6.7143 Term P-Value Genes GO:0007155 Cell Adhesion 1.96×10
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Sex Differences in Glutamate Receptor Gene Expression in Major Depression and Suicide
Molecular Psychiatry (2015) 20, 1057–1068 © 2015 Macmillan Publishers Limited All rights reserved 1359-4184/15 www.nature.com/mp IMMEDIATE COMMUNICATION Sex differences in glutamate receptor gene expression in major depression and suicide AL Gray1, TM Hyde2,3, A Deep-Soboslay2, JE Kleinman2 and MS Sodhi1,4 Accumulating data indicate that the glutamate system is disrupted in major depressive disorder (MDD), and recent clinical research suggests that ketamine, an antagonist of the N-methyl-D-aspartate (NMDA) glutamate receptor (GluR), has rapid antidepressant efficacy. Here we report findings from gene expression studies of a large cohort of postmortem subjects, including subjects with MDD and controls. Our data reveal higher expression levels of the majority of glutamatergic genes tested in the dorsolateral prefrontal cortex (DLPFC) in MDD (F21,59 = 2.32, P = 0.006). Posthoc data indicate that these gene expression differences occurred mostly in the female subjects. Higher expression levels of GRIN1, GRIN2A-D, GRIA2-4, GRIK1-2, GRM1, GRM4, GRM5 and GRM7 were detected in the female patients with MDD. In contrast, GRM5 expression was lower in male MDD patients relative to male controls. When MDD suicides were compared with MDD non-suicides, GRIN2B, GRIK3 and GRM2 were expressed at higher levels in the suicides. Higher expression levels were detected for several additional genes, but these were not statistically significant after correction for multiple comparisons. In summary, our analyses indicate a generalized disruption of the regulation of the GluRs in the DLPFC of females with MDD, with more specific GluR alterations in the suicides and in the male groups. -
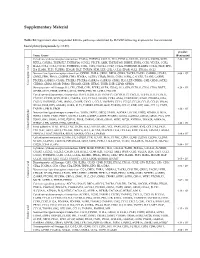
Supplementary Material
Supplementary Material Table S1: Significant downregulated KEGGs pathways identified by DAVID following exposure to five cinnamon- based phenylpropanoids (p < 0.05). p-value Term: Genes (Benjamini) Cytokine-cytokine receptor interaction: FASLG, TNFSF14, CXCL11, IL11, FLT3LG, CCL3L1, CCL3L3, CXCR6, XCR1, 2.43 × 105 RTEL1, CSF2RA, TNFRSF17, TNFRSF14, CCNL2, VEGFB, AMH, TNFRSF10B, INHBE, IFNB1, CCR3, VEGFA, CCR2, IL12A, CCL1, CCL3, CXCL5, TNFRSF25, CCR1, CSF1, CX3CL1, CCL7, CCL24, TNFRSF1B, IL12RB1, CCL21, FIGF, EPO, IL4, IL18R1, FLT1, TGFBR1, EDA2R, HGF, TNFSF8, KDR, LEP, GH2, CCL13, EPOR, XCL1, IFNA16, XCL2 Neuroactive ligand-receptor interaction: OPRM1, THRA, GRIK1, DRD2, GRIK2, TACR2, TACR1, GABRB1, LPAR4, 9.68 × 105 GRIK5, FPR1, PRSS1, GNRHR, FPR2, EDNRA, AGTR2, LTB4R, PRSS2, CNR1, S1PR4, CALCRL, TAAR5, GABRE, PTGER1, GABRG3, C5AR1, PTGER3, PTGER4, GABRA6, GABRA5, GRM1, PLG, LEP, CRHR1, GH2, GRM3, SSTR2, Chlorogenic acid Chlorogenic CHRM3, GRIA1, MC2R, P2RX2, TBXA2R, GHSR, HTR2C, TSHR, LHB, GLP1R, OPRD1 Hematopoietic cell lineage: IL4, CR1, CD8B, CSF1, FCER2, GYPA, ITGA2, IL11, GP9, FLT3LG, CD38, CD19, DNTT, 9.29 × 104 GP1BB, CD22, EPOR, CSF2RA, CD14, THPO, EPO, HLA-DRA, ITGA2B Cytokine-cytokine receptor interaction: IL6ST, IL21R, IL19, TNFSF15, CXCR3, IL15, CXCL11, TGFB1, IL11, FLT3LG, CXCL10, CCR10, XCR1, RTEL1, CSF2RA, IL21, CCNL2, VEGFB, CCR8, AMH, TNFRSF10C, IFNB1, PDGFRA, EDA, CXCL5, TNFRSF25, CSF1, IFNW1, CNTFR, CX3CL1, CCL5, TNFRSF4, CCL4, CCL27, CCL24, CCL25, CCL23, IFNA6, IFNA5, FIGF, EPO, AMHR2, IL2RA, FLT4, TGFBR2, EDA2R, -

Ion Channels
