Supplementary Tables 2C-E
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Cellular Responses to Erbb-2 Overexpression in Human Mammary Luminal Epithelial Cells: Comparison of Mrna and Protein Expression
British Journal of Cancer (2004) 90, 173 – 181 & 2004 Cancer Research UK All rights reserved 0007 – 0920/04 $25.00 www.bjcancer.com Cellular responses to ErbB-2 overexpression in human mammary luminal epithelial cells: comparison of mRNA and protein expression SL White1, S Gharbi1, MF Bertani1, H-L Chan1, MD Waterfield1 and JF Timms*,1 1 Ludwig Institute for Cancer Research, Wing 1.1, Cruciform Building, Gower Street, London WCIE 6BT, UK Microarray analysis offers a powerful tool for studying the mechanisms of cellular transformation, although the correlation between mRNA and protein expression is largely unknown. In this study, a microarray analysis was performed to compare transcription in response to overexpression of the ErbB-2 receptor tyrosine kinase in a model mammary luminal epithelial cell system, and in response to the ErbB-specific growth factor heregulin b1. We sought to validate mRNA changes by monitoring changes at the protein level using a parallel proteomics strategy, and report a surprisingly high correlation between transcription and translation for the subset of genes studied. We further characterised the identified targets and relate differential expression to changes in the biological properties of ErbB-2-overexpressing cells. We found differential regulation of several key cell cycle modulators, including cyclin D2, and downregulation of a large number of interferon-inducible genes, consistent with increased proliferation of the ErbB-2- overexpressing cells. Furthermore, differential expression of genes involved in extracellular matrix modelling and cellular adhesion was linked to altered adhesion of these cells. Finally, we provide evidence for enhanced autocrine activation of MAPK signalling and the AP-1 transcription complex. -

Integration of 198 Chip-Seq Datasets Reveals Human Cis-Regulatory Regions
Integration of 198 ChIP-seq datasets reveals human cis-regulatory regions Hamid Bolouri & Larry Ruzzo Thanks to Stephen Tapscott, Steven Henikoff & Zizhen Yao Slides will be available from: http://labs.fhcrc.org/bolouri/ Email [email protected] for manuscript (Bolouri & Ruzzo, submitted) Kleinjan & van Heyningen, Am. J. Hum. Genet., 2005, (76)8–32 Epstein, Briefings in Func. Genom. & Protemoics, 2009, 8(4)310-16 Regulation of SPi1 (Sfpi1, PU.1 protein) expression – part 1 miR155*, miR569# ~750nt promoter ~250nt promoter The antisense RNA • causes translational stalling • has its own promoter • requires distal SPI1 enhancer • is transcribed with/without SPI1. # Hikami et al, Arthritis & Rheumatism, 2011, 63(3):755–763 * Vigorito et al, 2007, Immunity 27, 847–859 Ebralidze et al, Genes & Development, 2008, 22: 2085-2092. Regulation of SPi1 expression – part 2 (mouse coordinates) Bidirectional ncRNA transcription proportional to PU.1 expression PU.1/ELF1/FLI1/GLI1 GATA1 GATA1 Sox4/TCF/LEF PU.1 RUNX1 SP1 RUNX1 RUNX1 SP1 ELF1 NF-kB SATB1 IKAROS PU.1 cJun/CEBP OCT1 cJun/CEBP 500b 500b 500b 500b 500b 750b 500b -18Kb -14Kb -12Kb -10Kb -9Kb Chou et al, Blood, 2009, 114: 983-994 Hoogenkamp et al, Molecular & Cellular Biology, 2007, 27(21):7425-7438 Zarnegar & Rothenberg, 2010, Mol. & cell Biol. 4922-4939 An NF-kB binding-site variant in the SPI1 URE reduces PU.1 expression & is GGGCCTCCCC correlated with AML GGGTCTTCCC Bonadies et al, Oncogene, 2009, 29(7):1062-72. SATB1 binding site A distal single nucleotide polymorphism alters long- range regulation of the PU.1 gene in acute myeloid leukemia Steidl et al, J Clin Invest. -

