Ancient Dna: a Multifunctional Tool for Resolving Anthropological Questions
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Mutation Rate Switch Inside Eurasian Mitochondrial Haplogroups: Impact of Selection and Consequences for Dating Settlement in Europe
Mutation Rate Switch inside Eurasian Mitochondrial Haplogroups: Impact of Selection and Consequences for Dating Settlement in Europe Denis Pierron1, Ivan Chang4, Amal Arachiche1, Margit Heiske1, Olivier Thomas1, Marine Borlin1, Erwan Pennarun5, Pacal Murail3, Didier Thoraval2, Christophe Rocher1, Thierry Letellier1* 1 Laboratoire de Physiopathologie Mitochondriale U688, INSERM - Universite´ Victor Segalen-Bordeaux 2, Bordeaux, France, 2 Institut de Biochimie et Ge´ne´tique Cellulaires UMR 5095, CNRS - Universite´ Victor Segalen-Bordeaux 2, Bordeaux, France, 3 Laboratoire d’Anthropologie des Populations du Passe´ PACEA UMR 5199, CNRS - Universite´ Bordeaux 1, Talence, France, 4 Institute of Genomic Biology, University of California Irvine, Irvine, California, United States of America, 5 Department of Evolutionary Biology, Institute of Molecular and Cell Biology, University of Tartu and Estonian Biocentre, Tartu, Estonia Abstract R-lineage mitochondrial DNA represents over 90% of the European population and is significantly present all around the planet (North Africa, Asia, Oceania, and America). This lineage played a major role in migration ‘‘out of Africa’’ and colonization in Europe. In order to determine an accurate dating of the R lineage and its sublineages, we analyzed 1173 individuals and complete mtDNA sequences from Mitomap. This analysis revealed a new coalescence age for R at 54.500 years, as well as several limitations of standard dating methods, likely to lead to false interpretations. These findings highlight the association of a striking under-accumulation of synonymous mutations, an over-accumulation of non- synonymous mutations, and the phenotypic effect on haplogroup J. Consequently, haplogroup J is apparently not a Neolithic group but an older haplogroup (Paleolithic) that was subjected to an underestimated selective force. -

The Polymerase Chain Reaction (PCR): the Second Generation of DNA Analysis Methods Takes the Stand, 9 Santa Clara High Tech
Santa Clara High Technology Law Journal Volume 9 | Issue 1 Article 8 January 1993 The olP ymerase Chain Reaction (PCR): The Second Generation of DNA Analysis Methods Takes the Stand Kamrin T. MacKnight Follow this and additional works at: http://digitalcommons.law.scu.edu/chtlj Part of the Law Commons Recommended Citation Kamrin T. MacKnight, The Polymerase Chain Reaction (PCR): The Second Generation of DNA Analysis Methods Takes the Stand, 9 Santa Clara High Tech. L.J. 287 (1993). Available at: http://digitalcommons.law.scu.edu/chtlj/vol9/iss1/8 This Comment is brought to you for free and open access by the Journals at Santa Clara Law Digital Commons. It has been accepted for inclusion in Santa Clara High Technology Law Journal by an authorized administrator of Santa Clara Law Digital Commons. For more information, please contact [email protected]. THE POLYMERASE CHAIN REACTION (PCR): THE SECOND GENERATION OF DNA ANALYSIS METHODS TAKES THE STAND Kamrin T. MacKnightt TABLE OF CONTENTS INTRODUCTION ........................................... 288 BASIC GENETICS AND DNA REPLICATION ................. 289 FORENSIC DNA ANALYSIS ................................ 292 Direct Sequencing ....................................... 293 Restriction FragmentLength Polymorphism (RFLP) ...... 294 Introduction .......................................... 294 Technology ........................................... 296 Polymerase Chain Reaction (PCR) ....................... 300 H istory ............................................... 300 Technology .......................................... -

