Novel Immunotherapeutic Strategies For
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genome-Wide DNA Methylation Profiling Identifies Differential Methylation in Uninvolved Psoriatic Epidermis
Genome-Wide DNA Methylation Profiling Identifies Differential Methylation in Uninvolved Psoriatic Epidermis Deepti Verma, Anna-Karin Ekman, Cecilia Bivik Eding and Charlotta Enerbäck The self-archived postprint version of this journal article is available at Linköping University Institutional Repository (DiVA): http://urn.kb.se/resolve?urn=urn:nbn:se:liu:diva-147791 N.B.: When citing this work, cite the original publication. Verma, D., Ekman, A., Bivik Eding, C., Enerbäck, C., (2018), Genome-Wide DNA Methylation Profiling Identifies Differential Methylation in Uninvolved Psoriatic Epidermis, Journal of Investigative Dermatology, 138(5), 1088-1093. https://doi.org/10.1016/j.jid.2017.11.036 Original publication available at: https://doi.org/10.1016/j.jid.2017.11.036 Copyright: Elsevier http://www.elsevier.com/ Genome-Wide DNA Methylation Profiling Identifies Differential Methylation in Uninvolved Psoriatic Epidermis Deepti Verma*a, Anna-Karin Ekman*a, Cecilia Bivik Edinga and Charlotta Enerbäcka *Authors contributed equally aIngrid Asp Psoriasis Research Center, Department of Clinical and Experimental Medicine, Division of Dermatology, Linköping University, Linköping, Sweden Corresponding author: Charlotta Enerbäck Ingrid Asp Psoriasis Research Center, Department of Clinical and Experimental Medicine, Linköping University SE-581 85 Linköping, Sweden Phone: +46 10 103 7429 E-mail: [email protected] Short title Differential methylation in psoriasis Abbreviations CGI, CpG island; DMS, differentially methylated site; RRBS, reduced representation bisulphite sequencing Keywords (max 6) psoriasis, epidermis, methylation, Wnt, susceptibility, expression 1 ABSTRACT Psoriasis is a chronic inflammatory skin disease with both local and systemic components. Genome-wide approaches have identified more than 60 psoriasis-susceptibility loci, but genes are estimated to explain only one third of the heritability in psoriasis, suggesting additional, yet unidentified, sources of heritability. -
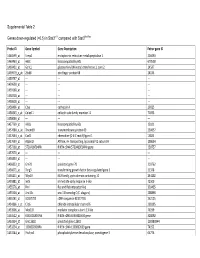
0.5) in Stat3∆/∆ Compared with Stat3flox/Flox
Supplemental Table 2 Genes down-regulated (<0.5) in Stat3∆/∆ compared with Stat3flox/flox Probe ID Gene Symbol Gene Description Entrez gene ID 1460599_at Ermp1 endoplasmic reticulum metallopeptidase 1 226090 1460463_at H60c histocompatibility 60c 670558 1460431_at Gcnt1 glucosaminyl (N-acetyl) transferase 1, core 2 14537 1459979_x_at Zfp68 zinc finger protein 68 24135 1459747_at --- --- --- 1459608_at --- --- --- 1459168_at --- --- --- 1458718_at --- --- --- 1458618_at --- --- --- 1458466_at Ctsa cathepsin A 19025 1458345_s_at Colec11 collectin sub-family member 11 71693 1458046_at --- --- --- 1457769_at H60a histocompatibility 60a 15101 1457680_a_at Tmem69 transmembrane protein 69 230657 1457644_s_at Cxcl1 chemokine (C-X-C motif) ligand 1 14825 1457639_at Atp6v1h ATPase, H+ transporting, lysosomal V1 subunit H 108664 1457260_at 5730409E04Rik RIKEN cDNA 5730409E04Rik gene 230757 1457070_at --- --- --- 1456893_at --- --- --- 1456823_at Gm70 predicted gene 70 210762 1456671_at Tbrg3 transforming growth factor beta regulated gene 3 21378 1456211_at Nlrp10 NLR family, pyrin domain containing 10 244202 1455881_at Ier5l immediate early response 5-like 72500 1455576_at Rinl Ras and Rab interactor-like 320435 1455304_at Unc13c unc-13 homolog C (C. elegans) 208898 1455241_at BC037703 cDNA sequence BC037703 242125 1454866_s_at Clic6 chloride intracellular channel 6 209195 1453906_at Med13l mediator complex subunit 13-like 76199 1453522_at 6530401N04Rik RIKEN cDNA 6530401N04 gene 328092 1453354_at Gm11602 predicted gene 11602 100380944 1453234_at -