The MER41 Family of Hervs Is Uniquely Involved in the Immune-Mediated Regulation of Cognition/Behavior-Related Genes
bioRxiv preprint doi: https://doi.org/10.1101/434209; this version posted October 3, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. The MER41 family of HERVs is uniquely involved in the immune-mediated regulation of cognition/behavior-related genes: pathophysiological implications for autism spectrum disorders Serge Nataf*1, 2, 3, Juan Uriagereka4 and Antonio Benitez-Burraco 5 1CarMeN Laboratory, INSERM U1060, INRA U1397, INSA de Lyon, Lyon-Sud Faculty of Medicine, University of Lyon, Pierre-Bénite, France. 2 University of Lyon 1, Lyon, France. 3Banque de Tissus et de Cellules des Hospices Civils de Lyon, Hôpital Edouard Herriot, Lyon, France. 4Department of Linguistics and School of Languages, Literatures & Cultures, University of Maryland, College Park, USA. 5Department of Spanish, Linguistics, and Theory of Literature (Linguistics). Faculty of Philology. University of Seville, Seville, Spain * Corresponding author: [email protected] bioRxiv preprint doi: https://doi.org/10.1101/434209; this version posted October 3, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. ABSTRACT Interferon-gamma (IFNa prototypical T lymphocyte-derived pro-inflammatory cytokine, was recently shown to shape social behavior and neuronal connectivity in rodents. STAT1 (Signal Transducer And Activator Of Transcription 1) is a transcription factor (TF) crucially involved in the IFN pathway. -

Comprehensive Epigenomic Profiling of Human Alveolar Epithelial
bioRxiv preprint doi: https://doi.org/10.1101/2021.04.14.439786; this version posted April 15, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. FULL TITLE: Comprehensive epigenomic profiling of human alveolar epithelial differentiation identifies key epigenetic states and transcription factor co-regulatory networks for maintenance of distal lung identity SHORT TITLE: Epigenomic profiling of human alveolar epithelial differentiation Beiyun Zhou1,2,3, Theresa Ryan Stueve3,4,5, Evan A. Mihalakakos3,4,5, Lin Miao3,4,5, Daniel J Mullen3,4,5, Yinchong Wang3,4,5, Yixin Liu1, Jiao Luo1, Evelyn Tran3,4,5, Kim D. Siegmund6, Sean K. Lynch7, Amy L. Ryan1,2,8, Ite A. Offringa3,4,5,#, Zea Borok1,2,3,5,9,#, Crystal N. Marconett3,4,5* 1Division of Pulmonary, Critical Care and Sleep Medicine, Department of Medicine, 2Hastings Center for Pulmonary Research, 3Norris Comprehensive Cancer Center, Departments of 4Surgery, 5Biochemistry and Molecular Medicine, and 6Preventive Medicine, Keck School of Medicine, University of Southern California, USA 90089 7Department of Engineering, Test Manufacturing Group, MAXIM Integrated Products, Sunnyvale, California, USA 95134 8Department of Stem Cell Biology and Regenerative Medicine, University of Southern California, Los Angeles, CA USA 90089 9Division of Pulmonary, Critical Care and Sleep Medicine, Department of Medicine, University of California, San Diego, La Jolla, CA USA 92093. #These authors contributed equally * Corresponding/Senior author contact information: Crystal Marconett, Ph.D., Norris Comprehensive Cancer Center, Keck School of Medicine, University of Southern California, NTT 6418A M/C 9176, Los Angeles CA 90089-9520, USA. -

AP-1 Imprints a Reversible Transcriptional Program of Senescent Cells
