Adaptation of Host Transmission Cycle During Pathogen Speciation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Characterization of Functional Residues for Catalysis and Kinetics of Aminoacylhistidine Dipeptidase from Vibrio Alginolyticus
國 立 交 通 大 學 生物科技研究所 碩士論文 溶藻弧菌胺醯組胺酸雙胜肽酶之生化特性分析 及其功能性胺基酸之研究 Characterization of Functional Residues for Catalysis and Kinetics of Aminoacylhistidine Dipeptidase from Vibrio alginolyticus 研究生: 陳怡親 指導教授: 吳東昆 博士 中華民國九十六年七月 Characterization of Functional Residues for Catalysis and Kinetics of Aminoacylhistidine Dipeptidase from Vibrio alginolyticus 研究生:陳怡親 Student: Yi-Chin Chen 指導教授:吳東昆 博士 Advisor: Dr. Tung-Kung Wu 國 立 交 通 大 學 生物科技研究所 碩士論文 A Thesis Submitted to Department of Biological Science and Technology College of Science National Chiao Tung University in partial Fulfillment of the Requirements for the Degree of Master in Biological Science and Technology July, 2007 Hsinchu, Taiwan, Republic of China 中華民國九十六年七月 溶藻弧菌胺醯組胺酸雙胜肽酶之生化特性分析及其功能性胺基酸 之研究 研究生:陳怡親 指導教授:吳東昆 博士 國立交通大學 生物科技研究所碩士班 摘要 胺醯組胺酸雙胜肽酶(PepD, EC 3.4.13.3)為胜肽酶家族M20 中的一員。胜肽酶家族M20 中的酵素皆屬於金屬雙胜肽酶,而經由研究發現可被應用於抗菌、癌症的臨床治療及神 經傳導物質的調控等方面。過去對於細菌中胺醯組胺酸雙胜肽酶的研究很少,只針對其 序列和部分生化特性進行探討,並無其生理角色或活性區胺基酸相關之研究。本論文將 溶藻弧菌pepD基因殖入pET-28a(+)質體中,表現出N端帶有His-tag之重組蛋白,並利用 Ni-NTA親和層析管柱純化之。純化出的蛋白質對於特定的Xaa-His雙胜肽(例如: L-carnosine)具有水解的能力,但無水解三胜肽之活性。經酵素動力學研究,溶藻弧菌 -1 PepD蛋白對雙胜肽L-carnosine之Km與kcat值分別為 0.36 mM與 8.6 min 。經由序列分析 預測溶藻弧菌PepD蛋白上胺基酸位置His80、Asp82、Asp119、Glu149、Glu150、Asp173 及His461 為活性區胺基酸。其中將所預測影響金屬鍵結之胺基酸Asp119 以及扮演催化 角色之胺基酸Glu149 分別進行飽和定點突變,發現大部分之突變蛋白皆失去或降低原 有之活性。此外,以同屬M20 胜肽酶家族之PepV蛋白結晶結構為模板做出溶藻弧菌PepD 蛋白之同源模擬,顯示出相同之活性區胺基酸。因此,根據本論文實驗結果將首次提出 胺醯組胺酸雙胜肽酶活性區胺基酸之分佈情形與其可能扮演之功能。 I Characterization of Functional Residues for Catalysis and Kinetics of Aminoacylhistidine Dipeptidase from Vibrio alginolyticus Student : Yi-Chin Chen Advisor : Dr. Tung-Kung Wu Institute of Biological Science and Technology National Chiao Tung University Abstract Proteins of the aminoacylase-1/metallopeptidase 20 (Acyl/M20) family were characterized to contain a zinc-binding domain at their active site. Aminoacylhistidine dipeptidase (PepD, EC 3.4.13.3) is a member of peptidase family M20 which catalyzes the cleavage and release of N-terminal amino acid, usually are neutral or hydrophobic residue, from Xaa-His peptide or polypeptide. -

A Mycobacterium Avium Subsp. Paratuberculosis Predicted Serine Protease Is Associated with Acid Stress and Intraphagosomal Survival
University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln U.S. Department of Agriculture: Agricultural Publications from USDA-ARS / UNL Faculty Research Service, Lincoln, Nebraska 2016 A Mycobacterium avium subsp. paratuberculosis Predicted Serine Protease Is Associated with Acid Stress and Intraphagosomal Survival Abirami Kugadas University of Minnesota, Boston Elise A. Lamont University of Minnesota, Saint Paul John Bannantine USDA ARS National Animal Disease Center, [email protected] Fernanda Miyagaki Shoyama Michigan State University, [email protected] Evan Brenner USDA ARS National Animal Disease Center See next page for additional authors Follow this and additional works at: https://digitalcommons.unl.edu/usdaarsfacpub Part of the Agriculture Commons Kugadas, Abirami; Lamont, Elise A.; Bannantine, John; Shoyama, Fernanda Miyagaki; Brenner, Evan; Janagama, Harish K.; and Sreevatsan, Srinand, "A Mycobacterium avium subsp. paratuberculosis Predicted Serine Protease Is Associated with Acid Stress and Intraphagosomal Survival" (2016). Publications from USDA-ARS / UNL Faculty. 