BPS Complete Catalog
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Gene Expression Polarization
Transcriptional Profiling of the Human Monocyte-to-Macrophage Differentiation and Polarization: New Molecules and Patterns of Gene Expression This information is current as of September 27, 2021. Fernando O. Martinez, Siamon Gordon, Massimo Locati and Alberto Mantovani J Immunol 2006; 177:7303-7311; ; doi: 10.4049/jimmunol.177.10.7303 http://www.jimmunol.org/content/177/10/7303 Downloaded from Supplementary http://www.jimmunol.org/content/suppl/2006/11/03/177.10.7303.DC1 Material http://www.jimmunol.org/ References This article cites 61 articles, 22 of which you can access for free at: http://www.jimmunol.org/content/177/10/7303.full#ref-list-1 Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision by guest on September 27, 2021 • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2006 by The American Association of Immunologists All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Transcriptional Profiling of the Human Monocyte-to-Macrophage Differentiation and Polarization: New Molecules and Patterns of Gene Expression1 Fernando O. -

Acidic Phospholipids,47, 58 Acidification Steps, 23 Adsorbent, 53
Index acidic phospholipids,47, 58 ATPase, 22, 107 acidification steps, 23 auto-oxidation, 48,49 adsorbent, 53 aequorin, 95 aggregated mitochondria, 23 bacterial cells, 10 alcohols, 49 bacteriorhodopsin (BR), 218,232 alkaline hydrolysis, I bee venom, 121 alkyl esters, 57, 59 binders,53 amide bond, 129 silica gel, 53, 65, 176 amide linkage, 14 silica gel H, 176 amino alcohol, II bioactive phospholipids, 144 amino-group labelling reagents, 121 blanching, 50 aminopeptidase, 17 buoyant density, 16 aminophospholipid, 113, 118 butyl hydroxy toluene (BHT), 49,212 aminophospholipid pump, 119, 140 aminophospholipid translocase, 11 3 amphotericin B (AmB), 107 Cal+ -ATPases, 28, 107 N-(1-deoxyD-fructos-1-yl) AmB, 107 Cal+ -uptake, 27 anchored lipids, 38 campesterol, 104 angular amplitude, 89 calciferol, 78 anilino-8-naphthalene sulfonate (ANS), Candida albicans, 59, 60, 61, 76, 78, 102, 94, 103 104, 108 animal cells, 10 caproic acid, 129, 130 animal tissues, 9 carotenoids, 37 anion transporter, 135 cell debris, 33 anisotropy, 81 , 89, 95, 96, 97, 102, 104, cell disintegrator, 19 105 ceramide, 14, 129, 194 antagonists, 178 ceramide monohexosides ( CMH), 60 antioxidant, 49, 50 cerebroside, 14 apolar lipids, 56, 57, 69 chemical probes,112 arachidonic acid (AA), 144,153, 161 chloroplast, 32, 33 artificial membrane, 83 isolation, 32, 33 ascending chromatography, 54 purification, 33 asymmetric topology, 128 cholesterol, 14, 15,212 asymmetry, 112, 113,119 choline, 7, 9 atebrin, 95 chromatograms, 55 Index 249 chromatographic analyses, 52 ELISA, 146, 148 -

Downloaded from GEO
bioRxiv preprint doi: https://doi.org/10.1101/2020.08.17.252007; this version posted November 3, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. Oxylipin metabolism is controlled by mitochondrial b-oxidation during bacterial inflammation. Mariya Misheva1, Konstantinos Kotzamanis1*, Luke C Davies1*, Victoria J Tyrrell1, Patricia R S Rodrigues1, Gloria A Benavides2, Christine Hinz1, Robert C Murphy3, Paul Kennedy4, Philip R Taylor1,5, Marcela Rosas1, Simon A Jones1, Sumukh Deshpande1, Robert Andrews1, Magdalena A Czubala1, Mark Gurney1, Maceler Aldrovandi1, Sven W Meckelmann1, Peter Ghazal1, Victor Darley-Usmar2, Daniel White1, and Valerie B O’Donnell1 1Systems Immunity Research Institute and Division of Infection