bioRxiv preprint doi: https://doi.org/10.1101/633594; this version posted May 9, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. AP-1 Imprints a Reversible Transcriptional Program of Senescent Cells Ricardo Iván Martínez-Zamudio1,5,8, Pierre-François Roux1,5, José Américo N L F de Freitas1,4, Lucas Robinson1,4, Gregory Doré1, Bin Sun6,7, Jesús Gil6,7, Utz Herbig8 and Oliver Bischof1,2,3,9* 1 Institut Pasteur, Laboratory of Nuclear Organization and Oncogenesis, Department of Cell Biology and Infection, 75015 Paris, France 2 INSERM, U993, 75015 Paris, France 3 Equipe Labellisée Fondation ARC pour la recherche sur le cancer, Paris, France 4 Université de Paris, Sorbonne Paris Cité, Paris, France 5 These authors contributed equally to this work 6 MRC London Institute of Medical Sciences (LMS), Du Cane Road, London, W12 0NN, UK. 7 Institute of Clinical Sciences (ICS), Faculty of Medicine, Imperial College London, Du Cane Road, London W12 0NN, UK. 8 Department of Microbiology, Biochemistry & Molecular Genetics, Rutgers Biomedical & Health Sciences, Rutgers University, 205 South Orange Avenue, Newark, NJ 07103, USA 9 Lead contact *Correspondence: [email protected] bioRxiv preprint doi: https://doi.org/10.1101/633594; this version posted May 9, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. SUMMARY Senescent cells play important physiological- and pathophysiological roles in tumor suppression, tissue regeneration, and aging. -

Brd4-Bound Enhancers Drive Cell-Intrinsic Sex Differences in Glioblastoma
Brd4-bound enhancers drive cell-intrinsic sex differences in glioblastoma Najla Kfourya,b,1, Zongtai Qic,d,1, Briana C. Pragere,f, Michael N. Wilkinsonc,d, Lauren Broestla,g, Kristopher C. Berretth, Arnav Moudgilc,d,g, Sumithra Sankararamanc,d, Xuhua Chenc,d, Jason Gertzh, Jeremy N. Riche,i, Robi D. Mitrac,d,2,3, and Joshua B. Rubina,j,2,3 aDepartment of Pediatrics, School of Medicine, Washington University in St. Louis, St. Louis, MO 63110; bDepartment of Neurological Surgery, University of California San Diego, La Jolla, CA 92037; cDepartment of Genetics, School of Medicine, Washington University in St. Louis, St. Louis, MO 63110; dCenter for Genome Sciences and Systems Biology, Washington University in St. Louis, St. Louis, MO 63110; eDivision of Regenerative Medicine, Department of Medicine, University of California San Diego, La Jolla, CA 92037; fCleveland Clinic Lerner College of Medicine, Cleveland, OH 44195; gMedical Scientist Training Program, School of Medicine, Washington University in St. Louis, St. Louis, MO 63110; hDepartment of Oncological Sciences, Huntsman Cancer Institute, University of Utah, Salt Lake City, UT 84112; iDepartment of Neurosciences, University of California San Diego, La Jolla, CA 92037; and jDepartment of Neuroscience, School of Medicine, Washington University in St. Louis, St. Louis, MO 63110 Edited by Rene Bernards, The Netherlands Cancer Institute, Amsterdam, The Netherlands, and approved December 25, 2020 (received for review August 21, 2020) Sex can be an important determinant of cancer phenotype, and cancer risk and progression, with the ultimate goal of incorporating exploring sex-biased tumor biology holds promise for identifying sex-informed approaches to treatment to improve survival of all novel therapeutic targets and new approaches to cancer treatment. -

Identification of Genomic Targets of Krüppel-Like Factor 9 in Mouse Hippocampal