2391. https://digitalcommons.unl.edu/usdaarsfacpub/2391 This Article is brought to you for free and open access by the U.S. Department of Agriculture: Agricultural Research Service, Lincoln, Nebraska at DigitalCommons@University of Nebraska - Lincoln. It has been accepted for inclusion in Publications from USDA-ARS / UNL Faculty by an authorized administrator of DigitalCommons@University of Nebraska - Lincoln. Authors Abirami Kugadas, Elise A. Lamont, John Bannantine, Fernanda Miyagaki Shoyama, Evan Brenner, Harish K. Janagama, and Srinand Sreevatsan This article is available at DigitalCommons@University of Nebraska - Lincoln: https://digitalcommons.unl.edu/ usdaarsfacpub/2391 ORIGINAL RESEARCH published: 22 August 2016 doi: 10.3389/fcimb.2016.00085 A Mycobacterium avium subsp. paratuberculosis Predicted Serine Protease Is Associated with Acid Stress and Intraphagosomal Survival Abirami Kugadas 1†, Elise A. -

Disease-Related Cellular Protein Networks Differentially Affected
www.nature.com/scientificreports OPEN Disease‑related cellular protein networks diferentially afected under diferent EGFR mutations in lung adenocarcinoma Toshihide Nishimura1,8*, Haruhiko Nakamura1,2,8, Ayako Yachie3,8, Takeshi Hase3,8, Kiyonaga Fujii1,8, Hirotaka Koizumi4, Saeko Naruki4, Masayuki Takagi4, Yukiko Matsuoka3, Naoki Furuya5, Harubumi Kato6,7 & Hisashi Saji2 It is unclear how epidermal growth factor receptor EGFR major driver mutations (L858R or Ex19del) afect downstream molecular networks and pathways. This study aimed to provide information on the infuences of these mutations. The study assessed 36 protein expression profles of lung adenocarcinoma (Ex19del, nine; L858R, nine; no Ex19del/L858R, 18). Weighted gene co-expression network analysis together with analysis of variance-based screening identifed 13 co-expressed modules and their eigen proteins. Pathway enrichment analysis for the Ex19del mutation demonstrated involvement of SUMOylation, epithelial and mesenchymal transition, ERK/mitogen- activated protein kinase signalling via phosphorylation and Hippo signalling. Additionally, analysis for the L858R mutation identifed various pathways related to cancer cell survival and death. With regard to the Ex19del mutation, ROCK, RPS6KA1, ARF1, IL2RA and several ErbB pathways were upregulated, whereas AURK and GSKIP were downregulated. With regard to the L858R mutation, RB1, TSC22D3 and DOCK1 were downregulated, whereas various networks, including VEGFA, were moderately upregulated. In all mutation types, CD80/CD86 (B7), MHC, CIITA and IFGN were activated, whereas CD37 and SAFB were inhibited. Costimulatory immune-checkpoint pathways by B7/CD28 were mainly activated, whereas those by PD-1/PD-L1 were inhibited. Our fndings may help identify potential therapeutic targets and develop therapeutic strategies to improve patient outcomes. -

Dysregulation of Macrophage PEPD in Obesity Determines Adipose Tissue Fbro-Infammation and Insulin Resistance
Dysregulation of macrophage PEPD in obesity determines adipose tissue bro-inammation and insulin resistance Vanessa Pellegrinelli ( [email protected] ) Institute of Metabolic Science Segio Rodriguez-Cuenca MRC Metabolic Diseases Unit, University of Cambridge Christine Rouault Sorbonne Université, INSERM, Nutrition and obesities; systemic approaches (NutriOmics) Hanna Schilbert Genetics and Genomics of Plants, Center for Biotechnology (CeBiTec) & Faculty of Biology, Bielefeld University Sam Virtue University of Cambridge Jose Maria Moreno-Navarrete Institut d'Investigació Biomèdica de Girona Guillaume Bidault University of Cambridge https://orcid.org/0000-0002-8396-9962 Maria del Carmen Vázquez Borrego Maimonides Institute of Biomedical Research of Cordoba (IMIBIC) Ana Rita Dias Metabolic Research Laboratories, Institute of Metabolic Science, Addenbrooke’s Hospital, University of Cambridge Boas Pucker Genetics and Genomics of Plants, Center for Biotechnology (CeBiTec) & Faculty of Biology, Bielefeld University Martin Dale University of Cambridge Metabolic Research Laboratories Mark Campbell University of Cambridge Stefania Carobbio Metabolic Research Laboratories, Institute of Metabolic Science, Addenbrooke’s Hospital, University of Cambridge Judith Aron-Wisnewsky Page 1/30 Assistance-Publique Hôpitaux de Paris, Nutrition department, Pitié-Salpêtrière hospital Silvia Mora Department of Molecular and Cellular Physiology, Institute of Translational Medicine, The University of Liverpool Mauro Masiero The Beijer Laboratory and Department -