Mittelalterliche Migrationen Als Gegenstand Der "Genetic History"
Jörg Feuchter Mittelalterliche Migrationen als Gegenstand der ‚Genetic History‘ Zusammenfassung Genetic History untersucht historische Fragen mit der Quelle DNA. Migrationen sind ihr Hauptgegenstand, woraus sich eine große Bedeutung für die Mediävistik ergibt. Doch bis vor kurzem waren Historiker nicht beteiligt. Der Beitrag gibt eine Definition der neuen Dis- ziplin (), untersucht die Rolle des Konzeptes Migration in der Populationsgenetik () und beschreibt den Stand der Migrationsforschung in der Mediävistik (). Anschließend gibt er einen kurzen Überblick über die von der Genetic History untersuchten Migrationsräu- me (), betrachtet exemplarisch zunächst eine Studie zur angelsächsischen Migration nach Britannien () und dann ein aktuelles Projekt, in dem erstmals ein Mediävist die Leitung innehat (). Der Beitrag endet mit einigen allgemeinen Beobachtungen und Postulaten (). Keywords: Genetik; DNA; Mittelalter; Angelsachsen; Langobarden; Völkerwanderung. Genetic History is the study of historical questions with DNA as a source. Migrations are its main subject. It is thus very relevant to Medieval Studies. Yet until recently historians have not been involved. The contribution provides a definition of the new discipline (), explores the role of migration as a concept in Population Genetics () and describes the state of migration studies in Medieval History (). It then sets out for an overview of medieval migratory areas studied by Genetic History () and takes an exemplary look first () at a study on Anglo-Saxon migration to Britain and later () at a current project where for the first time a medieval historian has taken the lead. The contribution ends () with some general observations and stipulations. Keywords: Genetics; DNA; Middle Ages; Anglo-Saxons; Lombards; Migration Period. Danksagung: Der Verfasser dankt Prof. Dr. Veronika Lipphardt (University College Frei- burg) und der von ihr von bis geleiteten Nachwuchsgruppe Twentieth Century Felix Wiedemann, Kerstin P. -

Ancient Mitochondrial DNA from Pre-Historic
Grand Valley State University ScholarWorks@GVSU Masters Theses Graduate Research and Creative Practice 4-30-2011 Ancient Mitochondrial DNA From Pre-historic Southeastern Europe: The rP esence of East Eurasian Haplogroups Provides Evidence of Interactions with South Siberians Across the Central Asian Steppe Belt Jeremy R. Newton Grand Valley State University Follow this and additional works at: http://scholarworks.gvsu.edu/theses Part of the Cell Biology Commons, and the Molecular Biology Commons Recommended Citation Newton, Jeremy R., "Ancient Mitochondrial DNA From Pre-historic Southeastern Europe: The rP esence of East Eurasian Haplogroups Provides Evidence of Interactions with South Siberians Across the Central Asian Steppe Belt" (2011). Masters Theses. 5. http://scholarworks.gvsu.edu/theses/5 This Thesis is brought to you for free and open access by the Graduate Research and Creative Practice at ScholarWorks@GVSU. It has been accepted for inclusion in Masters Theses by an authorized administrator of ScholarWorks@GVSU. For more information, please contact [email protected]. ANCIENT MITOCHONDRIAL DNA FROM PRE-HISTORIC SOUTH- EASTERN EUROPE: THE PRESENCE OF EAST EURASIAN HAPLOGROUPS PROVIDES EVIDENCE OF INTERACTIONS WITH SOUTH SIBERIANS ACROSS THE CENTRAL ASIAN STEPPE BELT A thesis submittal in partial fulfillment of the requirements for the degree of Master of Science By Jeremy R. Newton To Cell and Molecular Biology Department Grand Valley State University Allendale, MI April, 2011 “Not all those who wander are lost.” J.R.R. Tolkien iii ACKNOWLEDGEMENTS I would like to extend my sincerest thanks to every person who has motivated, directed, and encouraged me throughout this thesis project. I especially thank my graduate advisor, Dr. -
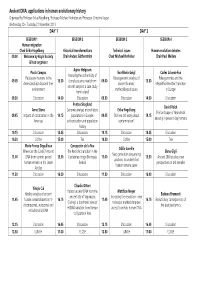
Final Programme
Ancient DNA: applications in human evolutionary history Organised by Professor Erika Hagelberg, Professor Michael Hofreiter and Professor Christine Keyser Wednesday 20 - Thursday 21 November 2013 DAY 1 DAY 2 SESSION 1 SESSION 2 SESSION 3 SESSION 4 Human migration Chair Erika Hagelberg Historical transformations Technical issues Human evolution debates 09.00 Welcome by Royal Society Chair Anders Götherström Chair Michael Hofreiter Chair Paul Mellars & lead organiser Agnar Helgason Paula Campos Eva-Maria Geigl Carles Lalueza-Fox Assessing the authenticity of Pleistocene humans in the Palaeogenomic analysis of Paleogenomics and the 09.05 13.30 clonal sequence reads from 09.00 13.30 Americans/East Asia and their ancient humans: Mesolithic-Neolithic transition ancient samples: a case study environment methodological issues in Europe from Iceland 09.30 Discussion 14.00 Discussion 09.30 Discussion 14.00 Discussion Pontus Skoglund David Reich Anne Stone Genomic analysis of prehistoric Erika Hagelberg The landscape of Neandertal 09.45 Impacts of colonization in the 14.15 populations in Europe: 09.45 Shall we still worry about 14.15 ancestry in present-day humans Americas authentication and population contamination? history 10.15 Discussion 14.45 Discussion 10.15 Discussion 14.45 Discussion 10.30 Coffee 15.00 Tea 10.30 Coffee 15.00 Tea Marie-France Deguilloux Concepción de la Rua Odille Loreille Where are the Caribs? Ancient The Neolithic transition in the Elena Gigli Next generation sequencing 11.00 DNA from ceramic period 15.30 Cantabrian fringe -

