Methyl-Cpg-Binding Domain 9 (MBD9) Is Required for H2A.Z Incorporation Into Chromatin at a Subset of H2A.Z-Enriched Regions in the Arabidopsis Genome
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Bromodomain Factors of BET Family Are New Essential Actors Of
Bromodomain factors of BET family are new essential actors of pericentric heterochromatin transcriptional activation in response to heat shock Edwige Col, Neda Hoghoughi, Solenne Dufour, Jessica Penin, Sivan Koskas, Virginie Faure, Maria Ouzounova, Hector Hernandez-Vargash, Nicolas Reynoird, Sylvain Daujat, et al. To cite this version: Edwige Col, Neda Hoghoughi, Solenne Dufour, Jessica Penin, Sivan Koskas, et al.. Bromodomain factors of BET family are new essential actors of pericentric heterochromatin transcriptional acti- vation in response to heat shock. Scientific Reports, Nature Publishing Group, 2017, 7, pp.5418. 10.1038/s41598-017-05343-8. hal-02989794 HAL Id: hal-02989794 https://hal.archives-ouvertes.fr/hal-02989794 Submitted on 5 Nov 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. www.nature.com/scientificreports OPEN Bromodomain factors of BET family are new essential actors of pericentric heterochromatin Received: 5 August 2016 Accepted: 30 May 2017 transcriptional activation in Published: xx xx xxxx response to heat shock Edwige Col1, Neda Hoghoughi1, Solenne Dufour1, Jessica Penin1, Sivan Koskas1, Virginie Faure1, Maria Ouzounova2, Hector Hernandez-Vargash2, Nicolas Reynoird1, Sylvain Daujat4, Eric Folco1, Marc Vigneron3, Robert Schneider4,5, André Verdel1, Saadi Khochbin1, Zdenko Herceg2, Cécile Caron1 & Claire Vourc’h1 The heat shock response is characterized by the transcriptional activation of both hsp genes and noncoding and repeated satellite III DNA sequences located at pericentric heterochromatin. -

Functional Roles of Bromodomain Proteins in Cancer
cancers Review Functional Roles of Bromodomain Proteins in Cancer Samuel P. Boyson 1,2, Cong Gao 3, Kathleen Quinn 2,3, Joseph Boyd 3, Hana Paculova 3 , Seth Frietze 3,4,* and Karen C. Glass 1,2,4,* 1 Department of Pharmaceutical Sciences, Albany College of Pharmacy and Health Sciences, Colchester, VT 05446, USA; [email protected] 2 Department of Pharmacology, Larner College of Medicine, University of Vermont, Burlington, VT 05405, USA; [email protected] 3 Department of Biomedical and Health Sciences, University of Vermont, Burlington, VT 05405, USA; [email protected] (C.G.); [email protected] (J.B.); [email protected] (H.P.) 4 University of Vermont Cancer Center, Burlington, VT 05405, USA * Correspondence: [email protected] (S.F.); [email protected] (K.C.G.) Simple Summary: This review provides an in depth analysis of the role of bromodomain-containing proteins in cancer development. As readers of acetylated lysine on nucleosomal histones, bromod- omain proteins are poised to activate gene expression, and often promote cancer progression. We examined changes in gene expression patterns that are observed in bromodomain-containing proteins and associated with specific cancer types. We also mapped the protein–protein interaction network for the human bromodomain-containing proteins, discuss the cellular roles of these epigenetic regu- lators as part of nine different functional groups, and identify bromodomain-specific mechanisms in cancer development. Lastly, we summarize emerging strategies to target bromodomain proteins in cancer therapy, including those that may be essential for overcoming resistance. Overall, this review provides a timely discussion of the different mechanisms of bromodomain-containing pro- Citation: Boyson, S.P.; Gao, C.; teins in cancer, and an updated assessment of their utility as a therapeutic target for a variety of Quinn, K.; Boyd, J.; Paculova, H.; cancer subtypes. -