4-6 Weeks Old Female C57BL/6 Mice Obtained from Jackson Labs Were Used for Cell Isolation
Methods Mice: 4-6 weeks old female C57BL/6 mice obtained from Jackson labs were used for cell isolation. Female Foxp3-IRES-GFP reporter mice (1), backcrossed to B6/C57 background for 10 generations, were used for the isolation of naïve CD4 and naïve CD8 cells for the RNAseq experiments. The mice were housed in pathogen-free animal facility in the La Jolla Institute for Allergy and Immunology and were used according to protocols approved by the Institutional Animal Care and use Committee. Preparation of cells: Subsets of thymocytes were isolated by cell sorting as previously described (2), after cell surface staining using CD4 (GK1.5), CD8 (53-6.7), CD3ε (145- 2C11), CD24 (M1/69) (all from Biolegend). DP cells: CD4+CD8 int/hi; CD4 SP cells: CD4CD3 hi, CD24 int/lo; CD8 SP cells: CD8 int/hi CD4 CD3 hi, CD24 int/lo (Fig S2). Peripheral subsets were isolated after pooling spleen and lymph nodes. T cells were enriched by negative isolation using Dynabeads (Dynabeads untouched mouse T cells, 11413D, Invitrogen). After surface staining for CD4 (GK1.5), CD8 (53-6.7), CD62L (MEL-14), CD25 (PC61) and CD44 (IM7), naïve CD4+CD62L hiCD25-CD44lo and naïve CD8+CD62L hiCD25-CD44lo were obtained by sorting (BD FACS Aria). Additionally, for the RNAseq experiments, CD4 and CD8 naïve cells were isolated by sorting T cells from the Foxp3- IRES-GFP mice: CD4+CD62LhiCD25–CD44lo GFP(FOXP3)– and CD8+CD62LhiCD25– CD44lo GFP(FOXP3)– (antibodies were from Biolegend). In some cases, naïve CD4 cells were cultured in vitro under Th1 or Th2 polarizing conditions (3, 4). -

Supp Table 6.Pdf
Supplementary Table 6. Processes associated to the 2037 SCL candidate target genes ID Symbol Entrez Gene Name Process NM_178114 AMIGO2 adhesion molecule with Ig-like domain 2 adhesion NM_033474 ARVCF armadillo repeat gene deletes in velocardiofacial syndrome adhesion NM_027060 BTBD9 BTB (POZ) domain containing 9 adhesion NM_001039149 CD226 CD226 molecule adhesion NM_010581 CD47 CD47 molecule adhesion NM_023370 CDH23 cadherin-like 23 adhesion NM_207298 CERCAM cerebral endothelial cell adhesion molecule adhesion NM_021719 CLDN15 claudin 15 adhesion NM_009902 CLDN3 claudin 3 adhesion NM_008779 CNTN3 contactin 3 (plasmacytoma associated) adhesion NM_015734 COL5A1 collagen, type V, alpha 1 adhesion NM_007803 CTTN cortactin adhesion NM_009142 CX3CL1 chemokine (C-X3-C motif) ligand 1 adhesion NM_031174 DSCAM Down syndrome cell adhesion molecule adhesion NM_145158 EMILIN2 elastin microfibril interfacer 2 adhesion NM_001081286 FAT1 FAT tumor suppressor homolog 1 (Drosophila) adhesion NM_001080814 FAT3 FAT tumor suppressor homolog 3 (Drosophila) adhesion NM_153795 FERMT3 fermitin family homolog 3 (Drosophila) adhesion NM_010494 ICAM2 intercellular adhesion molecule 2 adhesion NM_023892 ICAM4 (includes EG:3386) intercellular adhesion molecule 4 (Landsteiner-Wiener blood group)adhesion NM_001001979 MEGF10 multiple EGF-like-domains 10 adhesion NM_172522 MEGF11 multiple EGF-like-domains 11 adhesion NM_010739 MUC13 mucin 13, cell surface associated adhesion NM_013610 NINJ1 ninjurin 1 adhesion NM_016718 NINJ2 ninjurin 2 adhesion NM_172932 NLGN3 neuroligin -