and Immunity, and School of Medicine, Cardiff University, UK, 2Department of Pathology, University of Alabama at Birmingham, Birmingham, AL 35294, USA, 3Department of Pharmacology, University of Colorado Denver, Aurora, CO 80045, USA, 4Cayman Chemical 1180 E Ellsworth Rd, Ann Arbor, MI 48108, United States, 5UK Dementia Research Institute at Cardiff, Cardiff University, UK Address correspondence: Valerie O’Donnell, [email protected] or Daniel White, [email protected], Systems Immunity Research Institute, Cardiff University *Both authors contributed equally to the study 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.08.17.252007; this version posted November 3, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. -

Bioluminescent Properties of Semi-Synthetic Obelin and Aequorin Activated by Coelenterazine Analogues with Modifications of C-2, C-6, and C-8 Substituents
International Journal of Molecular Sciences Article Bioluminescent Properties of Semi-Synthetic Obelin and Aequorin Activated by Coelenterazine Analogues with Modifications of C-2, C-6, and C-8 Substituents 1, 2,3, 1 2, Elena V. Eremeeva y , Tianyu Jiang y , Natalia P. Malikova , Minyong Li * and Eugene S. Vysotski 1,* 1 Photobiology Laboratory, Institute of Biophysics SB RAS, Federal Research Center “Krasnoyarsk Science Center SB RAS”, Krasnoyarsk 660036, Russia; [email protected] (E.V.E.); [email protected] (N.P.M.) 2 Key Laboratory of Chemical Biology (MOE), Department of Medicinal Chemistry, School of Pharmaceutical Sciences, Shandong University, Jinan 250012, China; [email protected] 3 State Key Laboratory of Microbial Technology, Shandong University–Helmholtz Institute of Biotechnology, Shandong University, Qingdao 266237, China * Correspondence: [email protected] (M.L.); [email protected] (E.S.V.); Tel.: +86-531-8838-2076 (M.L.); +7-(391)-249-44-30 (E.S.V.); Fax: +86-531-8838-2076 (M.L.); +7-(391)-290-54-90 (E.S.V.) These authors contributed equally to this work. y Received: 23 June 2020; Accepted: 27 July 2020; Published: 30 July 2020 Abstract: Ca2+-regulated photoproteins responsible for bioluminescence of a variety of marine organisms are single-chain globular proteins within the inner cavity of which the oxygenated coelenterazine, 2-hydroperoxycoelenterazine, is tightly bound. Alongside with native coelenterazine, photoproteins can also use its synthetic analogues as substrates to produce flash-type bioluminescence. However, information on the effect of modifications of various groups of coelenterazine and amino acid environment of the protein active site on the bioluminescent properties of the corresponding semi-synthetic photoproteins is fragmentary and often controversial. -

Bioluminescence Related Publications from JNC Corporation
Bioluminescence related publications since 1985 110) Inouye, S. and Hojo, H. (2018) Revalidation of recombinant aequorin as a light emission standard: Estimation of specific activity of Gaussia luciferase. Biochem. Biophys. Res. Commun. 507: 242-245. 109) Yokawa, S., Suzuki, T., Hayashi, A., Inouye, S., Inoh, Y. and Furuno, T. (2018) Video-rate bioluminescence imaging of degranulation of mast cells attached to the extracellular matrix. Front. Cell Dev. Biol. 6: 74. 108) Inouye, S., Tomabechi, Y., Hosoya, T., Sekine, S. and Shirouzu, M. (2018) Slow luminescence kinetics of semi-synthetic aequorin:expression, purification and structure determination of cf3-aequorin. J. Biochem. 164(3): 247–255. 