DERIVED HUMAN MESENCHYMAL STEM CELLS Patricia Janicki, Stephane Boeuf, Eric Steck, Marcus Egermann, Philip Kasten§ and Wiltrud Richter*
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Association Analyses of Known Genetic Variants with Gene
ASSOCIATION ANALYSES OF KNOWN GENETIC VARIANTS WITH GENE EXPRESSION IN BRAIN by Viktoriya Strumba A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy (Bioinformatics) in The University of Michigan 2009 Doctoral Committee: Professor Margit Burmeister, Chair Professor Huda Akil Professor Brian D. Athey Assistant Professor Zhaohui S. Qin Research Statistician Thomas Blackwell To Sam and Valentina Dmitriy and Elizabeth ii ACKNOWLEDGEMENTS I would like to thank my advisor Professor Margit Burmeister, who tirelessly guided me though seemingly impassable corridors of graduate work. Throughout my thesis writing period she provided sound advice, encouragement and inspiration. Leading by example, her enthusiasm and dedication have been instrumental in my path to becoming a better scientist. I also would like to thank my co-advisor Tom Blackwell. His careful prodding always kept me on my toes and looking for answers, which taught me the depth of careful statistical analysis. His diligence and dedication have been irreplaceable in most difficult of projects. I also would like to thank my other committee members: Huda Akil, Brian Athey and Steve Qin as well as David States. You did not make it easy for me, but I thank you for believing and not giving up. Huda’s eloquence in every subject matter she explained have been particularly inspiring, while both Huda’s and Brian’s valuable advice made the completion of this dissertation possible. I would also like to thank all the members of the Burmeister lab, both past and present: Sandra Villafuerte, Kristine Ito, Cindy Schoen, Karen Majczenko, Ellen Schmidt, Randi Burns, Gang Su, Nan Xiang and Ana Progovac. -

MRPL11 Antibody A
Revision 1 C 0 2 - t MRPL11 Antibody a e r o t S Orders: 877-616-CELL (2355) [email protected] Support: 877-678-TECH (8324) 9 9 Web: [email protected] 1 www.cellsignal.com 2 # 3 Trask Lane Danvers Massachusetts 01923 USA For Research Use Only. Not For Use In Diagnostic Procedures. Applications: Reactivity: Sensitivity: MW (kDa): Source: UniProt ID: Entrez-Gene Id: WB, IP H Mk Endogenous 21 Rabbit Q9Y3B7 65003 Product Usage Information Application Dilution Western Blotting 1:1000 Immunoprecipitation 1:50 Storage Supplied in 10 mM sodium HEPES (pH 7.5), 150 mM NaCl, 100 µg/ml BSA and 50% glycerol. Store at –20°C. Do not aliquot the antibody. Specificity / Sensitivity MRPL11 Antibody detects endogenous levels of total MRPL11 protein. Species Reactivity: Human, Monkey Source / Purification Polyclonal antibodies are produced by immunizing animals with a synthetic peptide corresponding to the sequence of human MRPL11. Antibodies are purified by peptide affinity chromatography. Background A subset of mitochondrial proteins are synthesized on the ribosomes within mitochondria (1). The 55S mammalian mitochondrial ribosomes are composed of a 28S small subunit and a 39S large subunit (1). Over 40 protein components have been identified from the large subunit of the human mitochondrial ribosome (1). The mitochondrial ribosomal protein L11 (MRPL11) is one such component (1). In animals, plants and fungi, this protein is translated from a gene in the nuclear genome (2). 1. Koc, E.C. et al. (2001) J Biol Chem 276, 43958-69. 2. Handa, H. et al. (2001) Mol Genet Genomics 265, 569-75. -
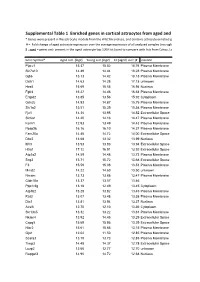
Supplemental Table 1 Enriched Genes in Cortical Astrocytes from Aged
Supplemental Table 1 Enriched genes in cortical astrocytes from aged and young-adult mice * Genes were present in the astrocyte module from the WGCNA analysis, and contains astrocyte enriched genes compared to microglia and oligodendrocytes # = Fold change of aged astrocyte expression over the average expression of all analyzed samples (microglia, astrocytes: young, old, with and without myelin contamination) $ ; aged = genes only present in the aged astrocyte top 1000 list (used to compare with lists from Cahoy, Lovatt, Doyle; see Fig. 4B), all = genes present in all astrocyte top 1000 lists Gene Symbol* Aged astr. (log2) Young astr.(log2) FC (aged/ aver.)# Location Ptprz1 15.37 15.02 18.76 Plasma Membrane Slc7a10 14.49 14.44 18.28 Plasma Membrane Gjb6 15.13 14.42 18.18 Plasma Membrane Dclk1 14.63 14.28 17.18 unknown Hes5 15.69 15.55 16.94 Nucleus Fgfr3 15.27 14.46 16.54 Plasma Membrane Entpd2 13.85 13.56 15.92 Cytoplasm Grin2c 14.93 14.87 15.75 Plasma Membrane Slc1a2 15.51 15.39 15.58 Plasma Membrane Fjx1 14.36 13.98 14.52 Extracellular Space Slc6a1 14.20 14.16 14.47 Plasma Membrane Kcnk1 12.93 13.49 14.43 Plasma Membrane Ppap2b 16.16 16.10 14.37 Plasma Membrane Fam20a 14.48 14.72 14.00 Extracellular Space Dbx2 13.68 13.32 13.99 Nucleus Itih3 13.93 13.93 13.94 Extracellular Space Htra1 17.12 16.91 13.92 Extracellular Space Atp1a2 14.59 14.48 13.73 Plasma Membrane Scg3 15.71 15.72 13.68 Extracellular Space F3 15.59 15.08 13.51 Plasma Membrane Mmd2 14.22 14.60 13.50 unknown Nrcam 13.73 13.88 13.47 Plasma Membrane Cldn10a 13.37 13.57 13.46 -