Single-Cell Transcriptomics of Allo-Reactive CD4 T Cells Over Time
Single-cell transcriptomics of allo-reactive CD4+ T cells over time reveals divergent fates during gut GVHD Jessica A. Engel, … , Motoko Koyama, Ashraful Haque JCI Insight. 2020. https://doi.org/10.1172/jci.insight.137990. Research In-Press Preview Hematology Immunology Graphical abstract Find the latest version: https://jci.me/137990/pdf 1 Single-cell transcriptomics of allo-reactive CD4+ T cells over time reveals 2 divergent fates during gut GVHD. 3 4 Jessica A. Engel*1, Hyun Jae Lee*1, Cameron G. Williams*1, Rachel Kuns1, Stuart Olver1, 5 Lianne I. M. Lansink1, Megan S. F. Soon1, Stacey B. Andersen2, Joseph E. Powell3,4, Valentine 6 Svensson5, Sarah A. Teichmann6, Geoffrey R Hill1,7,8, Antiopi Varelias1, 9, Motoko Koyama7 7 and Ashraful Haque1,10. 8 9 Affiliations: 10 1QIMR Berghofer Medical Research Institute, Herston, Brisbane, Queensland 4006, Australia 11 2Institute for Molecular Bioscience, University of Queensland, St Lucia, Australia 12 3Garvan-Weizmann Centre for Cellular Genomics, Sydney, Australia. 13 4UNSW Cellular Genomics Futures Institute, University of New South Wales, Sydney, 14 Australia 15 5Flagship Labs 60 Inc, Cambridge, Mass, USA. 16 6Wellcome Sanger Institute, Wellcome Genome Campus, Hinxton, Cambridgeshire, UK 17 7Clinical Research Division, Fred Hutchinson Cancer Research Center, Seattle, WA 98109, 18 USA 19 8Division of Medical Oncology, University of Washington, Seattle, WA 98109, USA 20 9Faculty of Medicine, University of Queensland, St Lucia, Queensland, Australia 21 10Department of Microbiology and Immunology, University of Melbourne, located at The Peter 22 Doherty Institute for Infection and Immunity, 792 Elizabeth Street, Parkville, Victoria 3000, 23 Australia 24 25 *These authors contributed equally. -

Tetraspanins in the Regulation of Mast Cell Function
Medical Microbiology and Immunology (2020) 209:531–543 https://doi.org/10.1007/s00430-020-00679-x REVIEW Tetraspanins in the regulation of mast cell function Zane Orinska1 · Philipp M. Hagemann1 · Ivana Halova2 · Petr Draber2 Received: 17 February 2020 / Accepted: 6 May 2020 / Published online: 7 June 2020 © The Author(s) 2020 Abstract Mast cells (MCs) are long-living immune cells highly specialized in the storage and release of diferent biologically active compounds and are involved in the regulation of innate and adaptive immunity. MC degranulation and replacement of MC granules are accompanied by active membrane remodelling. Tetraspanins represent an evolutionary conserved family of transmembrane proteins. By interacting with lipids and other membrane and intracellular proteins, they are involved in organisation of membrane protein complexes and act as “molecular facilitators” connecting extracellular and cytoplasmic signaling elements. MCs express diferent tetraspanins and MC degranulation is accompanied by changes in membrane organisation. Therefore, tetraspanins are very likely involved in the regulation of MC exocytosis and membrane reorganisa- tion after degranulation. Antiviral response and production of exosomes are further aspects of MC function characterized by dynamic changes