107) Inouye, S. (2018) Single-step purification of recombinant Gaussia luciferase from serum-containing culture medium of mammalian cells. Protein Expr. Purif. 141: 32-38. 106) Inouye, S. and Sahara-Miura, Y. (2017) A fusion protein of the synthetic IgG-binding domain and aequorin: Expression and purification from E. coli cells and its application. Protein Expr. Purif. 137: 58-63. 105) Suzuki, T., Kanamori, T. and Inouye, S. (2017) Quantitative visualization of synchronized insulin secretion from 3D-cultured cells. Biochem. Biophys. Res. Commun. 486: 886-892. 104) Yokawa, S., Suzuki, T., Inouye, S., Inoh, Y., Suzuki, R., Kanamori, T., Furuno, T. and Hirashima, N. (2017) Visualization of glucagon secretion from pancreatic α cells by bioluminescence video microscopy: Identification of secretion sites in the intercellular contact regions. Biochem. Biophys. Res. Commun. 485: 725-730. 103) Inouye, S. and Suzuki, T. (2016) Protein expression of preferred human codon-optimized Gaussia luciferase genes with an artificial open-reading frame in mammalian and bacterial cells. -

Platelet-Neutrophil Interaction
PLATELET-NEUTROPHIL INTERACTION: NEUTROPHILS MODULATE PLATELET FUNCTION Richard William FAINT 1992 A thesis submitted to the University of London for the degree of Ph.D The Department of Haematology University College and Middlesex School of Medicine University College London 98 Chenies Mews London WC1E 6HX ProQuest Number: U049250 All rights reserved INFORMATION TO ALL USERS The quality of this reproduction is dependent upon the quality of the copy submitted. In the unlikely event that the author did not send a com plete manuscript and there are missing pages, these will be noted. Also, if material had to be removed, a note will indicate the deletion. uest ProQuest U049250 Published by ProQuest LLC(2017). Copyright of the Dissertation is held by the Author. All rights reserved. This work is protected against unauthorized copying under Title 17, United States C ode Microform Edition © ProQuest LLC. ProQuest LLC. 789 East Eisenhower Parkway P.O. Box 1346 Ann Arbor, Ml 48106- 1346 2 ABSTRACT Platelets are vital for the arrest of bleeding following tissue injury. Neutrophils play an important role in host defence against bacterial infection. In the milieu of the acute inflammatory response both cellular and non-cellular elements may interact to modify behaviour. Evidence suggests that leukocytes may play an active role in the modulation of platelet function. This interaction may be abnormal in certain pathological states. This study examined the effect of purified neutrophils upon both washed platelets and platelet-rich plasma, as well as in whole blood in vitro. Neutrophils were found to alter platelet behaviour by several mechanisms. These included transcellular metabolism of eicosanoids. -

Antibody List
產品編號 產品名稱 PA569955 1110059E24Rik Polyclonal Antibody PA569956 1110059E24Rik Polyclonal Antibody PA570131 1190002N15Rik Polyclonal Antibody 01-1234-42 123count eBeads Counting Beads MA512242 14.3.3 Pan Monoclonal Antibody (CG15) LFMA0074 14-3-3 beta Monoclonal Antibody (60C10) LFPA0077 14-3-3 beta Polyclonal Antibody PA137002 14-3-3 beta Polyclonal Antibody PA14647 14-3-3 beta Polyclonal Antibody PA515477 14-3-3 beta Polyclonal Antibody PA517425 14-3-3 beta Polyclonal Antibody PA522264 14-3-3 beta Polyclonal Antibody PA529689 14-3-3 beta Polyclonal Antibody MA134561 14-3-3 beta/epsilon/zeta Monoclonal Antibody (3C8) MA125492 14-3-3 beta/zeta Monoclonal Antibody (22-IID8B) MA125665 14-3-3 beta/zeta Monoclonal Antibody (4E2) 702477 14-3-3 delta/zeta