History of Macromolecular Crystallography in India Through
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Subtiligase-Catalyzed Peptide Ligation Amy M
Review Cite This: Chem. Rev. 2020, 120, 3127−3160 pubs.acs.org/CR Subtiligase-Catalyzed Peptide Ligation Amy M. Weeks*,† and James A. Wells*,†,‡ † Department of Pharmaceutical Chemistry, University of California, San Francisco, San Francisco, California 94143, United States ‡ Department of Cellular and Molecular Pharmacology, University of California, San Francisco, San Francisco, California 94143, United States ABSTRACT: Subtiligase-catalyzed peptide ligation is a powerful approach for site- specific protein bioconjugation, synthesis and semisynthesis of proteins and peptides, and chemoproteomic analysis of cellular N termini. Here, we provide a comprehensive review of the subtiligase technology, including its development, applications, and impacts on protein science. We highlight key advantages and limitations of the tool and compare it to other peptide ligase enzymes. Finally, we provide a perspective on future applications and challenges and how they may be addressed. CONTENTS 6.1. Subtiligase-Catalyzed Thioester and Thioa- cid Synthesis for Peptide and Protein 1. Introduction 3128 Bioconjugation 3138 2. Using Proteases in Reverse for Peptide Bond 6.2. Peptide Segment Condensation 3139 Formation 3129 6.3. Peptide Cyclization 3140 2.1. Protease-Catalyzed Peptide Bond Synthesis 6.4. Total Protein Synthesis 3140 under Thermodynamic Control 3129 7. Application of Subtiligase for Site-Specific 2.2. Protease-Catalyzed Peptide Bond Synthesis Protein Bioconjugation 3141 under Kinetic Control 3129 7.1. Sequence and Structural Requirements for 3. Protein Engineering of Subtilisin for Improved N-Terminal Modification by Subtiligase 3142 Peptide Bond Synthesis 3129 7.1.1. Characterization of Sequence and 3.1. Mutation of the Catalytic Serine to Cysteine 3129 Structural Requirements 3142 3.2. -

Cysteine Proteinases of Microorganisms and Viruses
ISSN 00062979, Biochemistry (Moscow), 2008, Vol. 73, No. 1, pp. 113. © Pleiades Publishing, Ltd., 2008. Original Russian Text © G. N. Rudenskaya, D. V. Pupov, 2008, published in Biokhimiya, 2008, Vol. 73, No. 1, pp. 317. REVIEW Cysteine Proteinases of Microorganisms and Viruses G. N. Rudenskaya1* and D. V. Pupov2 1Faculty of Chemistry and 2Faculty of Biology, Lomonosov Moscow State University, 119991 Moscow, Russia; fax: (495) 9393181; Email: [email protected] Received May 7, 2007 Revision received July 18, 2007 Abstract—This review considers properties of secreted cysteine proteinases of protozoa, bacteria, and viruses and presents information on the contemporary taxonomy of cysteine proteinases. Literature data on the structure and physicochemical and enzymatic properties of these enzymes are reviewed. High interest in cysteine proteinases is explained by the discovery of these enzymes mostly in pathogenic organisms. The role of the proteinases in pathogenesis of several severe diseases of human and animals is discussed. DOI: 10.1134/S000629790801001X Key words: cysteine proteinases, properties, protozoa, bacteria, viruses Classification and Catalytic Mechanism papain and related peptidases showed that the catalytic of Cysteine Proteinases residues are arranged in the following order in the polypeptide chain: Cys, His, and Asn. Also, a glutamine Cysteine proteinases are peptidyl hydrolases in residue preceding the catalytic cysteine is also important which the role of the nucleophilic group of the active site for catalysis. This residue is probably involved in the for is performed by the sulfhydryl group of a cysteine residue. mation of the oxyanion cavity of the enzyme. The cat Cysteine proteinases were first discovered and investigat alytic cysteine residue is usually followed by a residue of ed in tropic plants. -

Serine Proteases with Altered Sensitivity to Activity-Modulating