of membrane organization. In this review, we pay a particular attention to tetraspanin gene expression in diferent human and murine MC populations, discuss tetraspanin involvement in regulation of key MC signaling com- plexes, and analyze the potential contribution of tetraspanins to MC antiviral response and exosome production. In-depth knowledge of tetraspanin-mediated molecular mechanisms involved in diferent aspects of the regulation of MC response will be benefcial for patients with allergies, characterized by overwhelming MC reactions. -

T Cells Regulatory + CD25 + T Cell Targets by CD4 + Distinct Molecular
Distinct Molecular Program Imposed on CD4 + T Cell Targets by CD4+CD25+ Regulatory T Cells This information is current as Teresa L. Sukiennicki and Deborah J. Fowell of September 28, 2021. J Immunol 2006; 177:6952-6961; ; doi: 10.4049/jimmunol.177.10.6952 http://www.jimmunol.org/content/177/10/6952 Downloaded from References This article cites 52 articles, 25 of which you can access for free at: http://www.jimmunol.org/content/177/10/6952.full#ref-list-1 Why The JI? Submit online. http://www.jimmunol.org/ • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average by guest on September 28, 2021 Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2006 by The American Association of Immunologists All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Distinct Molecular Program Imposed on CD4؉ T Cell Targets by CD4؉CD25؉ Regulatory T Cells1 Teresa L. Sukiennicki and Deborah J. Fowell2 CD4؉CD25؉ regulatory T cells (Tregs) are key modulators of immunity, but their mechanism of action is unclear. To elucidate the molecular consequences of Treg encounter, we analyzed changes in gene expression in CD4؉ T cell targets activated in the presence or absence of CD4؉CD25؉ Tregs. -

Impaired Expression of Tetraspanin 32 (TSPAN32) in Memory T Cells of Patients with Multiple Sclerosis
brain sciences Article Impaired Expression of Tetraspanin 32 (TSPAN32) in Memory T Cells of Patients with Multiple Sclerosis Maria Sofia Basile 1, Emanuela Mazzon 2 , Katia Mangano 1 , Manuela Pennisi 1, Maria Cristina Petralia 2, Salvo Danilo Lombardo 1 , Ferdinando Nicoletti 1 , Paolo Fagone 1,* and Eugenio Cavalli 2 1 Department of Biomedical and Biotechnological Sciences, University of Catania, Via S. Sofia 89, 95123 Catania, Italy; sofi[email protected] (M.S.B.); [email protected] (K.M.); [email protected] (M.P.); [email protected] (S.D.L.); [email protected] (F.N.) 2 IRCCS Centro Neurolesi “Bonino-Pulejo”, Via Provinciale Palermo, Contrada Casazza, 98124 Messina, Italy; [email protected] (E.M.); [email protected] (M.C.P.); [email protected] (E.C.) * Correspondence: [email protected] Received: 29 November 2019; Accepted: 14 January 2020; Published: 17 January 2020 Abstract: Tetraspanins are a conserved family of proteins involved in a number of biological processes. We have previously shown that Tetraspanin-32 (TSPAN32) is significantly downregulated upon activation of T helper cells via anti-CD3/CD28 stimulation. On the other hand, TSPAN32 is marginally modulated in activated Treg cells. A role for TSPAN32 in controlling the development of autoimmune responses is consistent with our observation that encephalitogenic T cells from myelin oligodendrocyte glycoprotein (MOG)-induced experimental autoimmune encephalomyelitis (EAE) mice exhibit significantly lower levels of TSPAN32 as compared to naïve T cells. In the present study, by making use of ex vivo and in silico analysis, we aimed to better characterize the pathophysiological and diagnostic/prognostic role of TSPAN32 in T cell immunity and in multiple sclerosis (MS). -