Profile of Alec J. Jeffreys
Profile of Alec J. Jeffreys s one of the great contributors to modern genetics, Sir Alec Jeffreys was born with curiosity in his genes as the son and Agrandson of prolific inventors. Jeffreys displayed an insatiable quest for knowl- edge, and his father fostered his son’s budding scientific interests with gifts of a microscope and chemistry set, the latter of which produced one of Jeffreys’ most memorable scientific ventures. ‘‘Those were back in the happy days of chemis- try,’’ Jeffreys points out, ‘‘where you could go down to your local pharmacist and get virtually everything you wanted.’’ The end result of that chemistry experiment was the detonation of his aunt’s apple tree and a set of scars that Jeffreys still bears to- day. ‘‘You learn science very fast that way,’’ he says, ‘‘but it was quite fun.’’ Jeffreys’ scientific curiosity only in- creased after the apple tree incident, and years later it would lead to one of the Alec J. Jeffreys most widely used applications in genetics: DNA fingerprinting. Among other uses, the DNA fingerprinting technique has freys go there, so Jeffreys entered the turn led to another groundbreaking find- helped solve numerous criminal investi- university in 1968. He was at first intent ing about the composition of eukaryotic gations, settle countless paternity dis- on pursuing a degree in biochemistry DNA: introns (3). ‘‘I was 27 at the time, putes, and spark a resurgence of interest but soon decided to alter his course. so still a real rookie, with the ability to in the forensic sciences. That achieve- ‘‘This was no criticism of the way it was detect single-copy DNA by Southern blot ment alone is worthy of merit, contrib- taught,’’ he says of biochemistry at Ox- hybridization and a nice paper on introns uting to Jeffreys’ receiving three high ford, ‘‘but rather a reflection of the fac- under my belt, which is, yeah, not a bad distinctions in 2005: the Albert Lasker ulty’s research, which leaned heavily to start,’’ he says. -

The DNA Fingerprinting Story Mark a Jobling
Jobling Investigative Genetics 2013, 4:20 http://www.investigativegenetics.com/content/4/1/20 OPINION Open Access Curiosity in the genes: the DNA fingerprinting story Mark A Jobling It is unusual for a scientific field to be associated with a The first DNA fingerprinting application was in par- single individual, but in the case of the subject of the the- entage testing [6]; normally it is the father who is in matic series now being launched in Investigative Genetics, doubt, but this unusual and challenging case was a ma- this is surely so; Alec Jeffreys (Figure 1) is DNA finger- ternity test, with paternal DNA unavailable. British nurse printing. Having invented the method, he coined the per- Christiana Sarbah’s 13-year-old son Andrew was denied fect name for it - how different things might have been if re-entry to the United Kingdom after a visit to Ghana, it had been called the tandem-repeat-based identification the immigration authorities suspecting that he was not technique (or something similarly dull). He realized its her child. Given three undisputed children for comparison, potential and immediately applied, developed and refined it was possible to reconstruct the absent father’sDNA it. He then followed his nose to unravel the mystery of fingerprint, and to strongly support the claimed maternity the madly mutable minisatellites that make up DNA over alternative relationships such as aunt-nephew - some- fingerprints, and eventually to understand the engines of thing that was not achieved with traditional protein poly- genome variability that reside in recombination hotspots. morphisms such as blood groups. -