UC Davis UC Davis Previously Published Works Title THE CONCISE GUIDE TO PHARMACOLOGY 2019/20: Ion channels. Permalink https://escholarship.org/uc/item/1442g5hg Journal British journal of pharmacology, 176 Suppl 1(S1) ISSN 0007-1188 Authors Alexander, Stephen PH Mathie, Alistair Peters, John A et al. Publication Date 2019-12-01 DOI 10.1111/bph.14749 License https://creativecommons.org/licenses/by/4.0/ 4.0 Peer reviewed eScholarship.org Powered by the California Digital Library University of California S.P.H. Alexander et al. The Concise Guide to PHARMACOLOGY 2019/20: Ion channels. British Journal of Pharmacology (2019) 176, S142–S228 THE CONCISE GUIDE TO PHARMACOLOGY 2019/20: Ion channels Stephen PH Alexander1 , Alistair Mathie2 ,JohnAPeters3 , Emma L Veale2 , Jörg Striessnig4 , Eamonn Kelly5, Jane F Armstrong6 , Elena Faccenda6 ,SimonDHarding6 ,AdamJPawson6 , Joanna L Sharman6 , Christopher Southan6 , Jamie A Davies6 and CGTP Collaborators 1School of Life Sciences, University of Nottingham Medical School, Nottingham, NG7 2UH, UK 2Medway School of Pharmacy, The Universities of Greenwich and Kent at Medway, Anson Building, Central Avenue, Chatham Maritime, Chatham, Kent, ME4 4TB, UK 3Neuroscience Division, Medical Education Institute, Ninewells Hospital and Medical School, University of Dundee, Dundee, DD1 9SY, UK 4Pharmacology and Toxicology, Institute of Pharmacy, University of Innsbruck, A-6020 Innsbruck, Austria 5School of Physiology, Pharmacology and Neuroscience, University of Bristol, Bristol, BS8 1TD, UK 6Centre for Discovery Brain Science, University of Edinburgh, Edinburgh, EH8 9XD, UK Abstract The Concise Guide to PHARMACOLOGY 2019/20 is the fourth in this series of biennial publications. The Concise Guide provides concise overviews of the key properties of nearly 1800 human drug targets with an emphasis on selective pharmacology (where available), plus links to the open access knowledgebase source of drug targets and their ligands (www.guidetopharmacology.org), which provides more detailed views of target and ligand properties. -

Neuroplasticity Pathways and Protein-Interaction Networks Are
Waller et al. BMC Neurosci (2017) 18:56 DOI 10.1186/s12868-017-0376-x BMC Neuroscience RESEARCH ARTICLE Open Access Neuroplasticity pathways and protein‑interaction networks are modulated by vortioxetine in rodents Jessica A. Waller1, Sara Holm Nygaard2, Yan Li1, Kristian Gaarn du Jardin3, Joseph A. Tamm4, Aicha Abdourahman4, Betina Elfving3, Alan L. Pehrson1, Connie Sánchez1* and Rasmus Wernersson2,5* Abstract Background: The identifcation of biomarkers that predict susceptibility to major depressive disorder and treatment response to antidepressants is a major challenge. Vortioxetine is a novel multimodal antidepressant that possesses pro-cognitive properties and diferentiates from other conventional antidepressants on various cognitive and plastic- ity measures. The aim of the present study was to identify biological systems rather than single biomarkers that may underlie vortioxetine’s treatment efects. Results: We show that the biological systems regulated by vortioxetine are overlapping between mouse and rat in response to distinct treatment regimens and in diferent brain regions. Furthermore, analysis of complexes of physically-interacting proteins reveal that biomarkers involved in transcriptional regulation, neurodevelopment, neu- roplasticity, and endocytosis are modulated by vortioxetine. A subsequent qPCR study examining the expression of targets in the protein–protein interactome space in response to chronic vortioxetine treatment over a range of doses provides further biological validation that vortioxetine engages neuroplasticity networks. Thus, the same biology is regulated in diferent species and sexes, diferent brain regions, and in response to distinct routes of administration and regimens. Conclusions: A recurring theme, based on the present study as well as previous fndings, is that networks related to synaptic plasticity, synaptic transmission, signal transduction, and neurodevelopment are modulated in response to vortioxetine treatment. -