Histone-Binding of DPF2 Mediates Its Repressive Role in Myeloid Differentiation
Histone-binding of DPF2 mediates its repressive role in myeloid differentiation Ferdinand M. Hubera,1, Sarah M. Greenblattb,1, Andrew M. Davenporta,1, Concepcion Martinezb,YeXub,LyP.Vuc, Stephen D. Nimerb,2, and André Hoelza,2 aDivision of Chemistry and Chemical Engineering, California Institute of Technology, Pasadena, CA 91125; bSylvester Comprehensive Cancer Center, University of Miami Miller School of Medicine, Miami, FL 33136; and cMolecular Pharmacology Program, Memorial Sloan Kettering Cancer Center, New York, NY 10065 Edited by Douglas C. Rees, Howard Hughes Medical Institute, California Institute of Technology, Pasadena, CA, and approved April 26, 2017 (received for review January 6, 2017) Double plant homeodomain finger 2 (DPF2) is a highly evolution- RUNX1 form a methylation-dependent repressive complex in arily conserved member of the d4 protein family that is ubiqui- AML, although it remains unclear whether the two proteins bind tously expressed in human tissues and was recently shown to each other directly or act concertedly as part of a larger complex. inhibit the myeloid differentiation of hematopoietic stem/progen- Here, we present the crystal structure of the human DPF2 itor and acute myelogenous leukemia cells. Here, we present the tandem PHD finger domain at a 1.6-Å resolution. We demon- crystal structure of the tandem plant homeodomain finger domain strate that the DPF2 tandem PHD finger domain binds acetylated of human DPF2 at 1.6-Å resolution. We show that DPF2 interacts H3 and H4 histone tails, identify the primary determinants of with the acetylated tails of both histones 3 and 4 via bipartite histone recognition, and confirm these interactions in vivo. -

Function of Bromodomain and Extra-Terminal Motif Proteins (Bets) in Gata1-Mediated Transcription
University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations 2015 Function of Bromodomain and Extra-Terminal Motif Proteins (bets) in Gata1-Mediated Transcription Aaron James Stonestrom University of Pennsylvania, [email protected] Follow this and additional works at: https://repository.upenn.edu/edissertations Part of the Molecular Biology Commons, and the Pharmacology Commons Recommended Citation Stonestrom, Aaron James, "Function of Bromodomain and Extra-Terminal Motif Proteins (bets) in Gata1-Mediated Transcription" (2015). Publicly Accessible Penn Dissertations. 1148. https://repository.upenn.edu/edissertations/1148 This paper is posted at ScholarlyCommons. https://repository.upenn.edu/edissertations/1148 For more information, please contact [email protected]. Function of Bromodomain and Extra-Terminal Motif Proteins (bets) in Gata1-Mediated Transcription Abstract Bromodomain and Extra-Terminal motif proteins (BETs) associate with acetylated histones and transcription factors. While pharmacologic inhibition of this ubiquitous protein family is an emerging therapeutic approach for neoplastic and inflammatory disease, the mechanisms through which BETs act remain largely uncharacterized. Here we explore the role of BETs in the physiologically relevant context of erythropoiesis driven by the transcription factor GATA1. First, we characterize functions of the BET family as a whole using a pharmacologic approach. We find that BETs are broadly required for GATA1-mediated transcriptional activation, but that repression is largely BET-independent. BETs support activation by facilitating both GATA1 occupancy and transcription downstream of its binding. Second, we test the specific olesr of BETs BRD2, BRD3, and BRD4 in GATA1-activated transcription. BRD2 and BRD4 are required for efficient anscriptionaltr activation by GATA1. Despite co-localizing with the great majority of GATA1 binding sites, we find that BRD3 is not equirr ed for GATA1-mediated transcriptional activation. -

Download Product Insert (PDF)
PRODUCT INFORMATION BRD3 bromodomains 1 and 2 (human, recombinant) Item No. 14864 Overview and Properties Synonyms: Bromodomain containing protein 3, ORFX, RING3L, RING3-like protein Source: Recombinant N-terminal GST-tagged protein expressed in E. coli Amino Acids: 2-434 Uniprot No.: Q15059 Molecular Weight: 75.0 kDa Storage: -80°C (as supplied) Stability: ≥2 years Purity: batch specific (≥85% estimated by SDS-PAGE) Supplied in: 50 mM Tris, pH 8.0, with 150 mM sodium chloride and 20% glycerol Protein Concentration: batch specific mg/ml Activity: batch specific U/ml Specific Activity: batch specific U/mg Information represents the product specifications. Batch specific analytical results are provided on each certificate of analysis. Image 1 2 3 4 · · · · · · ·250 kDa · · · · · · ·150 kDa · · · · · · ·100 kDa · · · · · · ·75 kDa · · · · · · ·50 kDa · · · · · · ·37 kDa · · · · · · ·25 kDa · · · · · · ·20 kDa · · · · · · ·15 kDa Lane 1: BRD3 (4 µg) Lane 2: BRD3 (6 µg) Lane 3: BRD3 (8 µg) Lane 4: MW Markers WARNING CAYMAN CHEMICAL THIS PRODUCT IS FOR RESEARCH ONLY - NOT FOR HUMAN OR VETERINARY DIAGNOSTIC OR THERAPEUTIC USE. 1180 EAST ELLSWORTH RD SAFETY DATA ANN ARBOR, MI 48108 · USA This material should be considered hazardous until further information becomes available. Do not ingest, inhale, get in eyes, on skin, or on clothing. Wash thoroughly after handling. Before use, the user must review the complete Safety Data Sheet, which has been sent via email to your institution. PHONE: [800] 364-9897 WARRANTY AND LIMITATION OF REMEDY [734] 971-3335 Buyer agrees to purchase the material subject to Cayman’s Terms and Conditions. Complete Terms and Conditions including Warranty and Limitation of Liability information can be found on our website. -