Past Human Migrations in East Asia: Matching Archaeology, Linguistics
16 A genetic perspective on the origins and dispersal of the Austronesians Mitochondrial DNA variation from Madagascar to Easter Island Erika Hagelberg, Murray Cox, Wulf Schiefenhövel, and Ian Frame Introduction This chapter describes the study of one genetic system, mitochondrial DNA (mtDNA), within the Austronesian-speaking world, and discusses whether the patterns of genetic variation at this genetic locus are consistent with the principal models of the settlement and linguistic diversity in this vast geographical region. Mitochondria are the seat of cellular metabolism and have their own complement of DNA. MtDNA is a tiny proportion of the genetic makeup of a human being, and has both advantages and disadvantages in population studies. It is inherited maternally, in contrast with the nuclear chromosomal DNA inherited from both parents. (Note: the mitochondria of the spermatozoon are eliminated from the egg soon after fertilization, so offspring do not inherit paternal mtDNA, although there are exceptions to this rule, as shown by Schwartz and Vissing 2002.) The maternal mode of inheritance permits the study of female lineages through time. Despite the small size of the mitochondrial genome, each mitochondrion has many thousands of copies of mtDNA, and there are many mitochondria in each cell. The abundance of mtDNA in cells has been exploited in anthropological research on ancient, rare or degraded biological samples, including archaeological bones, old hair samples, or dried blood (Hagelberg 1994). MtDNA accumulates mutations at a relatively fast rate, and so is useful for the investigation of recent evolutionary events. These features have made mtDNA an excellent marker for geneticists interested in human evolutionary history (see Wallace 1995 for review). -

A Lecture by Professor Erika Hagelberg
H-Announce Is there such a thing as a Jewish genome? A lecture by Professor Erika Hagelberg Announcement published by Leah Sidebotham on Friday, April 1, 2016 Type: Lecture Date: April 12, 2016 Location: United Kingdom Subject Fields: Research and Methodology, Jewish History / Studies, Holocaust, Genocide, and Memory Studies, History of Science, Medicine, and Technology, Contemporary History New developments in DNA technology are having a huge impact on industry, medical science and forensic identification. They have also opened new opportunities for research in archaeology and human evolution, as we have seen in the recent identification of the skeleton of Richard III, and the advances in Neanderthal genetics. The technology is also aiding a growing industry in genetic genealogies, where human identity is defined in terms of patterns of DNA variants, obtained easily and cheaply from commercial companies, and giving rise to controversial notions, like that of a Jewish genome. In this talk Professor Hagelberg will describe the opportunities offered by human evolutionary genetics, as well as some of the limitations and more troubling aspects of research on genetic genealogies. Erika Hagelberg’s father came to the UK on a Kindertransport in 1939, aged 13. She studied biochemistry in London and gained a PhD at Cambridge. Her main field of research is the analysis of DNA from archaeological bones, and she has worked on several high-profile cases, including the identification of the skeletal remains of Joseph Mengele and of the Romanov family. She is professor of evolutionary biology at the University of Oslo, and Cheney Fellow in the Arts at the University of Leeds. -