Table 2. Significant
Table 2. Significant (Q < 0.05 and |d | > 0.5) transcripts from the meta-analysis Gene Chr Mb Gene Name Affy ProbeSet cDNA_IDs d HAP/LAP d HAP/LAP d d IS Average d Ztest P values Q-value Symbol ID (study #5) 1 2 STS B2m 2 122 beta-2 microglobulin 1452428_a_at AI848245 1.75334941 4 3.2 4 3.2316485 1.07398E-09 5.69E-08 Man2b1 8 84.4 mannosidase 2, alpha B1 1416340_a_at H4049B01 3.75722111 3.87309653 2.1 1.6 2.84852656 5.32443E-07 1.58E-05 1110032A03Rik 9 50.9 RIKEN cDNA 1110032A03 gene 1417211_a_at H4035E05 4 1.66015788 4 1.7 2.82772795 2.94266E-05 0.000527 NA 9 48.5 --- 1456111_at 3.43701477 1.85785922 4 2 2.8237185 9.97969E-08 3.48E-06 Scn4b 9 45.3 Sodium channel, type IV, beta 1434008_at AI844796 3.79536664 1.63774235 3.3 2.3 2.75319499 1.48057E-08 6.21E-07 polypeptide Gadd45gip1 8 84.1 RIKEN cDNA 2310040G17 gene 1417619_at 4 3.38875643 1.4 2 2.69163229 8.84279E-06 0.0001904 BC056474 15 12.1 Mus musculus cDNA clone 1424117_at H3030A06 3.95752801 2.42838452 1.9 2.2 2.62132809 1.3344E-08 5.66E-07 MGC:67360 IMAGE:6823629, complete cds NA 4 153 guanine nucleotide binding protein, 1454696_at -3.46081884 -4 -1.3 -1.6 -2.6026947 8.58458E-05 0.0012617 beta 1 Gnb1 4 153 guanine nucleotide binding protein, 1417432_a_at H3094D02 -3.13334396 -4 -1.6 -1.7 -2.5946297 1.04542E-05 0.0002202 beta 1 Gadd45gip1 8 84.1 RAD23a homolog (S. -