Identification of Genomic Targets of Krüppel-like Factor 9 in Mouse Hippocampal Neurons: Evidence for a role in modulating peripheral circadian clocks by Joseph R. Knoedler A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy (Neuroscience) in the University of Michigan 2016 Doctoral Committee: Professor Robert J. Denver, Chair Professor Daniel Goldman Professor Diane Robins Professor Audrey Seasholtz Associate Professor Bing Ye ©Joseph R. Knoedler All Rights Reserved 2016 To my parents, who never once questioned my decision to become the other kind of doctor, And to Lucy, who has pushed me to be a better person from day one. ii Acknowledgements I have a huge number of people to thank for having made it to this point, so in no particular order: -I would like to thank my adviser, Dr. Robert J. Denver, for his guidance, encouragement, and patience over the last seven years; his mentorship has been indispensable for my growth as a scientist -I would also like to thank my committee members, Drs. Audrey Seasholtz, Dan Goldman, Diane Robins and Bing Ye, for their constructive feedback and their willingness to meet in a frequently cold, windowless room across campus from where they work -I am hugely indebted to Pia Bagamasbad and Yasuhiro Kyono for teaching me almost everything I know about molecular biology and bioinformatics, and to Arasakumar Subramani for his tireless work during the home stretch to my dissertation -I am grateful for the Neuroscience Program leadership and staff, in particular -
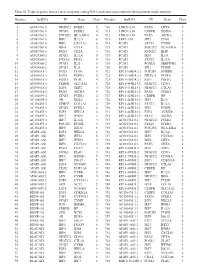
Table SI. Triple Negative Breast Cancer Long Non‑Coding RNA‑Mediated Transcriptional Dysregulation Triplet Network
Table SI. Triple negative breast cancer long non‑coding RNA‑mediated transcriptional dysregulation triplet network. Number lncRNA TF Gene Class Number lncRNA TF Gene Class 1 AC010761.8 NFATC2 FCER2 5 710 LINC01139 PAX5 CIITA 5 2 AC010761.8 STAT6 FCER2 4 711 LINC01139 CEBPB DDIT4 6 3 AC010761.8 POU2F2 HLA‑DRA 4 712 LINC01139 PAX5 AICDA 3 4 AC010761.8 IRF7 IL12A 6 713 LEF1‑AS1 SPI1 CD68 5 5 AC010761.8 IRF1 CXCL10 6 714 PCAT1 STAT1 TYMP 4 6 AC010761.8 IRF4 CIITA 5 715 PCAT1 POU2F2 HLA‑DRA 5 7 AC010761.8 PAX5 CIITA 4 716 PCAT1 RUNX2 IBSP 3 8 AC024560.2 STAT1 IL12A 5 717 PCAT1 IRF1 IL10 5 9 AC024560.2 FOXA2 PDX1 4 718 PCAT1 STAT2 IL12A 4 10 AC024560.2 STAT1 IL21 5 719 PCAT1 FOXD1 SERPINE1 3 11 AC024560.2 STAT2 IL21 5 720 PCAT1 SPI1 PTGIR 6 12 AC093818.1 PAX5 CD19 5 721 RP11‑169D4.2.1 POU2F1 GNRHR 3 13 AC093818.1 PAX5 FCER2 5 722 RP11‑169D4.2.1 NKX2‑5 PLOD1 4 14 AC093818.1 FOSL1 PLAU 4 723 RP11‑169D4.2.1 E2F1 POLE2 6 15 AC093818.1 CEBPB SLC11A1 6 724 RP11‑445K13.2 NFATC2 CD3G 5 16 AC093818.1 E2F1 TERT 3 725 RP11‑445K13.2 NFATC2 CTLA4 5 17 AC093818.1 PAX5 AICDA 5 726 RP11‑445K13.2 PAX5 FCER2 5 18 AC104699.1 STAT6 FCER2 4 727 RP11‑445K13.2 CEBPB HP 3 19 AC104699.1 AR KLK4 4 728 RP11‑445K13.2 IRF1 IL10 5 20 AC156455.1 CEBPZ COL1A1 2 729 RP11‑445K13.2 STAT1 IL12A 5 21 AC156455.1 STAT6 FCER2 4 730 RP11‑445K13.2 SPI1 PTGIR 6 22 AC156455.1 E2F1 HELLS 6 731 RP11‑445K13.2 E2F4 AURKB 6 23 AC156455.1 IRF7 IFNB1 3 732 RP11‑445K13.2 PAX5 AICDA 5 24 AC156455.1 IRF7 IL12A 3 733 AC010761.8.1 NFATC2 FCER2 5 25 AC156455.1 POU2F1 PTGIR 3 734 AC010761.8.1 STAT6 -

Supplementary Data