The Genes Encoding the Glutamate Receptor Subunits Kal And
Proc. Nati. Acad. Sci. USA Vol. 91, pp. 11849-11853, December 1994 Neurobiology The genes encoding the glutamate receptor subunits KAl and KA2 (GRIK4 and GRIK5) are located on separate chromosomes in human, mouse, and rat (lgand-gated ion channels/human chromosomes 19q13.2 and 11q22-23/mouse chromosomes 7 and 9/rat chromosomes 8 and 1) CLAUDE SZPIRER*, MONTSE MOLNOt, RACHELE ANTONACCI*, NANCY A. JENKINS§, PALMA FINELLI*, JOSIANE SZPIRER*, MICHELE RivIERE*, MARIANO ROCCHIt, DEBRA J. GILBERT§, NEAL G. COPELAND§, AND VITTORIO GALLOt¶ *Departement de Biologie Moleculaire, Universitd Libre de Bruxelles, B-1640 Rhode-St.-Genese, Belgium; tLaboratory of Cellular and Molecular Neurophysiology, National Institute of Child Health and Human Development, National Institutes of Health, Bethesda, MD 20892; tlstituto di Genetica, University of Bari, 70126 Bari, Italy; Mammalian Genetics Laboratory, Advanced BioScience Laboratories-Basic Research Program, National Cancer Institute, Frederick Cancer Research and Development Center, Frederick, MD 21702 Communicated by E. Costa, August 10, 1994 ABSTRACT The chromosomal alization of the human (AMPA)-preferring family (GluR1-4, or GluRA-D; GRIA and rat genes encoding the kalnate-preferring glutamate recep- gene family) and the two kainate-preferring families tor subunits KAl and KA2 (GRIK4 and GRIKS, respectively) (GluR5-7 and KA1 and KA2; in the two GRIK gene was determined by Southern analysis ofrat x mouse and human families). The kainate-preferring subunits KA1 and KA2 x mouse somatic cell hybrid panels and by fluorescence in situ display 68% identity in their amino acid sequence and code hybridization. The localization of the mouse genes (Grik4 and for proteins that do not form functional homomeric ionic Gri5) was established by interspecific backcross mapping. -

Kava for the Treatment of Generalised Anxiety Disorder (K-GAD): Study Protocol for a Randomised Controlled Trial Karen M
Savage et al. Trials (2015) 16:493 DOI 10.1186/s13063-015-0986-5 TRIALS STUDY PROTOCOL Open Access Kava for the treatment of generalised anxiety disorder (K-GAD): study protocol for a randomised controlled trial Karen M. Savage1,2*, Con K. Stough2, Gerard J. Byrne3, Andrew Scholey2, Chad Bousman2,4,5,6, Jenifer Murphy1, Patricia Macdonald3, Chao Suo7, Matthew Hughes8, Stuart Thomas9, Rolf Teschke10, Chengguo Xing11 and Jerome Sarris1,2 Abstract Background: Generalised anxiety disorder (GAD) is a chronic and pervasive condition that generates high levels of psychological stress, and it is difficult to treat in the long term. Current pharmacotherapeutic options for GAD are in some cases only modestly effective, and may elicit undesirable side effects. Through targeted actions on the gamma-aminobutyric acid (GABA) pathway, the South Pacific medicinal plant kava (Piper methysticum)isa non-addictive, non-hypnotic anxiolytic with the potential to treat GAD. The evidence for the efficacy of kava for treating anxiety has been affirmed through clinical trials and meta-analyses. Recent research has also served to lessen safety concerns regarding the use of kava due to hepatotoxic risk, which is reflected in a recent German court overturning the previous kava ban in that country (which may in turn influence a reinstatement by the European Union). The aim of current research is to assess the efficacy of an ‘aqueous noble cultivar rootstock extract’ of kava in GAD in a larger longer term study. In addition, we plan to investigate the pharmacogenomic influence of GABA transporters on response, effects of kava on gene expression, and for the first time, the neurobiological correlates of treatment response via functional and metabolic imaging. -