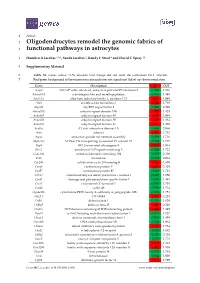
Oligodendrocytes Remodel the Genomic Fabrics of Functional Pathways in Astrocytes
1 Article 2 Oligodendrocytes remodel the genomic fabrics of 3 functional pathways in astrocytes 4 Dumitru A Iacobas 1,2,*, Sanda Iacobas 3, Randy F Stout 4 and David C Spray 2,5 5 Supplementary Material 6 Table S1. Genes whose >1.5x absolute fold-change did not meet the individual CUT criterion. 7 Red/green background of the expression ratio indicates not significant (false) up-/down-regulation. Gene Description X CUT Acap2 ArfGAP with coiled-coil, ankyrin repeat and PH domains 2 -1.540 1.816 Adamts18 a disintegrin-like and metallopeptidase -1.514 1.594 Akr1c12 aldo-keto reductase family 1, member C12 1.866 1.994 Alx3 aristaless-like homeobox 3 1.536 1.769 Alyref2 Aly/REF export factor 2 -1.880 2.208 Ankrd33b ankyrin repeat domain 33B 1.593 1.829 Ankrd45 ankyrin repeat domain 45 1.514 1.984 Ankrd50 ankyrin repeat domain 50 1.628 1.832 Ankrd61 ankyrin repeat domain 61 1.645 1.802 Arid1a AT rich interactive domain 1A -1.668 2.066 Artn artemin 1.524 1.732 Aspm abnormal spindle microtubule assembly -1.693 1.716 Atp6v1e1 ATPase, H+ transporting, lysosomal V1 subunit E1 -1.679 1.777 Bag4 BCL2-associated athanogene 4 1.723 1.914 Birc3 baculoviral IAP repeat-containing 3 -1.588 1.722 Ccdc104 coiled-coil domain containing 104 -1.819 2.130 Ccl2 chemokine -1.699 2.034 Cdc20b cell division cycle 20 homolog B 1.512 1.605 Cenpf centromere protein F 2.041 2.128 Cep97 centrosomal protein 97 -1.641 1.723 COX1 mitochondrially encoded cytochrome c oxidase I -1.607 1.650 Cpsf7 cleavage and polyadenylation specific factor 7 -1.635 1.891 Crct1 cysteine-rich -

Lineage-Specific Effector Signatures of Invariant NKT Cells Are Shared Amongst Δγ T, Innate Lymphoid, and Th Cells
Downloaded from http://www.jimmunol.org/ by guest on September 26, 2021 δγ is online at: average * The Journal of Immunology , 10 of which you can access for free at: 2016; 197:1460-1470; Prepublished online 6 July from submission to initial decision 4 weeks from acceptance to publication 2016; doi: 10.4049/jimmunol.1600643 http://www.jimmunol.org/content/197/4/1460 Lineage-Specific Effector Signatures of Invariant NKT Cells Are Shared amongst T, Innate Lymphoid, and Th Cells You Jeong Lee, Gabriel J. Starrett, Seungeun Thera Lee, Rendong Yang, Christine M. Henzler, Stephen C. Jameson and Kristin A. Hogquist J Immunol cites 41 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription http://www.jimmunol.org/content/suppl/2016/07/06/jimmunol.160064 3.DCSupplemental This article http://www.jimmunol.org/content/197/4/1460.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2016 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 26, 2021. The Journal of Immunology Lineage-Specific Effector Signatures of Invariant NKT Cells Are Shared amongst gd T, Innate Lymphoid, and Th Cells You Jeong Lee,* Gabriel J. -

Mirnas Documented to Play a Role in Hematopoietic Cell Lineage. Our