Antibody (1H9L19), ABfinity Rabbit Monoclonal 711507 14-3-3 delta/zeta Antibody (1HCLC), ABfinity Rabbit Oligoclonal 702241 14-3-3 epsilon Antibody (5H10L5), ABfinity Rabbit Monoclonal 711273 14-3-3 epsilon Antibody (5HCLC), ABfinity Rabbit Oligoclonal PA517104 14-3-3 epsilon Polyclonal Antibody PA528937 14-3-3 epsilon Polyclonal Antibody PA529773 14-3-3 epsilon Polyclonal Antibody PA575298 14-3-3 eta (Lys81) Polyclonal Antibody MA524792 14-3-3 eta Monoclonal Antibody PA528113 14-3-3 eta Polyclonal Antibody PA529774 14-3-3 eta Polyclonal Antibody PA546811 14-3-3 eta Polyclonal Antibody MA116588 14-3-3 gamma Monoclonal Antibody (HS23) MA116587 14-3-3 gamma Monoclonal Antibody (KC21) PA529690 14-3-3 gamma Polyclonal Antibody PA578233 14-3-3 gamma Polyclonal Antibody 510700 14-3-3 Pan Polyclonal -

Regulation of Tissue Inflammation by 12-Lipoxygenases
biomolecules Review Regulation of Tissue Inflammation by 12-Lipoxygenases Abhishek Kulkarni 1 , Jerry L. Nadler 2, Raghavendra G. Mirmira 1,* and Isabel Casimiro 1,* 1 Department of Medicine, The University of Chicago, Chicago, IL 60637, USA; [email protected] 2 Department of Medicine and Pharmacology, New York Medical College, Valhalla, NY 10595, USA; [email protected] * Correspondence: [email protected] (R.G.M.); [email protected] (I.C.) Abstract: Lipoxygenases (LOXs) are lipid metabolizing enzymes that catalyze the di-oxygenation of polyunsaturated fatty acids to generate active eicosanoid products. 12-lipoxygenases (12-LOXs) primarily oxygenate the 12th carbon of its substrates. Many studies have demonstrated that 12-LOXs and their eicosanoid metabolite 12-hydroxyeicosatetraenoate (12-HETE), have significant pathological implications in inflammatory diseases. Increased level of 12-LOX activity promotes stress (both oxidative and endoplasmic reticulum)-mediated inflammation, leading to damage in these tissues. 12-LOXs are also associated with enhanced cellular migration of immune cells—a characteristic of several metabolic and autoimmune disorders. Genetic depletion or pharmacological inhibition of the enzyme in animal models of various diseases has shown to be protective against disease development and/or progression in animal models in the setting of diabetes, pulmonary, cardiovascular, and metabolic disease, suggesting a translational potential of targeting the enzyme for the treatment of several disorders. In this article, we review the role of 12-LOXs in the pathogenesis of several diseases in which chronic inflammation plays an underlying role. Citation: Kulkarni, A.; Nadler, J.L.; Keywords: 12-lipoxygenases; 12-LOXs; 12/15-lipoxygenase; 12/15-LOX; lipoxygenases; eicosanoids; Mirmira, R.G.; Casimiro, I. -

Original Article Genetic Variants of Aloxs Genes in Polyunsaturated Fatty Acid/Arachidonic Acid Metabolism Associated with Type-2 Diabetes Development
Int J Clin Exp Med 2018;11(12):13797-13805 www.ijcem.com /ISSN:1940-5901/IJCEM0077987 Original Article Genetic variants of ALOXs genes in polyunsaturated fatty acid/arachidonic acid metabolism associated with type-2 diabetes development Jim Jinn-Chyuan Sheu1,3,4,5*, Ying-Ju Lin1,3*, Cherry Yin-Yi Chang2, Shih-Yin Chen1,3, Wen-Ling Liao1,4, Jai-Sing Yang1, Ming-Tsung Lai6, Chih-Mei Chen1, Chun-Cheng Tseng4, Tritium Hwang4, Ping-Ho Chen7, Fuu-Jen Tsai1,3 1Human Genetic Center, 2Department of Obstetrics and Gynecology, China Medical University Hospital, Taichung, Taiwan; 3School of Chinese Medicine, China Medical University, Taichung, Taiwan; 4Institute of Biomedical