Supplementary Figure 1 Supplementary Figure 2 CCR-10-3244.R1 Supplementary Figure Legends Supplementary Figure 1. B-Myb is overexpressed in primary AML blasts and B-CLL cells. Baseline B-Myb mRNA levels were determined by quantitative RT-PCR, after normalization to the level of housekeeping gene, in primary B-CLL (n=10) and AML (n=5) patient samples, and in normal CD19+ (n=5) and CD34+ (n=4) cell preparations. Each sample was determined in triplicate. Horizontal bars are median, upper and lower edges of box are 75th and 25th percentiles, lines extending from box are 10th and 90th percentiles. Supplementary Figure 2. Cytotoxicity by Nutlin-3 and Chlorambucil used alone or in combination in leukemic cells. The p53wild-type EHEB and SKW6.4 cells lines, and the p53mutated BJAB cell line were exposed to Nutlin-3 or Chlorambucil used either alone or in combination. (Nutl.+Chlor.). In A, upon treatment with Nutlin-3 or Chlorambucil, used either alone (both at 10 μM) or in combination (Nutl.+Chlor.), induction of apoptosis was quantitatively evaluated by Annexin V/PI staining, while E2F1 and pRb protein levels were analyzed by Western blot. Tubulin staining is shown as loading control. The average combination index (CI) values (analyzed by the method of Chou and Talalay) for effects of Chlorambucil+Nutlin-3 on cell viability are shown. ED indicates effect dose. In B, levels of B-Myb and E2F1 mRNA were analyzed by quantitative RT- PCR. Results are expressed as fold of B-Myb and E2F1 modulation in cells treated for 24 hours as indicated, with respect to the control untreated cultures set to 1 (hatched line). -

Identification and Analysis of Key Genes in Osteosarcoma Using Bioinformatics
ONCOLOGY LETTERS 15: 2789-2794, 2018 Identification and analysis of key genes in osteosarcoma using bioinformatics CHUNYU DIAO1, YONG XI2 and TAO XIAO1 1Department of Orthopedics, The Second Xiangya Hospital of Central South University, Changsha, Hunan 410011; 2Department of Orthopedics, Tongchuan People's Hospital, Tongchuan, Shaanxi 727000, P.R. China Received August 26, 2016; Accepted June 21, 2017 DOI: 10.3892/ol.2017.7649 Abstract. Osteosarcoma (OS) is an invasive malignant were identified between the metastatic and non‑metastatic neoplasm of the bones. The present study identified and samples. In the PPI network, upregulated epidermal growth analyzed key genes associated with OS. Expression profiling factor receptor (EGFR) exhibits a high degree and was there- of the dataset GSE49003, which included 6 metastatic and fore highly interconnected with other proteins. Enrichment 6 non-metastatic OS cell lines and was obtained from the analysis revealed that EGFR was enriched in cytoskeleton Gene Expression Omnibus, was performed. Following data organization, organic substance response and the signaling preprocessing, the differentially expressed genes (DEGs) pathway of focal adhesion. The TFs early growth response 1, were selected using the limma package in R. Subsequently, nuclear factor-κB complex subunits, peroxisome proliferator bidirectional hierarchical clustering using the pheatmap activated receptor α, signal transducer and activator of tran- package in R and an unpaired Students' t-test was performed scription 3 and MYC proto‑oncogene were identified in the for the DEGs. Based on the Search Tool for the Retrieval EGFR promoter region. Furthermore, multiple CpG islands, of Interacting Genes database and Cytoscape software, a starting from the 400 bp of the EGFR promoter sequence, protein-protein interaction (PPI) network for the DEGs was were predicted. -

Identification of Candidate Genes for Myeloma-Induced Osteocyte Death Based on Microarray Data Honglai Tian