Table S1: miRNAs documented to play a role in hematopoietic cell lineage. Our review of the literature summarizing miRNAs known to be involved in the development and proliferation of the hematopoietic lineage cells. miRNA Expression/function/target/regulator References miR-150 Elevated during developmental stages of B and T cell maturation. 16-19 Controls B cell differentiation, present in mature, resting T and B cells but decreased upon activation of naïve T or B cells. Plays a role in establishing lymphocyte identity. Very little is known about function in T cells. Regulators: Foxp3 Target: C-Myb miR-146a/b Upregulated in macropgahe inflammatory response. Differentially 17, 20 upregulated in murine Th1 subset but abolished in Th2 subset. Upregulated in response to TCRs stimulation, as well as by IL-1 and TNF. Highly expressed in murine T-regs and could play a role in establishing lymphocyte identity. Modulates activation induced cell death in activated T cells. Negative regulator of TLR and cytokine signaling pathway. Endotoxin tolerance. Antiviral role. Targets: IRAK1, IRAK2, TRAF6, FAF1 miR-16-1 Promote apoptosis by targeting Bcl2 expression, act as tumor 22 cluster suppressor RNAs. May block differentiation of later stage hematopoietic progenitor cells to mature cells. Downregulated in CLL. Target: BCL2. miR-155 Regulator of T and B cell maturation and innate immune response. 23-29 Expressed in primary mediastinal B-cell lymphoma, T and B cells, macrophages and DCs. Upregulated during B cell activation. Involved in T cell differentiation and indicated as a positive regulator of cytokine production. Activated by stimulating TLR3 and INFab receptors in bone derived macrophages (regulation of antimicrobial defense). -

TSPAN32) Expression in T Cell-Mediated Immune Responses and in Multiple Sclerosis
International Journal of Molecular Sciences Article Modulation of Tetraspanin 32 (TSPAN32) Expression in T Cell-Mediated Immune Responses and in Multiple Sclerosis Salvo Danilo Lombardo 1 , Emanuela Mazzon 2, Maria Sofia Basile 1, Giorgia Campo 1, Federica Corsico 1, Mario Presti 1 , Placido Bramanti 2, Katia Mangano 1 , Maria Cristina Petralia 2, Ferdinando Nicoletti 1,* and Paolo Fagone 1 1 Department of Biomedical and Biotechnological Sciences, University of Catania, 95123 Catania, Italy 2 IRCCS Centro Neurolesi Bonino Pulejo, C.da Casazza, 98124 Messina, Italy * Correspondence: [email protected]; Tel.: +39-095-478-1270 Received: 30 July 2019; Accepted: 30 August 2019; Published: 4 September 2019 Abstract: Tetraspanins are a conserved family of proteins involved in a number of biological processes including, cell–cell interactions, fertility, cancer metastasis and immune responses. It has previously been shown that TSPAN32 knockout mice have normal hemopoiesis and B-cell responses, but hyperproliferative T cells. Here, we show that TSPAN32 is expressed at higher levels in the lymphoid lineage as compared to myeloid cells. In vitro activation of T helper cells via anti-CD3/CD28 is associated with a significant downregulation of TSPAN32. Interestingly, engagement of CD3 is sufficient to modulate TSPAN32 expression, and its effect is potentiated by costimulation with anti-CD28, but not anti-CTLA4, -ICOS nor -PD1. Accordingly, we measured the transcriptomic levels of TSPAN32 in polarized T cells under Th1 and Th2 conditions and TSPAN32 resulted significantly reduced as compared with unstimulated cells. On the other hand, in Treg cells, TSPAN32 underwent minor changes upon activation. The in vitro data were finally translated into the context of multiple sclerosis (MS).