(19) & (11) EP 2 045 321 A2 (12) EUROPEAN PATENT APPLICATION (43) Date of publication: (51) Int Cl.: 08.04.2009 Bulletin 2009/15 C12N 9/00 (2006.01) C12N 15/00 (2006.01) C12Q 1/37 (2006.01) (21) Application number: 09150549.5 (22) Date of filing: 26.05.2006 (84) Designated Contracting States: • Haupts, Ulrich AT BE BG CH CY CZ DE DK EE ES FI FR GB GR 51519 Odenthal (DE) HU IE IS IT LI LT LU LV MC NL PL PT RO SE SI • Coco, Wayne SK TR 50737 Köln (DE) •Tebbe, Jan (30) Priority: 27.05.2005 EP 05104543 50733 Köln (DE) • Votsmeier, Christian (62) Document number(s) of the earlier application(s) in 50259 Pulheim (DE) accordance with Art. 76 EPC: • Scheidig, Andreas 06763303.2 / 1 883 696 50823 Köln (DE) (71) Applicant: Direvo Biotech AG (74) Representative: von Kreisler Selting Werner 50829 Köln (DE) Patentanwälte P.O. Box 10 22 41 (72) Inventors: 50462 Köln (DE) • Koltermann, André 82057 Icking (DE) Remarks: • Kettling, Ulrich This application was filed on 14-01-2009 as a 81477 München (DE) divisional application to the application mentioned under INID code 62. (54) Serine proteases with altered sensitivity to activity-modulating substances (57) The present invention provides variants of ser- screening of the library in the presence of one or several ine proteases of the S1 class with altered sensitivity to activity-modulating substances, selection of variants with one or more activity-modulating substances. A method altered sensitivity to one or several activity-modulating for the generation of such proteases is disclosed, com- substances and isolation of those polynucleotide se- prising the provision of a protease library encoding poly- quences that encode for the selected variants. -

Proteolytic Enzymes in Grass Pollen and Their Relationship to Allergenic Proteins
Proteolytic Enzymes in Grass Pollen and their Relationship to Allergenic Proteins By Rohit G. Saldanha A thesis submitted in fulfilment of the requirements for the degree of Masters by Research Faculty of Medicine The University of New South Wales March 2005 TABLE OF CONTENTS TABLE OF CONTENTS 1 LIST OF FIGURES 6 LIST OF TABLES 8 LIST OF TABLES 8 ABBREVIATIONS 8 ACKNOWLEDGEMENTS 11 PUBLISHED WORK FROM THIS THESIS 12 ABSTRACT 13 1. ASTHMA AND SENSITISATION IN ALLERGIC DISEASES 14 1.1 Defining Asthma and its Clinical Presentation 14 1.2 Inflammatory Responses in Asthma 15 1.2.1 The Early Phase Response 15 1.2.2 The Late Phase Reaction 16 1.3 Effects of Airway Inflammation 16 1.3.1 Respiratory Epithelium 16 1.3.2 Airway Remodelling 17 1.4 Classification of Asthma 18 1.4.1 Extrinsic Asthma 19 1.4.2 Intrinsic Asthma 19 1.5 Prevalence of Asthma 20 1.6 Immunological Sensitisation 22 1.7 Antigen Presentation and development of T cell Responses. 22 1.8 Factors Influencing T cell Activation Responses 25 1.8.1 Co-Stimulatory Interactions 25 1.8.2 Cognate Cellular Interactions 26 1.8.3 Soluble Pro-inflammatory Factors 26 1.9 Intracellular Signalling Mechanisms Regulating T cell Differentiation 30 2 POLLEN ALLERGENS AND THEIR RELATIONSHIP TO PROTEOLYTIC ENZYMES 33 1 2.1 The Role of Pollen Allergens in Asthma 33 2.2 Environmental Factors influencing Pollen Exposure 33 2.3 Classification of Pollen Sources 35 2.3.1 Taxonomy of Pollen Sources 35 2.3.2 Cross-Reactivity between different Pollen Allergens 40 2.4 Classification of Pollen Allergens 41 2.4.1 -

Supplementary Information
Supplementary information (a) (b) Figure S1. Resistant (a) and sensitive (b) gene scores plotted against subsystems involved in cell regulation. The small circles represent the individual hits and the large circles represent the mean of each subsystem. Each individual score signifies the mean of 12 trials – three biological and four technical. The p-value was calculated as a two-tailed t-test and significance was determined using the Benjamini-Hochberg procedure; false discovery rate was selected to be 0.1. Plots constructed using Pathway Tools, Omics Dashboard. Figure S2. Connectivity map displaying the predicted functional associations between the silver-resistant gene hits; disconnected gene hits not shown. The thicknesses of the lines indicate the degree of confidence prediction for the given interaction, based on fusion, co-occurrence, experimental and co-expression data. Figure produced using STRING (version 10.5) and a medium confidence score (approximate probability) of 0.4. Figure