NCAPD3 Antibody (C-Term) Affinity Purified Rabbit Polyclonal Antibody (Pab) Catalog # AP16786B
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 NCAPD3 Antibody (C-term) Affinity Purified Rabbit Polyclonal Antibody (Pab) Catalog # AP16786B Specification NCAPD3 Antibody (C-term) - Product Information Application WB,E Primary Accession P42695 Other Accession NP_056076.1 Reactivity Human Host Rabbit Clonality Polyclonal Isotype Rabbit Ig Calculated MW 168891 Antigen Region 1050-1078 NCAPD3 Antibody (C-term) - Additional Information NCAPD3 Antibody (C-term) (Cat. Gene ID 23310 #AP16786b) western blot analysis in K562 cell line lysates (35ug/lane).This Other Names Condensin-2 complex subunit D3, Non-SMC demonstrates the NCAPD3 antibody detected condensin II complex subunit D3, hCAP-D3, the NCAPD3 protein (arrow). NCAPD3, CAPD3, KIAA0056 Target/Specificity NCAPD3 Antibody (C-term) - Background This NCAPD3 antibody is generated from rabbits immunized with a KLH conjugated Condensin complexes I and II play essential synthetic peptide between 1050-1078 roles in amino acids from the C-terminal region of mitotic chromosome assembly and human NCAPD3. segregation. Both condensins contain 2 invariant structural maintenance of Dilution chromosome (SMC) WB~~1:1000 subunits, SMC2 (MIM 605576) and SMC4 (MIM 605575), but they contain Format different sets of non-SMC subunits. NCAPD3 is Purified polyclonal antibody supplied in PBS 1 of 3 non-SMC with 0.09% (W/V) sodium azide. This subunits that define condensin II (Ono et al., antibody is purified through a protein A 2003 [PubMed column, followed by peptide affinity 14532007]). purification. NCAPD3 Antibody (C-term) - References Storage Maintain refrigerated at 2-8°C for up to 2 Rose, J.E., et al. -

A Commercial Antibody to the Human Condensin II Subunit NCAPH2 Cross-Reacts with a SWI/SNF Complex Component
bioRxiv preprint doi: https://doi.org/10.1101/2020.11.07.372599; this version posted November 9, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. A commercial antibody to the human condensin II subunit NCAPH2 cross-reacts with a SWI/SNF complex component Erin E. Cutts1*, Gillian C Taylor2*, Mercedes Pardo1, Lu Yu1, Jimi C Wills3, Jyoti S. Choudhary1, Alessandro Vannini1#, Andrew J Wood2# 1 Division of Structural Biology, The Institute of Cancer Research, London SW7 3RP, United Kingdom 2 MRC Human Genetics Unit, Institute of Genetics and Molecular Medicine, The University of Edinburgh, Edinburgh, EH4 2XU, UK. 3 Cancer Research UK Edinburgh Centre, Institute of Genetics and Molecular Medicine, University of Edinburgh, Edinburgh, EH4 2XU, UK. * Equal contribution # correspondence to: [email protected], [email protected]. Summary Condensin complexes compact and disentangle chromosomes in preparation for cell division. Commercially available antibodies raised against condensin subunits have been widely used to characterise their cellular interactome. Here we have assessed the specificity of a polyclonal antibody (Bethyl A302- 276A) that is commonly used as a probe for NCAPH2, the kleisin subunit of condensin II, in mammalian cells. We find that, in addition to its intended target, this antibody cross-reacts with one or more components of the SWI/SNF family of chromatin remodelling complexes in an NCAPH2- independent manner. This cross-reactivity with an abundant chromatin- associated factor is likely to affect the interpretation of protein and chromatin immunoprecipitation experiments that make use of this antibody probe. -

Condensation of Prometaphase Chromo-Somes
Condensation of Prometaphase Chromo- somes D'Eustachio, P., Matthews, L., Orlic-Milacic, M. European Bioinformatics Institute, New York University Langone Medical Center, Ontario Institute for Cancer Research, Oregon Health and Science University. The contents of this document may be freely copied and distributed in any media, provided the authors, plus the institutions, are credited, as stated under the terms of Creative Commons Attribution 4.0 Inter- national (CC BY 4.0) License. For more information see our license. 09/03/2019 Introduction Reactome is open-source, open access, manually curated and peer-reviewed pathway database. Pathway annotations are authored by expert biologists, in collaboration with Reactome editorial staff and cross- referenced to many bioinformatics databases. A system of evidence tracking ensures that all assertions are backed up by the primary literature. Reactome is used by clinicians, geneticists, genomics research- ers, and molecular biologists to interpret the results of high-throughput experimental studies, by bioin- formaticians seeking to develop novel algorithms for mining knowledge from genomic studies, and by systems biologists building predictive models of normal and disease variant pathways. The development of Reactome is supported by grants from the US National Institutes of Health (P41 HG003751), University of Toronto (CFREF Medicine by Design), European Union (EU STRP, EMI-CD), and the European Molecular Biology Laboratory (EBI Industry program). Literature references Fabregat, A., Sidiropoulos, K., Viteri, G., Forner, O., Marin-Garcia, P., Arnau, V. et al. (2017). Reactome pathway ana- lysis: a high-performance in-memory approach. BMC bioinformatics, 18, 142. ↗ Sidiropoulos, K., Viteri, G., Sevilla, C., Jupe, S., Webber, M., Orlic-Milacic, M. -