(P -Value<0.05, Fold Change≥1.4), 4 Vs. 0 Gy Irradiation
Table S1: Significant differentially expressed genes (P -Value<0.05, Fold Change≥1.4), 4 vs. 0 Gy irradiation Genbank Fold Change P -Value Gene Symbol Description Accession Q9F8M7_CARHY (Q9F8M7) DTDP-glucose 4,6-dehydratase (Fragment), partial (9%) 6.70 0.017399678 THC2699065 [THC2719287] 5.53 0.003379195 BC013657 BC013657 Homo sapiens cDNA clone IMAGE:4152983, partial cds. [BC013657] 5.10 0.024641735 THC2750781 Ciliary dynein heavy chain 5 (Axonemal beta dynein heavy chain 5) (HL1). 4.07 0.04353262 DNAH5 [Source:Uniprot/SWISSPROT;Acc:Q8TE73] [ENST00000382416] 3.81 0.002855909 NM_145263 SPATA18 Homo sapiens spermatogenesis associated 18 homolog (rat) (SPATA18), mRNA [NM_145263] AA418814 zw01a02.s1 Soares_NhHMPu_S1 Homo sapiens cDNA clone IMAGE:767978 3', 3.69 0.03203913 AA418814 AA418814 mRNA sequence [AA418814] AL356953 leucine-rich repeat-containing G protein-coupled receptor 6 {Homo sapiens} (exp=0; 3.63 0.0277936 THC2705989 wgp=1; cg=0), partial (4%) [THC2752981] AA484677 ne64a07.s1 NCI_CGAP_Alv1 Homo sapiens cDNA clone IMAGE:909012, mRNA 3.63 0.027098073 AA484677 AA484677 sequence [AA484677] oe06h09.s1 NCI_CGAP_Ov2 Homo sapiens cDNA clone IMAGE:1385153, mRNA sequence 3.48 0.04468495 AA837799 AA837799 [AA837799] Homo sapiens hypothetical protein LOC340109, mRNA (cDNA clone IMAGE:5578073), partial 3.27 0.031178378 BC039509 LOC643401 cds. [BC039509] Homo sapiens Fas (TNF receptor superfamily, member 6) (FAS), transcript variant 1, mRNA 3.24 0.022156298 NM_000043 FAS [NM_000043] 3.20 0.021043295 A_32_P125056 BF803942 CM2-CI0135-021100-477-g08 CI0135 Homo sapiens cDNA, mRNA sequence 3.04 0.043389246 BF803942 BF803942 [BF803942] 3.03 0.002430239 NM_015920 RPS27L Homo sapiens ribosomal protein S27-like (RPS27L), mRNA [NM_015920] Homo sapiens tumor necrosis factor receptor superfamily, member 10c, decoy without an 2.98 0.021202829 NM_003841 TNFRSF10C intracellular domain (TNFRSF10C), mRNA [NM_003841] 2.97 0.03243901 AB002384 C6orf32 Homo sapiens mRNA for KIAA0386 gene, partial cds. -

Discovery of a Hidden Transient State in All Bromodomain Families
Discovery of a hidden transient state in all bromodomain families Lluís Raicha, Katharina Meierb, Judith Güntherc, Clara D. Christc, Frank Noéa,d,1, and Simon Olssona,1,2 aDepartment of Mathematics and Computer Science, Freie Universität Berlin, 14195 Berlin, Germany; bComputational Molecular Design, Pharmaceuticals, R&D, Bayer AG, 42096 Wuppertal, Germany; cComputational Molecular Design, Pharmaceuticals, R&D, Bayer AG, 13342 Berlin, Germany; and dDepartment of Chemistry, Rice University, Houston, TX 77005 Edited by David Baker, University of Washington, Seattle, WA, and approved December 1, 2020 (received for review August 17, 2020) Bromodomains (BDs) are small protein modules that interact with comprised by a four-helix bundle (αZ, αA, αB,andαC) with two acetylated marks in histones. These posttranslational modifica- principal loops that connect them (ZA- and BC-loops), con- tions are pivotal to regulate gene expression, making BDs prom- forming a hydrophobic pocket that is suited for acetyl-lysine ising targets to treat several diseases. While the general structure binding (Fig. 1) (24). The BC-loop contains a well-conserved hy- of BDs is well known, their dynamical features and their interplay drogen bond donor or “reader”—usually an asparagine—that is with other macromolecules are poorly understood, hampering the key for recognition. The ZA-loop is the longest loop and the one rational design of potent and selective inhibitors. Here, we com- that displays more variability in terms of sequence and structure, bine extensive molecular dynamics simulations, Markov state presenting short helices, turns, and even hairpin insertions in BD modeling, and available structural data to reveal a transiently family VIII (Fig. -