S3. Connectivity map displaying the predicted functional associations between the silver-sensitive gene hits; disconnected gene hits not shown. The thicknesses of the lines indicate the degree of confidence prediction for the given interaction, based on fusion, co-occurrence, experimental and co-expression data. Figure produced using STRING (version 10.5) and a medium confidence score (approximate probability) of 0.4. Figure S4. Metabolic overview of the pathways in Escherichia coli. The pathways involved in silver-resistance are coloured according to respective normalized score. Each individual score represents the mean of 12 trials – three biological and four technical. Amino acid – upward pointing triangle, carbohydrate – square, proteins – diamond, purines – vertical ellipse, cofactor – downward pointing triangle, tRNA – tee, and other – circle. -
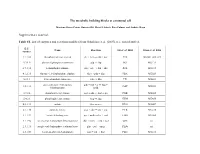
The Metabolic Building Blocks of a Minimal Cell Supplementary
The metabolic building blocks of a minimal cell Mariana Reyes-Prieto, Rosario Gil, Mercè Llabrés, Pere Palmer and Andrés Moya Supplementary material. Table S1. List of enzymes and reactions modified from Gabaldon et. al. (2007). n.i.: non identified. E.C. Name Reaction Gil et. al. 2004 Glass et. al. 2006 number 2.7.1.69 phosphotransferase system glc + pep → g6p + pyr PTS MG041, 069, 429 5.3.1.9 glucose-6-phosphate isomerase g6p ↔ f6p PGI MG111 2.7.1.11 6-phosphofructokinase f6p + atp → fbp + adp PFK MG215 4.1.2.13 fructose-1,6-bisphosphate aldolase fbp ↔ gdp + dhp FBA MG023 5.3.1.1 triose-phosphate isomerase gdp ↔ dhp TPI MG431 glyceraldehyde-3-phosphate gdp + nad + p ↔ bpg + 1.2.1.12 GAP MG301 dehydrogenase nadh 2.7.2.3 phosphoglycerate kinase bpg + adp ↔ 3pg + atp PGK MG300 5.4.2.1 phosphoglycerate mutase 3pg ↔ 2pg GPM MG430 4.2.1.11 enolase 2pg ↔ pep ENO MG407 2.7.1.40 pyruvate kinase pep + adp → pyr + atp PYK MG216 1.1.1.27 lactate dehydrogenase pyr + nadh ↔ lac + nad LDH MG460 1.1.1.94 sn-glycerol-3-phosphate dehydrogenase dhp + nadh → g3p + nad GPS n.i. 2.3.1.15 sn-glycerol-3-phosphate acyltransferase g3p + pal → mag PLSb n.i. 2.3.1.51 1-acyl-sn-glycerol-3-phosphate mag + pal → dag PLSc MG212 acyltransferase 2.7.7.41 phosphatidate cytidyltransferase dag + ctp → cdp-dag + pp CDS MG437 cdp-dag + ser → pser + 2.7.8.8 phosphatidylserine synthase PSS n.i. cmp 4.1.1.65 phosphatidylserine decarboxylase pser → peta PSD n.i. -
Q 297 Suppl USE
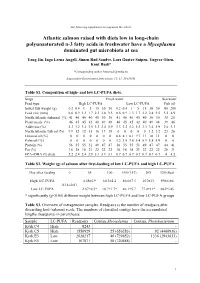
The following supplement accompanies the article Atlantic salmon raised with diets low in long-chain polyunsaturated n-3 fatty acids in freshwater have a Mycoplasma dominated gut microbiota at sea Yang Jin, Inga Leena Angell, Simen Rød Sandve, Lars Gustav Snipen, Yngvar Olsen, Knut Rudi* *Corresponding author: [email protected] Aquaculture Environment Interactions 11: 31–39 (2019) Table S1. Composition of high- and low LC-PUFA diets. Stage Fresh water Sea water Feed type High LC-PUFA Low LC-PUFA Fish oil Initial fish weight (g) 0.2 0.4 1 5 15 30 50 0.2 0.4 1 5 15 30 50 80 200 Feed size (mm) 0.6 0.9 1.3 1.7 2.2 2.8 3.5 0.6 0.9 1.3 1.7 2.2 2.8 3.5 3.5 4.9 North Atlantic fishmeal (%) 41 40 40 40 40 30 30 41 40 40 40 40 30 30 35 25 Plant meals (%) 46 45 45 42 40 49 48 46 45 45 42 40 49 48 39 46 Additives (%) 3.3 3.2 3.2 3.5 3.3 3.4 3.9 3.3 3.2 3.2 3.5 3.3 3.4 3.9 2.6 3.3 North Atlantic fish oil (%) 9.9 12 12 15 16 17 18 0 0 0 0 0 1.2 1.2 23 26 Linseed oil (%) 0 0 0 0 0 0 0 6.8 8.1 8.1 9.7 11 10 11 0 0 Palm oil (%) 0 0 0 0 0 0 0 3.2 3.8 3.8 5.4 5.9 5.8 5.9 0 0 Protein (%) 56 55 55 51 49 47 47 56 55 55 51 49 47 47 44 41 Fat (%) 16 18 18 21 22 22 22 16 18 18 21 22 22 22 28 31 EPA+DHA (% diet) 2.2 2.4 2.4 2.9 3.1 3.1 3.1 0.7 0.7 0.7 0.7 0.7 0.7 0.7 4 4.2 Table S2. -

Recognition of Class II MHC Peptide Ligands That Contain Β-Amino Acids Ross W