The Hypothalamus As a Hub for SARS-Cov-2 Brain Infection and Pathogenesis
bioRxiv preprint doi: https://doi.org/10.1101/2020.06.08.139329; this version posted June 19, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. The hypothalamus as a hub for SARS-CoV-2 brain infection and pathogenesis Sreekala Nampoothiri1,2#, Florent Sauve1,2#, Gaëtan Ternier1,2ƒ, Daniela Fernandois1,2 ƒ, Caio Coelho1,2, Monica ImBernon1,2, Eleonora Deligia1,2, Romain PerBet1, Vincent Florent1,2,3, Marc Baroncini1,2, Florence Pasquier1,4, François Trottein5, Claude-Alain Maurage1,2, Virginie Mattot1,2‡, Paolo GiacoBini1,2‡, S. Rasika1,2‡*, Vincent Prevot1,2‡* 1 Univ. Lille, Inserm, CHU Lille, Lille Neuroscience & Cognition, DistAlz, UMR-S 1172, Lille, France 2 LaBoratorY of Development and PlasticitY of the Neuroendocrine Brain, FHU 1000 daYs for health, EGID, School of Medicine, Lille, France 3 Nutrition, Arras General Hospital, Arras, France 4 Centre mémoire ressources et recherche, CHU Lille, LiCEND, Lille, France 5 Univ. Lille, CNRS, INSERM, CHU Lille, Institut Pasteur de Lille, U1019 - UMR 8204 - CIIL - Center for Infection and ImmunitY of Lille (CIIL), Lille, France. # and ƒ These authors contriButed equallY to this work. ‡ These authors directed this work *Correspondence to: [email protected] and [email protected] Short title: Covid-19: the hypothalamic hypothesis 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.06.08.139329; this version posted June 19, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. -

Association Between the Ionotropic Glutamate Receptor Kainate 3
Molecular Psychiatry (2002) 7, 416–418 2002 Nature Publishing Group All rights reserved 1359-4184/02 $25.00 www.nature.com/mp ORIGINAL RESEARCH ARTICLE Association between the ionotropic glutamate receptor kainate 3 (GRIK3) ser310ala polymorphism and schizophrenia S Begni1, M Popoli2, S Moraschi1, S Bignotti3, GB Tura3 and M Gennarelli1 1Genetics Unit, IRCCS ‘S Giovanni di Dio’, Fatebenefratelli, 25123 Brescia, Italy; 2Center of Neuropharmacology, Institute of Pharmacological Sciences, University of Milan, 20133 Milan, Italy; 3Psychiatric Rehabilitation Unit, IRCCS ‘S Giovanni di Dio’, Fatebenefratelli, 25123 Brescia, Italy Keywords: association study; cSNP; expression; kainate through ionotropic glutamate receptors influences the receptor; neuronal plasticity regulation of transcriptional, translational and post- Schizophrenia is a severe psychiatric illness character- translational processes fundamental for the function of ised by disturbance of thought, hallucination and brain cells.3 delusions.1 Several studies have suggested that dys- One of the hypotheses about the molecular mech- functions in the glutamatergic transmission are linked anisms leading to schizophrenia is the presence of an to the pathogenesis of schizophrenia, and in particular excessive activation of glutamate receptors. An an excessive activation of glutamate receptors seems to increase in the basic metabolic activity along the gluta- be related to the disruption of neuronal ionic gradients matergic axons has been demonstrated by PET scan leading to excitotoxicity.2–7 -

Recurrent Copy Number Changes in Mentally Retarded Children Harbour Genes Involved in Cellular Localization and the Glutamate Receptor Complex