Prolidase Deficiency
Prolidase deficiency Author: Professor Jaak Jaeken1 Creation Date: August 2001 Update: January 2004 Scientific Editor: Professor Jean-Marie Saudubray 1Center for Metabolic Disease, University Hospital Gasthuisberg, Herestraat 49, B-3000 Leuven, Belgium. [email protected] Abstract Keywords Disease name Excluded diseases Diagnostic criterium Differential diagnosis Prevalence Clinical description Management Etiology Diagnostic methods Genetic counseling Antenatal diagnosis Unresolved questions References Abstract Prolidase deficiency is a very rare autosomal recessive disease. Prevalence remains unknown, some 40 cases have been reported. It is characterized by mild to severe skin lesions particularly on the face, palms, lower legs and soles, besides variable other features. Patients excrete massive amounts of iminodipeptides in urine due to a deficiency of the exopeptidase prolidase. The activity of this enzyme can be measured in hemolysates, leukocytes or fibroblasts. Mutations in the PEPD gene on chromosome 19p13.2 have been observed. The skin ulcers respond partially to local treatment. Keywords prolidase, iminodipeptiduria Disease name Clinical description Prolidase deficiency First symptoms start between birth and young adult age. Most characteristic are the skin Excluded diseases lesions ranging from mild to severe, including Lathyrism recalcitrant ulcerations. They are mostly located on the face, the palms, the lower legs and the Diagnostic criterium soles. Other features are facial dysmorphy, Prolidase deficiency psychomotor retardation and recurrent infections. Differential diagnosis Systemic lupus erythematosus Management This is limited to treatment of the skin ulcers with Prevalence oral ascorbate, manganese (cofactor of Unknown; some 40 individuals have been prolidase), inhibitors of collagenase, and local reported. Jaeken J; Prolidase deficiency. Orphanet encyclopedia, January 2004. http://www.orpha.net/data/patho/GB/uk-prolidase.pdf 1 applications of L-proline-and glycine-containing References ointments. -
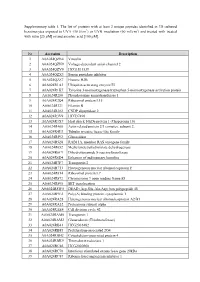
Supplementary Table 1. the List of Proteins with at Least 2 Unique
Supplementary table 1. The list of proteins with at least 2 unique peptides identified in 3D cultured keratinocytes exposed to UVA (30 J/cm2) or UVB irradiation (60 mJ/cm2) and treated with treated with rutin [25 µM] or/and ascorbic acid [100 µM]. Nr Accession Description 1 A0A024QZN4 Vinculin 2 A0A024QZN9 Voltage-dependent anion channel 2 3 A0A024QZV0 HCG1811539 4 A0A024QZX3 Serpin peptidase inhibitor 5 A0A024QZZ7 Histone H2B 6 A0A024R1A3 Ubiquitin-activating enzyme E1 7 A0A024R1K7 Tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein 8 A0A024R280 Phosphoserine aminotransferase 1 9 A0A024R2Q4 Ribosomal protein L15 10 A0A024R321 Filamin B 11 A0A024R382 CNDP dipeptidase 2 12 A0A024R3V9 HCG37498 13 A0A024R3X7 Heat shock 10kDa protein 1 (Chaperonin 10) 14 A0A024R408 Actin related protein 2/3 complex, subunit 2, 15 A0A024R4U3 Tubulin tyrosine ligase-like family 16 A0A024R592 Glucosidase 17 A0A024R5Z8 RAB11A, member RAS oncogene family 18 A0A024R652 Methylenetetrahydrofolate dehydrogenase 19 A0A024R6C9 Dihydrolipoamide S-succinyltransferase 20 A0A024R6D4 Enhancer of rudimentary homolog 21 A0A024R7F7 Transportin 2 22 A0A024R7T3 Heterogeneous nuclear ribonucleoprotein F 23 A0A024R814 Ribosomal protein L7 24 A0A024R872 Chromosome 9 open reading frame 88 25 A0A024R895 SET translocation 26 A0A024R8W0 DEAD (Asp-Glu-Ala-Asp) box polypeptide 48 27 A0A024R9E2 Poly(A) binding protein, cytoplasmic 1 28 A0A024RA28 Heterogeneous nuclear ribonucleoprotein A2/B1 29 A0A024RA52 Proteasome subunit alpha 30 A0A024RAE4 Cell division cycle 42 31 -