Recognition of Class II MHC Peptide Ligands That Contain β-Amino Acids Ross W. Cheloha, Andrew W. Woodham, Djenet Bousbaine, Tong Wang, Shi Liu, John Sidney, Alessandro Sette, Samuel This information is current as H. Gellman and Hidde L. Ploegh of October 1, 2021. J Immunol published online 7 August 2019 http://www.jimmunol.org/content/early/2019/08/06/jimmun ol.1900536 Downloaded from Supplementary http://www.jimmunol.org/content/suppl/2019/08/06/jimmunol.190053 Material 6.DCSupplemental http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average by guest on October 1, 2021 Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2019 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. Published August 7, 2019, doi:10.4049/jimmunol.1900536 The Journal of Immunology Recognition of Class II MHC Peptide Ligands That Contain b-Amino Acids Ross W. Cheloha,* Andrew W. Woodham,* Djenet Bousbaine,*,† Tong Wang,‡ Shi Liu,‡ John Sidney,x Alessandro Sette,x,{ Samuel H. -
![[Thesis Title Goes Here]](https://docslib.b-cdn.net/cover/3110/thesis-title-goes-here-2873110.webp)
[Thesis Title Goes Here]
BIOLOGICALLY ACTIVE ASSEMBLIES THAT ATTENUATE THROMBOSIS ON BLOOD-CONTACTING SURFACES A Dissertation Presented to The Academic Faculty by Zheng Qu In Partial Fulfillment of the Requirements for the Degree Doctor of Philosophy in Bioengineering Georgia Institute of Technology December 2012 BIOLOGICALLY ACTIVE ASSEMBLIES THAT ATTENUATE THROMBOSIS ON BLOOD-CONTACTING SURFACES Approved by: Dr. Elliot L. Chaikof, Advisor Dr. Larry V. McIntire The Wallace H. Coulter Department of The Wallace H. Coulter Department of Biomedical Engineering Biomedical Engineering Georgia Institute of Technology Georgia Institute of Technology Department of Surgery Beth Israel Deaconess Medical Center Dr. Julia E. Babensee Dr. W. Robert Taylor The Wallace H.Coulter Department of Division of Cardiology Biomedical Engineering Emory University School of Medicine Georgia Institute of Technology Dr. Stephen R. Hanson Department of Biomedical Engineering Oregon Health and Science University Date Approved: November 2, 2012 Dedicated to my parents, for their unconditional love. ACKNOWLEDGEMENTS The progress and advances made in this research were enabled by contributions from many colleagues and peers, as well as support from family and friends. It is these relationships that were forged during the course of my career that I cherish the most. First and foremost, I want to express my sincerest gratitude for Dr. Elliot Chaikof, for guiding me through the many challenges of scientific research with innovative ideas and patient advice, and above all, by always keeping the “big picture” in perspective. Dr. Chaikof has taken the time to personally support every step along the way of my career as a PhD candidate despite his huge commitments to the clinic. -

Structural and Mechanistic Investigations on Salmonella
Chittori et al. BMC Structural Biology 2012, 12:24 http://www.biomedcentral.com/1472-6807/12/24 RESEARCH ARTICLE Open Access Structural and mechanistic investigations on Salmonella typhimurium acetate kinase (AckA): identification of a putative ligand binding pocket at the dimeric interface Sagar Chittori1,3, Handanahal S Savithri2 and Mathur RN Murthy1* Abstract Background: Bacteria such as Escherichia coli and Salmonella typhimurium can utilize acetate as the sole source of carbon and energy. Acetate kinase (AckA) and phosphotransacetylase (Pta), key enzymes of acetate utilization pathway, regulate flux of metabolites in glycolysis, gluconeogenesis, TCA cycle, glyoxylate bypass and fatty acid metabolism. Results: Here we report kinetic characterization of S. typhimurium AckA (StAckA) and structures of its unliganded (Form-I, 2.70 Å resolution) and citrate-bound (Form-II, 1.90 Å resolution) forms. The enzyme showed broad substrate specificity with kcat/Km in the order of acetate > propionate > formate. Further, the Km for acetyl-phosphate was significantly lower than for acetate and the enzyme could catalyze the reverse reaction (i.e. ATP synthesis) more efficiently. ATP and Mg2+ could be substituted by other nucleoside 50-triphosphates (GTP, UTP and CTP) and divalent cations (Mn2+ and Co2+), respectively. Form-I StAckA represents the first structural report of an unliganded AckA. StAckA protomer consists of two domains with characteristic βββαβαβα topology of ASKHA superfamily of proteins. These domains adopt an intermediate conformation compared to that of open and closed forms of ligand-bound Methanosarcina thermophila AckA (MtAckA). Spectroscopic and structural analyses of StAckA further suggested occurrence of inter-domain motion upon ligand-binding. -