Neolithic Mitochondrial Haplogroup H Genomes and the Genetic Origins of Europeans
ARTICLE Received 20 Sep 2012 | Accepted 27 Feb 2013 | Published 16 Apr 2013 DOI: 10.1038/ncomms2656 Neolithic mitochondrial haplogroup H genomes and the genetic origins of Europeans Paul Brotherton1,2,*, Wolfgang Haak1,*, Jennifer Templeton1, Guido Brandt3, Julien Soubrier1, Christina Jane Adler1,w, Stephen M. Richards1, Clio Der Sarkissian1,w, Robert Ganslmeier4, Susanne Friederich4, Veit Dresely4, Mannis van Oven5, Rosalie Kenyon6, Mark B. Van der Hoek6, Jonas Korlach7, Khai Luong7, Simon Y.W. Ho8, Lluis Quintana-Murci9, Doron M. Behar10, Harald Meller4,KurtW.Alt3, Alan Cooper1 & The Genographic Consortiumz Haplogroup H dominates present-day Western European mitochondrial DNA variability (440%), yet was less common (B19%) among Early Neolithic farmers (B5450 BC) and virtually absent in Mesolithic hunter-gatherers. Here we investigate this major component of the maternal population history of modern Europeans and sequence 39 complete haplogroup H mitochondrial genomes from ancient human remains. We then compare this ‘real-time’ genetic data with cultural changes taking place between the Early Neolithic (B5450 BC) and Bronze Age (B2200 BC) in Central Europe. Our results reveal that the current diversity and distribution of haplogroup H were largely established by the Mid Neolithic (B4000 BC), but with substantial genetic contributions from subsequent pan-European cultures such as the Bell Beakers expanding out of Iberia in the Late Neolithic (B2800 BC). Dated haplogroup H genomes allow us to reconstruct the recent evolutionary history of haplogroup H and reveal a mutation rate 45% higher than current estimates for human mitochondria. 1 The Australian Centre for Ancient DNA, School of Earth and Environmental Sciences, University of Adelaide, Adelaide, South Australia 5005, Australia. -

Human Maternal Heritage in Andalusia (Spain): Its Composition
Hernández et al. BMC Genetics 2014, 15:11 http://www.biomedcentral.com/1471-2156/15/11 RESEARCH ARTICLE Open Access Human maternal heritage in Andalusia (Spain): its composition reveals high internal complexity and distinctive influences of mtDNA haplogroups U6 and L in the western and eastern side of region Candela L Hernández1, Guillermo Reales1, Jean-Michel Dugoujon2, Andrea Novelletto3, Juan Nicolás Rodríguez4, Pedro Cuesta5 and Rosario Calderón1* Abstract Background: The archeology and history of the ancient Mediterranean have shown that this sea has been a permeable obstacle to human migration. Multiple cultural exchanges around the Mediterranean have taken place with presumably population admixtures. A gravitational territory of those migrations has been the Iberian Peninsula. Here we present a comprehensive analysis of the maternal gene pool, by means of control region sequencing and PCR-RFLP typing, of autochthonous Andalusians originating from the coastal provinces of Huelva and Granada, located respectively in the west and the east of the region. Results: The mtDNA haplogroup composition of these two southern Spanish populations has revealed a wide spectrum of haplogroups from different geographical origins. The registered frequencies of Eurasian markers, together with the high incidence and diversification of African maternal lineages (15% of the total mitochondrial variability) among Huelva Andalusians when compared to its eastwards relatives of Granada and other Iberian populations, constitute relevant findings unknown up-to-date on the characteristics of mtDNA within Andalusia that testifies a female population substructure. Therefore, Andalusia must not be considered a single, unique population. Conclusions: The maternal legacy among Andalusians reflects distinctive local histories, pointing out the role of the westernmost territory of Peninsular Spain as a noticeable recipient of multiple and diverse human migrations. -

DNA and the Dead Sea Scrolls
Putting the Pieces Together: DNA and the Dead Sea Scrolls Scott R. Woodward A number of questions concerning the origin and production of the Dead Sea Scrolls may be addressed using DNA analysis. These documents were for the most part written on what is thought to be goat- or sheepskin parchment. Based on radiocarbon and other analyses, these manuscripts date between the mid–second century BCE and the rst century CE.1 Under most conditions it would be remarkable that organic material, like parchment, would survive intact after such a long period of time. However, some of the material is remarkably well preserved because of the unique climate and storage conditions at Qumran. Because these parchments were produced from animal skins, they may contain remnant DNA molecules. Within the last decade, new techniques in molecular biology have been developed that have made it possible to recover DNA from ancient sources. The molecular analysis of ancient DNA (aDNA) from the Judean desert parchment fragments would enable us to establish a genetic signature unique for each manuscript. The precision of the DNA analysis will allow us to identify the species, population, and individual animal from which the parchment was produced. Background The ability to recover biomolecules, most importantly DNA, from ancient remains has opened new research that has many implications.2 Access to aDNA provides the opportunity to study the genetic material of past organisms and identify individual and population histories. Unfortunately, the DNA recovered from archaeological specimens is of such a degraded nature that the usual techniques associated with DNA ngerprinting cannot be used.