Sci- ences, National Sun Yat-sen University, Kaohsiung, Taiwan; 5Department of Health and Nutrition Biotechnology, Asia University, Taichung, Taiwan; 6Department of Pathology, Taichung Hospital, Ministry of Health and Welfare, Taichung, Taiwan; 7School of Dentistry, Kaohsiung Medical University, Kaohsiung, Taiwan. *Equal contributors. Received April 16, 2018; Accepted July 24, 2018; Epub December 15, 2018; Published December 30, 2018 Abstract: Poly-unsaturated fatty acids (PUFAs)/arachidonic acids (AAs) and their derived eicosanoids play potent roles in triggering inflammation during obesity and diabetes development. Recent studies have indicated functional roles of ALOX5, ALOX12, ALOX12B, and ALOX15 in the development of insulin resistance and islet β-cell dysfunc- tion. However, the impact of their genetic variants on type 2 diabetes (T2D) development in Asian patients remains unclear. In this study, 1,682 healthy controls and 788 patients with T2D were enrolled for genotyping those four ALOX genes by the TaqMan method. A total of eight Han Chinese-specific SNPs (two SNps for each gene) were selected for this study. -

Fatty Acid Metabolism Mediated by 12/15-Lipoxygenase Is a Novel Regulator of Hematopoietic Stem Cell Function and Myelopoiesis
University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations Spring 2010 Fatty Acid Metabolism Mediated by 12/15-Lipoxygenase is a Novel Regulator of Hematopoietic Stem Cell Function and Myelopoiesis Michelle Kinder University of Pennsylvania, [email protected] Follow this and additional works at: https://repository.upenn.edu/edissertations Part of the Immunology and Infectious Disease Commons Recommended Citation Kinder, Michelle, "Fatty Acid Metabolism Mediated by 12/15-Lipoxygenase is a Novel Regulator of Hematopoietic Stem Cell Function and Myelopoiesis" (2010). Publicly Accessible Penn Dissertations. 88. https://repository.upenn.edu/edissertations/88 This paper is posted at ScholarlyCommons. https://repository.upenn.edu/edissertations/88 For more information, please contact [email protected]. Fatty Acid Metabolism Mediated by 12/15-Lipoxygenase is a Novel Regulator of Hematopoietic Stem Cell Function and Myelopoiesis Abstract Fatty acid metabolism governs critical cellular processes in multiple cell types. The goal of my dissertation was to investigate the intersection between fatty acid metabolism and hematopoiesis. Although fatty acid metabolism has been extensively studied in mature hematopoietic subsets during inflammation, in developing hematopoietic cells the role of fatty acid metabolism, in particular by 12/ 15-Lipoxygenase (12/15-LOX), was unknown. The observation that 12/15-LOX-deficient (Alox15) mice developed a myeloid leukemia instigated my studies since leukemias are often a consequence of dysregulated hematopoiesis. This observation lead to the central hypothesis of this dissertation which is that polyunsaturated fatty acid metabolism mediated by 12/15-LOX participates in hematopoietic development. Using genetic mouse models and in vitro and in vivo cell development assays, I found that 12/15-LOX indeed regulates multiple stages of hematopoiesis including the function of hematopoietic stem cells (HSC) and the differentiation of B cells, T cells, basophils, granulocytes and monocytes. -

Site-Specific Mutagenesis of the Calcium-Binding Photoprotein