European Journal of Human Genetics (2010) 18, 39–46 & 2010 Macmillan Publishers Limited All rights reserved 1018-4813/10 $32.00 www.nature.com/ejhg ARTICLE Recurrent copy number changes in mentally retarded children harbour genes involved in cellular localization and the glutamate receptor complex Martin Poot*,1, Marc J Eleveld1, Ruben van ‘t Slot1, Hans Kristian Ploos van Amstel1 and Ron Hochstenbach1 To determine the phenotypic significance of copy number changes (CNCs) in the human genome, we performed genome-wide segmental aneuploidy profiling by BAC-based array-CGH of 278 unrelated patients with multiple congenital abnormalities and mental retardation (MCAMR) and in 48 unaffected family members. In 20 patients, we found de novo CNCs composed of multiple consecutive probes. Of the 125 probes making up these probably pathogenic CNCs, 14 were also found as single CNCs in other patients and 5 in healthy individuals. Thus, these CNCs are not by themselves pathogenic. Almost one out of five patients and almost one out of six healthy individuals in our study cohort carried a gain or a loss for any one of the recently discovered microdeletion/microduplication loci, whereas seven patients and one healthy individual showed losses or gains for at least two different loci. The pathogenic burden resulting from these CNCs may be limited as they were found with similar frequencies among patients and healthy individuals (P¼0.165; Fischer’s exact test), and several individuals showed CNCs at multiple loci. CNCs occurring specifically in our study cohort were enriched for components of the glutamate receptor family (GRIA2, GRIA4, GRIK2 and GRIK4) and genes encoding proteins involved in guiding cell localization during development (ATP1A2, GIRK3, GRIA2, KCNJ3, KCNJ10, KCNK17 and KCNK5). -

Genetic Studies on the Tripartite Glutamate Synapse in the Pathophysiology and Therapeutics of Mood Disorders
Neuropsychopharmacology (2017) 42, 787–800 © 2017 American College of Neuropsychopharmacology. All rights reserved 0893-133X/17 www.neuropsychopharmacology.org Review Genetic Studies on the Tripartite Glutamate Synapse in the Pathophysiology and Therapeutics of Mood Disorders Rafael T de Sousa*,1,2, Alexandre A Loch2, André F Carvalho3, André R Brunoni2, Marie Reine Haddad4, 1 1 1,2,5 Ioline D Henter , Carlos A Zarate Jr and Rodrigo Machado-Vieira 1Experimental Therapeutics and Pathophysiology Branch, National Institute of Mental Health, NIH, Bethesda, MD, USA; 2Laboratory of Neuroscience, LIM- 27, Institute and Department of Psychiatry, University of São Paulo, São Paulo, Brazil; 3Translational Psychiatry Research Group 4 and Department of Clinical Medicine, Faculty of Medicine, Federal University of Ceará, Fortaleza, Brazil; Section on Translational Neuroscience, 5 National Institute of Child Health and Human Development, NIH, Bethesda, MD, USA; Center for Interdisciplinary Research on Applied Neurosciences (NAPNA), University of São Paulo, São Paulo, Brazil Both bipolar disorder (BD) and major depressive disorder (MDD) have high morbidity and share a genetic background. Treatment options for these mood disorders are currently suboptimal for many patients; however, specific genetic variables may be involved in both pathophysiology and response to treatment. Agents such as the glutamatergic modulator ketamine are effective in treatment-resistant mood disorders, underscoring the potential importance of the glutamatergic system as a target for improved therapeutics. Here we review genetic studies linking the glutamatergic system to the pathophysiology and therapeutics of mood disorders. We screened 763 original genetic studies of BD or MDD that investigated genes encoding targets of the pathway/mediators related to the so-called tripartite glutamate synapse, including pre- and post-synaptic neurons and glial cells; 60 papers were included in this review. -

