energy-converting [NiFe]-hydrogenases are phylogenetically related Annu. Rev. Biochem. 2010.79:507-536. Downloaded from www.annualreviews.org to complex I of the respiratory chain. Under conditions of nickel limitation, some methanogens synthesize a nickel-independent [Fe]- hydrogenase (instead of F420-reducing [NiFe]-hydrogenase) and by that reduce their nickel requirement. The [Fe]-hydrogenase harbors a unique iron-guanylylpyridinol cofactor (FeGP cofactor), in which a low-spin iron is ligated by two CO, one C(O)CH2-, one S-CH2-, and a sp2-hybridized pyridinol nitrogen. -

Xuejun Yu Dissertation Final
UNIVERSITY OF CALIFORNIA RIVERSIDE Conversion of Carbon Dioxide to Formate by a Formate Dehydrogenase from Cupriavidus necator A Dissertation submitted in partial satisfaction of the requirements for the degree of Doctor of Philosophy in Bioengineering by Xuejun Yu September 2018 Dissertation Committee: Dr. Ashok Mulchandani, Co-Chairperson Dr. Xin Ge, Co-Chairperson Dr. Russ Hille Copyright by Xuejun Yu 2018 The Dissertation of Xuejun Yu is approved: Committee Co-Chairperson Committee Co-Chairperson University of California, Riverside ACKNOWLEDGEMENTS I would like to express sincere appreciation to my advisor, Professor Ashok Mulchandani, for accepting me to be his student when I was suffering. Thank you very much for your delicate guidance, support and encouragement during the past years. You not only guide me on my PhD study, but also help me to be mature on my personality. From bottom of my heart, I feel very lucky to have you as professor. Also, many thanks go to my other committee members. Professor Xin Ge acted as my co-advisor and provided many valuable comments on my molecular cloning work. Professor Russ Hille has also provided insights, encouragements and advices as a member of my committee members. I have learned so many important insights from our meetings and discussions and thank you for bringing me into the molybdenum/tungsten enzyme conference. I am also thankful to the past and current group members, colleagues and friends - Dimitri Niks, Pankaj Ramnani, Feng Tan, Rabeay Hassan, Trupti Terse, Thien-Toan Tran, Claudia Chaves, Jia-wei Tay, Hui Wang, Pham Tung, Hilda Chan, Yingning Gao and Tynan Young. -

192ICM ICBIC Posters
Journal of Inorganic Biochemistry 96 (2003) 203 Monomeric TpPrMoVOSR complexes via the chemical reduction of TpPrMoVIOSR. David J Nielsen, School of Chemistry, University of Melbourne, Australia Christian J Doonan, School of Chemistry, University of Melbourne, Australia Graham N George, Stanford Synchrotron Radiation Laboratory, United States Hugh Harris, Stanford Synchrotron Radiation Laboratory, United States Charles G Young, University of Melbourne, Australia EPR evidence has suggested the presence of molybdenum(V) intermediates in the catalytic cycle of hydroxylase enzyme systems [1], and references therein], and as such these species are attractive targets for the synthesis of small-molecule model systems. Ongoing work in our group has allowed access to several stable and well characterised monomeric molybdenum(VI) oxo-thio complexes TpPrMoVIOSR (TpPr = hydridotris(3-isopropylpyrazol-1-yl)borate) with co-ligand R = eg. substituted phenolates [2], as shown below. These Mo(VI) complexes have proved amenable to chemical reduction using cobaltocene (CoCp2) yielding initially the Pr V [CoCp2][Tp Mo OSR] salts [1,2]. Solution and solid state sulfur X-ray absorption spectroscopy (XAS) on selected examples of the chemically reduced species shows pre-edge features attributable to the S 1s → Mo=S π* transition of a [MoOS]+ unit. Further spectroscopic investigations (EPR, IR) are consistent with the presence of a paramagnetic Mo(V) centre bearing a terminal thio ligand. Continuing spectroscopic, structural and reactivity investigations centred on these important species will be presented. References: [1] P. D. Smith, D. A. Slizys, G. N. George and C. G. Young, J. Amer. Chem. Soc., 122(12), 2000, 2946. [2] C. J. Doonan, Unpublished results. -

Van Heuvelen Department of Chemistry, Harvey Mudd College
Development of Bio-Inspired Catalysts for Dechlorination Reactions Prof. Katherine Van Heuvelen Department of Chemistry, Harvey Mudd College Abstract The nickel-containing cofactor F430 found in methyl-coenzyme M reductase (MCR) and the cobalt-containing cobalamin cofactor (Cbl) found in Vitamin B12 carry out the reductive dehalogenation of chlorinated alkenes, which can act as damaging pollutants in the environment. Both F430 and Cbl are found in biological systems and carry out this reaction under benign conditions using earth-abundant materials. This work centers on the preparation and investigation of small molecular model compounds that reproduce key geometric and electronic features of cofactors F430 and Cbl. In particular, I propose to: 1. Prepare a series of nickel- and cobalt-containing F430 model compounds designed to investigate the influence of the supporting ligand on reactivity. 2. Evaluate the reactivity of these complexes towards halogenated substrates. 3. Characterize reaction intermediates using a combination of spectroscopic (UV-visible, infrared, NMR) and computational (density functional theory, DFT) techniques in order to correlate geometric and electronic structure with reactivity. 4. Elucidate the reaction mechanism using insights gained from aims 1–3, ultimately applying a detailed understanding of the fundamental chemistry underlying dehalogenation to the rational design of an improved catalytic system to treat chlorinated pollutants before they enter the water supply. Start Date, Duration, and Location This research will be conducted at Harvey Mudd College over a ten-week period in the summer of 2016, which will run from May 23 – July 29. The Chemistry Department is in the process of hiring students for the summer of 2016 and the student working on this project will be identified later in the spring semester. -