Proc. Natd. Acad. Sci. USA Vol. 83, pp. 8107-8111, November 1986 Biochemistry Site-specific mutagenesis of the calcium-binding photoprotein aequorin (bioluninescence/oxygenase/oligonudeotide-directed mutagenesis/protein modification) FREDERICK I. TsuJI*, SATOSHI INOUYEtt, TOSHIO GOTO§, AND YOSHIYUKI SAKAKIt *Marine Biology Research Division, Scripps Institution of Oceanography, University of California-San Diego, La Jolla, CA 92093, and V. A. Medical Center Brentwood, Los Angeles, CA 90073; tResearch Laboratory for Genetic Information, Kyushu University, Fukuoka 812, Japan; and §Department of Agricultural Chemistry, Faculty of Agriculture, Nagoya University, Nagoya 464, Japan Communicated by Martin D. Kamen, July 21, 1986 ABSTRACT The luminescent protein aequorin from the which amino acid substitutions were made at the three jellyfish Aequoria victoria emits light by an intramolecular Ca2l-binding sites, at the three cysteines, and at histidine in reaction in the presence of a trace amount of Ca2 . In order to the hydrophobic region. understand the mechanism of the reaction, a study of structure-function relationships was undertaken with respect MATERIALS AND METHODS to modifying certain of its amino acid residues. This was done by carrying out oligonudeotide-directed site-specific mutagen- Enzymes and Chemicals. All restriction endonucleases, esis of apoaequorin cDNA and expressing the mutagenized Escherichia coli T4 DNA ligase, Klenow enzyme of DNA cDNA in Escherichia cofi. Amino acid substitutions were made polymerase 1, and T4 polynucleotide kinase were purchased at the three Ca2+-binding sites, the three cysteines, and a from Takara Shuzo (Kyoto, Japan). Tris, 2-mercaptoethanol, histidine in one of the hydrophobic regions. Subsequent assay and isopropyl-,fD-thiogalactopyranoside were obtained from of the modified aequorin showed that the Ca2 -binding sites, Wako Pure Chemicals (Osaka, Japan). -
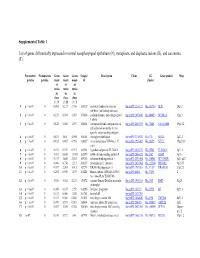
Supplemental Table 1 List of Genes Differentially Expressed In
Supplemental Table 1 List of genes differentially expressed in normal nasopharyngeal epithelium (N), metaplastic and displastic lesions (R), and carcinoma (T). Parametric Permutation Geom Geom Geom Unique Description Clone UG Gene symbol Map p-value p-value mean mean mean id cluster of of of ratios ratios ratios in in in class class class 1 : N 2 : R 3 : T 1 p < 1e-07 0 0.061 0.123 2.708 169329 secretory leukocyte protease IncytePD:2510171 Hs.251754 SLPI 20q12 inhibitor (antileukoproteinase) 2 p < 1e-07 0 0.125 0.394 1.863 163628 sodium channel, nonvoltage-gated IncytePD:1453049 Hs.446415 SCNN1A 12p13 1 alpha 3 p < 1e-07 0 0.122 0.046 1.497 160401 carcinoembryonic antigen-related IncytePD:2060355 Hs.73848 CEACAM6 19q13.2 cell adhesion molecule 6 (non- specific cross reacting antigen) 4 p < 1e-07 0 0.675 1.64 5.594 165101 monoglyceride lipase IncytePD:2174920 Hs.6721 MGLL 3q21.3 5 p < 1e-07 0 0.182 0.487 0.998 166827 nei endonuclease VIII-like 1 (E. IncytePD:1926409 Hs.28355 NEIL1 15q22.33 coli) 6 p < 1e-07 0 0.194 0.339 0.915 162931 hypothetical protein FLJ22418 IncytePD:2816379 Hs.36563 FLJ22418 1p11.1 7 p < 1e-07 0 1.313 0.645 13.593 162399 S100 calcium binding protein P IncytePD:2060823 Hs.2962 S100P 4p16 8 p < 1e-07 0 0.157 1.445 2.563 169315 selenium binding protein 1 IncytePD:2591494 Hs.334841 SELENBP1 1q21-q22 9 p < 1e-07 0 0.046 0.738 1.213 160115 prominin-like 1 (mouse) IncytePD:2070568 Hs.112360 PROML1 4p15.33 10 p < 1e-07 0 0.787 2.264 3.013 167294 HRAS-like suppressor 3 IncytePD:1969263 Hs.37189 HRASLS3 11q12.3 11 p < 1e-07 0 0.292 0.539 1.493 168221 Homo sapiens cDNA FLJ13510 IncytePD:64451 Hs.37896 2 fis, clone PLACE1005146.