Asymmetric Disulfanylbenzamides As Irreversible and Selective Inhibitors
Full Papers ChemMedChem doi.org/10.1002/cmdc.201900687 1 2 3 Asymmetric Disulfanylbenzamides as Irreversible and 4 5 Selective Inhibitors of Staphylococcus aureus Sortase A 6 [a] [b] [b] [a] 7 Fabian Barthels, Gabriella Marincola, Tessa Marciniak, Matthias Konhäuser, [a] [c] [d, e] [a, f] [d] 8 Stefan Hammerschmidt, Jan Bierlmeier, Ute Distler, Peter R. Wich, Stefan Tenzer, 9 Dirk Schwarzer,[c] Wilma Ziebuhr,[b] and Tanja Schirmeister*[a] 10 11 12 Staphylococcus aureus is one of the most frequent causes of methods allowed the experimentally observed binding affinities 13 nosocomial and community-acquired infections, with drug- and selectivities to be rationalized. The most active compounds 14 resistant strains being responsible for tens of thousands of were found to have single-digit micromolar Ki values and 15 deaths per year. S. aureus sortase A inhibitors are designed to caused up to a 66% reduction of S. aureus fibrinogen attach- 16 interfere with virulence determinants. We have identified ment at an effective inhibitor concentration of 10 μM. This new 17 disulfanylbenzamides as a new class of potent inhibitors against molecule class exhibited minimal cytotoxicity, low bacterial 18 sortase A that act by covalent modification of the active-site growth inhibition and impaired sortase-mediated adherence of 19 cysteine. A broad series of derivatives were synthesized to S. aureus cells. 20 derive structure-activity relationships (SAR). In vitro and in silico 21 22 Introduction target structures.[2] SrtA mediates the attachment of surface 23 proteins to the bacterial cell wall and it was shown that an S. 24 The ongoing spread of antibiotic resistance among Gram- aureus ΔSrtA mutant is clearly attenuated in mouse infection 25 positive bacteria such as Staphylococcus aureus highlights the models compared to the wild type.[3,4] SrtA inhibitors are likely 26 need for new treatment options beyond traditional antibiotics. -

Sorting out Sortases: Designing a Better Enzyme and Synthesizing Trapping Peptides Towards Capturing a Bound-State Structure
Western Washington University Western CEDAR WWU Honors Program Senior Projects WWU Graduate and Undergraduate Scholarship Spring 2020 Sorting Out Sortases: Designing a Better Enzyme and Synthesizing Trapping Peptides Towards Capturing a Bound-State Structure Katherine Johnston Western Washington University Follow this and additional works at: https://cedar.wwu.edu/wwu_honors Part of the Chemistry Commons Recommended Citation Johnston, Katherine, "Sorting Out Sortases: Designing a Better Enzyme and Synthesizing Trapping Peptides Towards Capturing a Bound-State Structure" (2020). WWU Honors Program Senior Projects. 398. https://cedar.wwu.edu/wwu_honors/398 This Project is brought to you for free and open access by the WWU Graduate and Undergraduate Scholarship at Western CEDAR. It has been accepted for inclusion in WWU Honors Program Senior Projects by an authorized administrator of Western CEDAR. For more information, please contact [email protected]. Sorting Out Sortases: Designing a Better Enzyme and Synthesizing Trapping Peptides Towards Capturing a Bound-State Structure By Katherine Johnston June 2020 A Thesis Presented to The Honors Department of Western Washington University In Partial Fulfillment Of the Requirements for University Honors Abstract Sortase A is a powerful protein engineering tool that cleaves proteins and attaches them to an acyl acceptor of choice. However, the most active wild type variants of sortase A only show high activity at a limited number of cleavage motifs, and so work is underway to create a variant of sortase A that shows high activity at a greater variety of cleavage sites. Additionally, current studies attempting to optimize the enzyme require a way to stabilize an unstructured loop for the crystallization, in order to collect X-ray diffraction structural data.