Deciphering the Mechanism of Enzymatic Methane Synthesis
Science Highlight – June 2021 Deciphering the Mechanism of Enzymatic Methane Synthesis Methane is the simplest organic compound with the highest energy content of any carbon- based fuel. Methane accounts for almost a quarter of U.S. energy consumption, with one-half of homes using natural gas as their heating fuel.1-2 Interestingly, more than 90% of all methane on earth is produced biogenically by methanogens which are responsible for enzymatic synthesis of 1 billion tons of methane per year.3-4 Much of the methane formed by methanogens is captured and used as an energy source by aerobic and anaerobic methanotrophic microbes.5-6 However, the increased mining of methane and industrial farming of cattle has created a mismatch between the sources and the sinks of methane, causing its atmospheric levels to double over the past two centuries. This is an environmental concern related to climate change because pound for pound, methane causes 25 times more 7 global warming over a 100-year period than CO2. Thus, understanding the biosynthesis of methane is imperative from basic energy, economic and environmental perspectives. Methane is formed by one of the few Ni containing cofactors (F430) as part of the active site of methyl coenzyme M reductase (MCR), a strictly anaerobic enzyme present in methanogenic archaea. MCR catalyzes the reaction of methyl-coenzyme M (CH3−SCoM) with coenzyme B (HSCoB) to form methane and the heterodisulfide CoMS−SCoB.8 Besides the importance of methane metabolism in our environment and global energy landscape, the basic chemistry and biology of alkane activation and formation by MCR are of fundamental interest. -

Changes in Soil Microbial Communities After Long-Term Warming Exposure" (2019)
University of Massachusetts Amherst ScholarWorks@UMass Amherst Doctoral Dissertations Dissertations and Theses October 2019 CHANGES IN SOIL MICROBIAL COMMUNITIES AFTER LONG- TERM WARMING EXPOSURE William G. Rodríguez-Reillo University of Massachusetts Amherst Follow this and additional works at: https://scholarworks.umass.edu/dissertations_2 Part of the Environmental Microbiology and Microbial Ecology Commons Recommended Citation Rodríguez-Reillo, William G., "CHANGES IN SOIL MICROBIAL COMMUNITIES AFTER LONG-TERM WARMING EXPOSURE" (2019). Doctoral Dissertations. 1757. https://doi.org/10.7275/15007293 https://scholarworks.umass.edu/dissertations_2/1757 This Open Access Dissertation is brought to you for free and open access by the Dissertations and Theses at ScholarWorks@UMass Amherst. It has been accepted for inclusion in Doctoral Dissertations by an authorized administrator of ScholarWorks@UMass Amherst. For more information, please contact [email protected]. CHANGES IN SOIL MICROBIAL COMMUNITIES AFTER LONG-TERM WARMING EXPOSURE A Dissertation Presented by WILLIAM GABRIEL RODRÍGUEZ-REILLO Submitted to the Graduate School of the University of Massachusetts Amherst in partial fulfillment of the requirements for the degree of DOCTOR OF PHILOSOPHY SEPTEMBER 2019 Organismic and Evolutionary Biology © Copyright by William Gabriel Rodríguez-Reillo 2019 All Rights Reserved CHANGES IN SOIL MICROBIAL COMMUNITIES AFTER LONG-TERM WARMING EXPOSURE A Dissertation Presented by WILLIAM GABRIEL RODRÍGUEZ-REILLO Approved as to style and content by: _________________________________________ Jeffrey L. Blanchard, Chair _________________________________________ Courtney Babbitt, Member _________________________________________ David Sela, Member _________________________________________ Kristina Stinson, Member ______________________________________ Paige Warren, Graduate Program Director Organismic and Evolutionary Biology DEDICATION To my parents, William Rodriguez Arce and Carmen L. Reillo Batista. A quienes aún en la distancia me mantuvieron en sus oraciones. -

Characterisation, Classification and Conformational Variability Of
Characterisation, Classification and Conformational Variability of Organic Enzyme Cofactors Julia D. Fischer European Bioinformatics Institute Clare Hall College University of Cambridge A thesis submitted for the degree of Doctor of Philosophy 11 April 2011 This dissertation is the result of my own work and includes nothing which is the outcome of work done in collaboration except where specifically indicated in the text. This dissertation does not exceed the word limit of 60,000 words. Acknowledgements I would like to thank all the members of the Thornton research group for their constant interest in my work, their continuous willingness to answer my academic questions, and for their company during my time at the EBI. This includes Saumya Kumar, Sergio Martinez Cuesta, Matthias Ziehm, Dr. Daniela Wieser, Dr. Xun Li, Dr. Irene Pa- patheodorou, Dr. Pedro Ballester, Dr. Abdullah Kahraman, Dr. Rafael Najmanovich, Dr. Tjaart de Beer, Dr. Syed Asad Rahman, Dr. Nicholas Furnham, Dr. Roman Laskowski and Dr. Gemma Holli- day. Special thanks to Asad for allowing me to use early development versions of his SMSD software and for help and advice with the KEGG API installation, to Roman for knowing where to find all kinds of data, to Dani for help with R scripts, to Nick for letting me use his E.C. tree program, to Tjaart for python advice and especially to Gemma for her constant advice and feedback on my work in all aspects, in particular the chemistry side. Most importantly, I would like to thank Prof. Janet Thornton for giving me the chance to work on this project, for all the time she spent in meetings with me and reading my work, for sharing her seemingly limitless knowledge and enthusiasm about the fascinating world of enzymes, and for being such an experienced and motivational advisor.