Detection of H3k4me3 Identifies Neurohiv Signatures, Genomic
viruses Article Detection of H3K4me3 Identifies NeuroHIV Signatures, Genomic Effects of Methamphetamine and Addiction Pathways in Postmortem HIV+ Brain Specimens that Are Not Amenable to Transcriptome Analysis Liana Basova 1, Alexander Lindsey 1, Anne Marie McGovern 1, Ronald J. Ellis 2 and Maria Cecilia Garibaldi Marcondes 1,* 1 San Diego Biomedical Research Institute, San Diego, CA 92121, USA; [email protected] (L.B.); [email protected] (A.L.); [email protected] (A.M.M.) 2 Departments of Neurosciences and Psychiatry, University of California San Diego, San Diego, CA 92103, USA; [email protected] * Correspondence: [email protected] Abstract: Human postmortem specimens are extremely valuable resources for investigating trans- lational hypotheses. Tissue repositories collect clinically assessed specimens from people with and without HIV, including age, viral load, treatments, substance use patterns and cognitive functions. One challenge is the limited number of specimens suitable for transcriptional studies, mainly due to poor RNA quality resulting from long postmortem intervals. We hypothesized that epigenomic Citation: Basova, L.; Lindsey, A.; signatures would be more stable than RNA for assessing global changes associated with outcomes McGovern, A.M.; Ellis, R.J.; of interest. We found that H3K27Ac or RNA Polymerase (Pol) were not consistently detected by Marcondes, M.C.G. Detection of H3K4me3 Identifies NeuroHIV Chromatin Immunoprecipitation (ChIP), while the enhancer H3K4me3 histone modification was Signatures, Genomic Effects of abundant and stable up to the 72 h postmortem. We tested our ability to use H3K4me3 in human Methamphetamine and Addiction prefrontal cortex from HIV+ individuals meeting criteria for methamphetamine use disorder or not Pathways in Postmortem HIV+ Brain (Meth +/−) which exhibited poor RNA quality and were not suitable for transcriptional profiling. -

Grik4 Antibody Cat
Grik4 Antibody Cat. No.: 4391 Western blot analysis of Grik4 in rat brain lysate with Grik4 antibody at (A) 0.5 (B) 1 and (C) 2 μg/mL. Immunohistochemical staining of human brain tissue Immunofluorescence of Grik4 in Human Brain cells with using Grik4 antibody at 2.5 μg/mL. Grik4 antibody at 20 μg/mL. Specifications HOST SPECIES: Rabbit SPECIES REACTIVITY: Human, Mouse, Rat Grik4 antibody was raised against a 18 amino acid synthetic peptide near the carboxy terminus of the mouse Grik4. IMMUNOGEN: The immunogen is located within the last 50 amino acids of Grik4. TESTED APPLICATIONS: ELISA, IF, IHC-P, WB September 30, 2021 1 https://www.prosci-inc.com/grik4-antibody-4391.html Grik4 antibody can be used for detection of Grik4 by Western blot at 0.5 - 2 μg/mL. Despite its predicted molecular weight, Grik4 often migrates at a lower molecular weight in SDS-PAGE. Antibody can also be used for immunohistochemistry starting at 2.5 μg/mL. For immunofluorescence start at 20 μg/mL. APPLICATIONS: Antibody validated: Western Blot in rat samples; Immunohistochemistry in human samples and Immunofluorescence in human samples. All other applications and species not yet tested. POSITIVE CONTROL: 1) Cat. No. 1463 - Rat Brain Tissue Lysate 2) Cat. No. 10-301 - Human Brain Tissue Slide Properties PURIFICATION: Grik4 Antibody is affinity chromatography purified via peptide column. CLONALITY: Polyclonal ISOTYPE: IgG CONJUGATE: Unconjugated PHYSICAL STATE: Liquid BUFFER: Grik4 Antibody is supplied in PBS containing 0.02% sodium azide. CONCENTRATION: 1 mg/mL Grik4 antibody can be stored at 4˚C for three months and -20˚C, stable for up to one STORAGE CONDITIONS: year.