Mechanisms Underlying Phenotypic Heterogeneity in Simplex Autism Spectrum Disorders
Mechanisms Underlying Phenotypic Heterogeneity in Simplex Autism Spectrum Disorders Andrew H. Chiang Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy under the Executive Committee of the Graduate School of Arts and Sciences COLUMBIA UNIVERSITY 2021 © 2021 Andrew H. Chiang All Rights Reserved Abstract Mechanisms Underlying Phenotypic Heterogeneity in Simplex Autism Spectrum Disorders Andrew H. Chiang Autism spectrum disorders (ASD) are a group of related neurodevelopmental diseases displaying significant genetic and phenotypic heterogeneity. Despite recent progress in ASD genetics, the nature of phenotypic heterogeneity across probands is not well understood. Notably, likely gene- disrupting (LGD) de novo mutations affecting the same gene often result in substantially different ASD phenotypes. We find that truncating mutations in a gene can result in a range of relatively mild decreases (15-30%) in gene expression due to nonsense-mediated decay (NMD), and show that more severe autism phenotypes are associated with greater decreases in expression. We also find that each gene with recurrent ASD mutations can be described by a parameter, phenotype dosage sensitivity (PDS), which characteriZes the relationship between changes in a gene’s dosage and changes in a given phenotype. Using simple linear models, we show that changes in gene dosage account for a substantial fraction of phenotypic variability in ASD. We further observe that LGD mutations affecting the same exon frequently lead to strikingly similar phenotypes in unrelated ASD probands. These patterns are observed for two independent proband cohorts and multiple important ASD-associated phenotypes. The observed phenotypic similarities are likely mediated by similar changes in gene dosage and similar perturbations to the relative expression of splicing isoforms. -

Identification of Multiple Risk Loci and Regulatory Mechanisms Influencing Susceptibility to Multiple Myeloma
Corrected: Author correction ARTICLE DOI: 10.1038/s41467-018-04989-w OPEN Identification of multiple risk loci and regulatory mechanisms influencing susceptibility to multiple myeloma Molly Went1, Amit Sud 1, Asta Försti2,3, Britt-Marie Halvarsson4, Niels Weinhold et al.# Genome-wide association studies (GWAS) have transformed our understanding of susceptibility to multiple myeloma (MM), but much of the heritability remains unexplained. 1234567890():,; We report a new GWAS, a meta-analysis with previous GWAS and a replication series, totalling 9974 MM cases and 247,556 controls of European ancestry. Collectively, these data provide evidence for six new MM risk loci, bringing the total number to 23. Integration of information from gene expression, epigenetic profiling and in situ Hi-C data for the 23 risk loci implicate disruption of developmental transcriptional regulators as a basis of MM susceptibility, compatible with altered B-cell differentiation as a key mechanism. Dysregu- lation of autophagy/apoptosis and cell cycle signalling feature as recurrently perturbed pathways. Our findings provide further insight into the biological basis of MM. Correspondence and requests for materials should be addressed to K.H. (email: [email protected]) or to B.N. (email: [email protected]) or to R.S.H. (email: [email protected]). #A full list of authors and their affliations appears at the end of the paper. NATURE COMMUNICATIONS | (2018) 9:3707 | DOI: 10.1038/s41467-018-04989-w | www.nature.com/naturecommunications 1 ARTICLE NATURE COMMUNICATIONS | DOI: 10.1038/s41467-018-04989-w ultiple myeloma (MM) is a malignancy of plasma cells consistent OR across all GWAS data sets, by genotyping an Mprimarily located within the bone marrow. -