Prolidase Deficiency: a New Genetic Cause of Combined Pulmonary Fibrosis and Emphysema Syndrome in the Adult
Early View Research letter Prolidase deficiency: a new genetic cause of combined pulmonary fibrosis and emphysema syndrome in the adult Vincent Cottin, Mouhamad Nasser, Julie Traclet, Lara Chalabreysse, Anne-Sophie Lèbre, Salim Si- Mohamed, François Philit, Françoise Thivolet-Béjui Please cite this article as: Cottin V, Nasser M, Traclet J, Chalabreysse L, et al. Prolidase deficiency: a new genetic cause of combined pulmonary fibrosis and emphysema syndrome in the adult. Eur Respir J 2020; in press (https://doi.org/10.1183/13993003.01952-2019). This manuscript has recently been accepted for publication in the European Respiratory Journal. It is published here in its accepted form prior to copyediting and typesetting by our production team. After these production processes are complete and the authors have approved the resulting proofs, the article will move to the latest issue of the ERJ online. Copyright ©ERS 2020 Prolidase deficiency: a new genetic cause of combined pulmonary fibrosis and emphysema syndrome in the adult Letter to the editor Authors Vincent Cottin 1a, Mouhamad Nasser 1, Julie Traclet 1, Lara Chalabreysse 2, Anne-Sophie Lèbre3, Salim Si-Mohamed 4, François Philit 5, Françoise Thivolet-Béjui 2 ORCID : V. Cottin : 0000-0002-5591-0955; M. Nasser : 0000-0001-8373-8032; A.S. Lebre : 0000-0002-7519-9822 Affiliations 1 National Coordinating Reference Centre for Rare Pulmonary Diseases, Louis Pradel Hospital, Hospices Civils de Lyon, UMR 754, Claude Bernard University Lyon 1, member of OrphaLung, RespiFil, and ERN-LUNG, Lyon, -

Genetics and Molecular Biology, 44, 2, E20200393 (2021) Copyright © Sociedade Brasileira De Genética
Genetics and Molecular Biology, 44, 2, e20200393 (2021) Copyright © Sociedade Brasileira de Genética. DOI: https://doi.org/10.1590/1678-4685-GMB-2020-0393 Research Article Human and Medical Genetics Structural analysis of new compound heterozygous variants in PEPD gene identified in a patient with Prolidase Deficiency diagnosed by exome sequencing Natália D. Linhares1,2*, Piotr Wilk3,4*, Elżbieta Wątor3,4, Meire A. Tostes5, Manfred S. Weiss3 and Sergio D. J. Pena1,2,6 1Universidade Federal de Minas Gerais, Faculdade de Medicina, Laboratório de Genômica Clínica, Belo Horizonte, MG, Brazil. 2Universidade Federal de Minas Gerais, Instituto de Ciências Biológicas, Departamento de Bioquímica e Imunologia, Belo Horizonte, MG, Brazil. 3Helmholtz-Zentrum Berlin, Macromolecular Crystallography (HZB-MX), Berlin, Germany. 4Jagiellonian University, Malopolska Centre of Biotechnology, Kraków, Poland. 5Hospital das Clínicas da Universidade Federal de Minas Gerais, Serviço de Hematologia, Belo Horizonte, MG, Brazil. 6Laboratório Gene - Núcleo de Genética Médica, Belo Horizonte, MG, Brazil. Abstract Prolidase Deficiency (PD) is an autosomal recessive rare disorder caused by loss or reduction of prolidase enzymatic activity due to variants in the PEPD gene. PD clinical features vary among affected individuals: skin ulcerations, recurrent infections, and developmental delay are common. In this study, we describe a 16-year-old boy with a mild PD phenotype comprising chronic eczema, recurrent infections and elevated IgE. Whole exome sequencing analysis revealed three PEPD variants: c.575T>C p.(Leu192Pro) inherited from the mother, and c.692_694del p.(Tyr231del) and c.1409G>A p.(Arg470His), both inherited from the father. The variant p.(Tyr231del) has been previously characterized by high-resolution X-ray structure analysis as altering protein dynamics/flexibility. -