Tian Journal of Orthopaedic Surgery and Research (2016) 11:81 DOI 10.1186/s13018-016-0411-0 RESEARCH ARTICLE Open Access Identification of candidate genes for myeloma-induced osteocyte death based on microarray data Honglai Tian Abstract Background: The study was aimed to investigate the molecular mechanisms of osteocyte death in multiple myeloma (MM) patients. Methods: GSE27372 was downloaded from Gene Expression Omnibus, including three HOB-01 (osteocyte cell line) control samples and three HOB-01 samples co-cultured with JJN3 (human MM cell line). After the differentially expressed genes (DEGs) were identified by Student’s t test method, enrichment analyses were performed for them using DAVID software. Using TRANSFAC, TSGene, and tumor-associated gene (TAG) databases, functional annotation was conducted for the DEGs. Additionally, protein-protein interaction (PPI) network and sub-network analyses were performed using STRING database and Cytoscape software. Results: Total 393 DEGs were identified, including 22 transcription factors (e.g., KLF4 and IRF8) and 37 TAGs. Enrichment analysis suggested that EGF, S1PR1, and NPY1R were enriched in the function of circulatory system development. EGF (degree = 31) and EGR1 (degree = 19) had high degrees and interactions in the PPI network. In the sub-network, S1PR1, C3AR1, and NPY1R could interact with each other. Conclusions: These DEGs might participate in the osteocyte apoptosis induced by myeloma cells. These findings might provide a theoretical basis for a better understanding of the osteolysis in MM patients. Keywords: Multiple myeloma, Osteocyte, Differentially expressed genes, Enrichment analysis, Protein-protein interaction network Background fully understand the potential molecular mechanisms of Multiple myeloma (MM) originates in neoplastic plasma osteolysis in MM patients. -

Supplementary Information for Inflammatory and Antiviral
Supplementary Information for Inflammatory and antiviral gene expression in Add Health: Molecular pathways to social disparities in disease emerge by young adulthood Steven W. Cole, Michael J. Shanahan, Lauren Gaydosh, & Kathleen Mullan Harris Corresponding authors: Kathleen Mullan Harris and Steven W. Cole Email: [email protected], [email protected] This PDF file includes: Supplementary text Figures S1 to S2 Tables S1 to S2 Legends for Datasets S1 to S2 SI References Other supplementary materials for this manuscript include the following: Datasets S1 to S2 www.pnas.org/cgi/doi/10.1073/pnas.1821367117 Methods Sample and survey procedures. Data come from the National Longitudinal Study of Adolescent to Adult Health (Add Health), a nationally representative study of U.S. adolescents in grades 7-12 at Wave I in 1994-1995 and followed into adulthood over five waves of data collection. We analyze data from the nationally representative Sample 1 subsample of Wave V, conducted in 2016-2017 when respondents were aged 32-42. Add Health administered Wave V using continuous interviewing over three years (2016-2018), and the sampling design selected 3 random subsamples of eligible Wave V respondents for interview in each year. The subsample of 1126 participants analyzed here are those who consented to provide a blood specimen for RNA analysis during the Sample 1 physical examination visit. Add Health developed this three subsample design in order to release preliminary Wave V data prior to the entire Wave V sample being interviewed (given it took 3 years) and released Sample 1 survey data in 2017. Sample 1 therefore represents a nationally representative sample because a randomly-selected subsample of a nationally-representative sample is also nationally representative (1).