Acetyltransferase Machinery Conserved in P300/CBP-Family Proteins
Oncogene (2002) 21, 2253 ± 2260 ã 2002 Nature Publishing Group All rights reserved 0950 ± 9232/02 $25.00 www.nature.com/onc SHORT REPORT Acetyltransferase machinery conserved in p300/CBP-family proteins L Wuchao Yuan1 and Antonio Giordano*,2 1Department of Physiology and Biophysics, Boston University School of Medicine, Boston, Massachusetts, MA 02118, USA; 2Department of Pathology, Anatomy and Cell Biology, Jeerson Medical College, Philadelphia, Pennsylvania, PA 19107, USA CREB-binding protein (CBP) and p300 are highly supports this proposal. Later, this protein family was conserved and functionally related transcription coacti- expanded after the discovery of Drosophila CBP (dCBP), vators and histone/protein acetyltransferases. They are which was also able to bind to E1A and coactivate tumor suppressors, participate in a wide variety of CREB-dependent transactivation (Akimaru et al., physiological events, and serve as integrators among 1997a). Recently, p300/CBP-like proteins of the plant dierent signal transduction pathways. In this study, 11 Arabidopsis thaliana have been reported (Bordoli et al., distinct proteins that have a high degree of homology 2001), suggesting that they are possible members of the with the amino acid sequence of p300 have been p300/CBP family. It is interesting to know how large the identi®ed in current protein databases. All of these 11 p300/CBP protein family could be and how this family of proteins belong to either animal or plant multicellular proteins is conserved evolutionarily. organisms (higher eucaryotes). Conservation of p300/ Taking advantage of recent progress in genomic CBP domains among these proteins was examined sequencing of dierent organisms, we performed a blast further by sequence alignment and pattern search. -
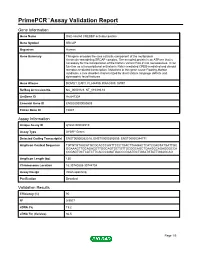
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name Snf2-related CREBBP activator protein Gene Symbol SRCAP Organism Human Gene Summary This gene encodes the core catalytic component of the multiprotein chromatin-remodeling SRCAP complex. The encoded protein is an ATPase that is necessary for the incorporation of the histone variant H2A.Z into nucleosomes. It can function as a transcriptional activator in Notch-mediated CREB-mediated and steroid receptor-mediated transcription. Mutations in this gene cause Floating-Harbor syndrome a rare disorder characterized by short stature language deficits and dysmorphic facial features. Gene Aliases DOMO1, EAF1, FLJ44499, KIAA0309, SWR1 RefSeq Accession No. NC_000016.9, NT_010393.16 UniGene ID Hs.647334 Ensembl Gene ID ENSG00000080603 Entrez Gene ID 10847 Assay Information Unique Assay ID qHsaCID0006510 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000262518, ENST00000395059, ENST00000344771 Amplicon Context Sequence TGTGTGTAACATGCGCACCCAGTTCCCTGACTTAAGACTCATCCAGTATGATTGC GGAAAGTTGCAGACGTTGGCAGTGCTGTTGCGGCAGCTCAAGGCAGAGGGCCA CCGAGTGCTCATCTTCACCCAGATGACCCGAATGCTGGATGTATTGGAGCAG Amplicon Length (bp) 130 Chromosome Location 16:30740838-30744704 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 90 R2 0.9977 cDNA Cq 19.2 cDNA Tm (Celsius) 86.5 Page 1/5 PrimePCR™Assay Validation Report gDNA Cq Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report SRCAP, Human -

Dynamics of Transcription Elongation Are Finely Tuned by Dozens of Regulatory Factors
bioRxiv preprint doi: https://doi.org/10.1101/2021.08.15.456358; this version posted August 15, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. Dynamics of transcription elongation are finely tuned by dozens of regulatory factors Mary Couvillion1*, Kevin M. Harlen1*, Kate C. Lachance1*, Kristine L. Trotta1, Erin Smith1, Christian Brion1, Brendan M. Smalec1, L. Stirling Churchman1 1Blavatnik Institute, Department of Genetics, Harvard Medical School, Boston, Massachusetts, USA 02115 *these authors contributed equally 1 bioRxiv preprint doi: https://doi.org/10.1101/2021.08.15.456358; this version posted August 15, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. ABSTRACT Understanding the complex network and dynamics that regulate transcription elongation requires the quantitative analysis of RNA polymerase II (Pol II) activity in a wide variety of regulatory environments. We performed native elongating transcript sequencing (NET-seq) in 41 strains of S. cerevisiae lacking known elongation regulators, including RNA processing factors, transcription elongation factors, chromatin modifiers, and remodelers. We found that the opposing effects of these factors balance transcription elongation dynamics. Different sets of factors tightly regulate Pol II progression across gene bodies so that Pol II density peaks at key points of RNA processing.