The Nav1.7 Sodium Channel: from Molecule to Man
REVIEWS The NaV1.7 sodium channel: from molecule to man Sulayman D. Dib-Hajj1,2, Yang Yang1,2, Joel A. Black1,2 and Stephen G. Waxman1,2 Abstract | The voltage-gated sodium channel NaV1.7 is preferentially expressed in peripheral somatic and visceral sensory neurons, olfactory sensory neurons and sympathetic ganglion neurons. NaV1.7 accumulates at nerve fibre endings and amplifies small subthreshold depolarizations, poising it to act as a threshold channel that regulates excitability. Genetic and functional studies have added to the evidence that NaV1.7 is a major contributor to pain signalling in humans, and homology modelling based on crystal structures of ion channels suggests an atomic-level structural basis for the altered gating of mutant NaV1.7 that causes pain. Neuropathic pain Voltage-gated sodium channels are essential for electro- that are not seen when these channels are expressed in 11 Pain resulting from lesions or genesis in excitable cells. Nine pore-forming α‑subunits HEK 293 cells . Methods are now available that allow the diseases of the somatosensory of such channels (referred to as channels hereinafter), expression and functional profiling of sodium channels in system. 1 NaV1.1–NaV1.9, have been identified in mammals . peripheral neurons, which more closely mimic the in vivo These isoforms share a common overall structural environment of such channels12. motif (FIG. 1). They are each composed of a long poly- In humans, gain‑of‑function mutations in SCN9A, neuropathic pain peptide (1,700–2,000 amino acids) that folds into four which encodes NaV1.7, lead to severe , homologous domains (DI–DIV) that are linked by three whereas loss‑of‑function mutations in this gene lead to loops (L1–L3), with each domain having six transmem- an indifference to pain13. -

Molecular Immunology Deiminated Proteins in Extracellular Vesicles
Molecular Immunology 117 (2020) 37–53 Contents lists available at ScienceDirect Molecular Immunology journal homepage: www.elsevier.com/locate/molimm Deiminated proteins in extracellular vesicles and serum of llama (Lama glama)—Novel insights into camelid immunity T Michael F. Criscitielloa,b, Igor Kraevc, Sigrun Langed,* a Comparative Immunogenetics Laboratory, Department of Veterinary Pathobiology, College of Veterinary Medicine and Biomedical Sciences, Texas A&M University, College Station, TX, 77843, USA b Department of Microbial Pathogenesis and Immunology, College of Medicine, Texas A&M Health Science Center, Texas A&M University, College Station, TX, 77843, USA c Electron Microscopy Suite, Faculty of Science, Technology, Engineering and Mathematics, Open University, Milton Keynes, MK7 6AA, UK d Tissue Architecture and Regeneration Research Group, School of Life Sciences, University of Westminster, London W1W 6UW, UK ARTICLE INFO ABSTRACT Keywords: Peptidylarginine deiminases (PADs) are phylogenetically conserved calcium-dependent enzymes which post- Peptidylarginine deiminases (PADs) translationally convert arginine into citrulline in target proteins in an irreversible manner, causing functional Protein deimination and structural changes in target proteins. Protein deimination causes generation of neo-epitopes, affects gene Llama (Lama glama) regulation and also allows for protein moonlighting. Furthermore, PADs have been found to be a phylogeneti- extracellular vesicles (EVs) cally conserved regulator for extracellular vesicle (EVs) release. EVs are found in most body fluids and parti- Innate immunity cipate in cellular communication via transfer of cargo proteins and genetic material. In this study, post-trans- Adaptive immunity fi Metabolism lationally deiminated proteins in serum and serum-EVs are described for the rst time in camelids, using the Complement llama (Lama glama L. -

A Single Nucleotide Change in the Prolidase Gene in Fibroblasts from Two Patients with Polypeptide Positive Prolidase Deficiency
A single nucleotide change in the prolidase gene in fibroblasts from two patients with polypeptide positive prolidase deficiency. Expression of the mutant enzyme in NIH 3T3 cells. A Tanoue, … , A Kitano, I Matsuda J Clin Invest. 1990;86(1):351-355. https://doi.org/10.1172/JCI114708. Research Article Prolidase deficiency is an autosomal recessive disorder characterized by mental retardation and various skin lesions. Cultured skin fibroblasts were obtained from two independent patients with abnormal prolidase. Using the polymerase chain reaction, we amplified the entire coding region of human prolidase mRNA derived from patients' fibroblasts. Nucleotide sequence analysis of amplified cDNA products revealed a G to A substitution at position 826 in exon 12, where aspartic acid was replaced by asparagine at the amino acid residue 276, in cells from both patients. An analysis of the DNA showed that the substitution was homozygous. An expression plasmid clone containing a normal human prolidase cDNA (pEPD-W) or mutant prolidase cDNA (pEPD-M) was prepared, transfected, and tested for expression in NIH 3T3 cells. Incorporation of pEPD-W and pEPD-M resulted in the synthesis of an immunological polypeptide that corresponded to human prolidase. Active human enzyme was detected in cells transfected with pEPD-W, but not in those transfected with pEPD-M. These results were compatible with our observation of fibroblasts and confirmed that the substitution was responsible for the enzyme deficiency. As active prolidase was recovered in prolidase-deficient fibroblasts transfected with pEPD-W, this restoration of prolidase activity after transfection means that gene replacement therapy for individuals with this human disorder can be given due consideration. -

Mycobacteria and Their Sweet Proteins an Overview of Protein
Tuberculosis 115 (2019) 1–13 Contents lists available at ScienceDirect Tuberculosis journal homepage: www.elsevier.com/locate/tube Review Mycobacteria and their sweet proteins: An overview of protein glycosylation and lipoglycosylation in M. tuberculosis T ∗ Carolina Mehaffy , John T. Belisle, Karen M. Dobos Department of Microbiology, Immunology and Pathology, Colorado State University, 1682 Campus delivery, Fort Collins, CO, USA ARTICLE INFO ABSTRACT Keywords: Post-translational modifications represent a key aspect of enzyme and protein regulation and function. Post- Mycobacterium translational modifications are involved in signaling and response to stress, adaptation to changing environ- Tuberculosis ments, regulation of toxic and damaged proteins, proteins localization and host-pathogen interactions. fi Post-translational modi cations Glycosylation in Mycobacterium tuberculosis (Mtb), is a post-translational modification often found in conjunction Protein glycosylation with acylation in mycobacterial proteins. Since the discovery of glycosylated proteins in the early 1980's, im- portant advances in our understanding of the mechanisms of protein glycosylation have been made. The number of known glycosylated substrates in Mtb has grown through the years, yet many questions remain. This review will explore the current knowledge on protein glycosylation in Mtb, causative agent of Tuberculosis and number one infectious killer in the world. The mechanism and significance of this post- translational modification, as well as maturation, export and acylation of glycosylated proteins will be reviewed. We expect to provide the reader with an overall view of protein glycosylation in Mtb, as well as the significance of this post-translational modification to the physiology and host-pathogen interactions of this important pa- thogen. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD011081 and 10.6019/PXD011081.