Supplementary Figures 1-14 and Supplementary References
SUPPORTING INFORMATION Spatial Cross-Talk Between Oxidative Stress and DNA Replication in Human Fibroblasts Marko Radulovic,1,2 Noor O Baqader,1 Kai Stoeber,3† and Jasminka Godovac-Zimmermann1* 1Division of Medicine, University College London, Center for Nephrology, Royal Free Campus, Rowland Hill Street, London, NW3 2PF, UK. 2Insitute of Oncology and Radiology, Pasterova 14, 11000 Belgrade, Serbia 3Research Department of Pathology and UCL Cancer Institute, Rockefeller Building, University College London, University Street, London WC1E 6JJ, UK †Present Address: Shionogi Europe, 33 Kingsway, Holborn, London WC2B 6UF, UK TABLE OF CONTENTS 1. Supplementary Figures 1-14 and Supplementary References. Figure S-1. Network and joint spatial razor plot for 18 enzymes of glycolysis and the pentose phosphate shunt. Figure S-2. Correlation of SILAC ratios between OXS and OAC for proteins assigned to the SAME class. Figure S-3. Overlap matrix (r = 1) for groups of CORUM complexes containing 19 proteins of the 49-set. Figure S-4. Joint spatial razor plots for the Nop56p complex and FIB-associated complex involved in ribosome biogenesis. Figure S-5. Analysis of the response of emerin nuclear envelope complexes to OXS and OAC. Figure S-6. Joint spatial razor plots for the CCT protein folding complex, ATP synthase and V-Type ATPase. Figure S-7. Joint spatial razor plots showing changes in subcellular abundance and compartmental distribution for proteins annotated by GO to nucleocytoplasmic transport (GO:0006913). Figure S-8. Joint spatial razor plots showing changes in subcellular abundance and compartmental distribution for proteins annotated to endocytosis (GO:0006897). Figure S-9. Joint spatial razor plots for 401-set proteins annotated by GO to small GTPase mediated signal transduction (GO:0007264) and/or GTPase activity (GO:0003924). -

Supplementary Materials
Supplementary materials Supplementary Table S1: MGNC compound library Ingredien Molecule Caco- Mol ID MW AlogP OB (%) BBB DL FASA- HL t Name Name 2 shengdi MOL012254 campesterol 400.8 7.63 37.58 1.34 0.98 0.7 0.21 20.2 shengdi MOL000519 coniferin 314.4 3.16 31.11 0.42 -0.2 0.3 0.27 74.6 beta- shengdi MOL000359 414.8 8.08 36.91 1.32 0.99 0.8 0.23 20.2 sitosterol pachymic shengdi MOL000289 528.9 6.54 33.63 0.1 -0.6 0.8 0 9.27 acid Poricoic acid shengdi MOL000291 484.7 5.64 30.52 -0.08 -0.9 0.8 0 8.67 B Chrysanthem shengdi MOL004492 585 8.24 38.72 0.51 -1 0.6 0.3 17.5 axanthin 20- shengdi MOL011455 Hexadecano 418.6 1.91 32.7 -0.24 -0.4 0.7 0.29 104 ylingenol huanglian MOL001454 berberine 336.4 3.45 36.86 1.24 0.57 0.8 0.19 6.57 huanglian MOL013352 Obacunone 454.6 2.68 43.29 0.01 -0.4 0.8 0.31 -13 huanglian MOL002894 berberrubine 322.4 3.2 35.74 1.07 0.17 0.7 0.24 6.46 huanglian MOL002897 epiberberine 336.4 3.45 43.09 1.17 0.4 0.8 0.19 6.1 huanglian MOL002903 (R)-Canadine 339.4 3.4 55.37 1.04 0.57 0.8 0.2 6.41 huanglian MOL002904 Berlambine 351.4 2.49 36.68 0.97 0.17 0.8 0.28 7.33 Corchorosid huanglian MOL002907 404.6 1.34 105 -0.91 -1.3 0.8 0.29 6.68 e A_qt Magnogrand huanglian MOL000622 266.4 1.18 63.71 0.02 -0.2 0.2 0.3 3.17 iolide huanglian MOL000762 Palmidin A 510.5 4.52 35.36 -0.38 -1.5 0.7 0.39 33.2 huanglian MOL000785 palmatine 352.4 3.65 64.6 1.33 0.37 0.7 0.13 2.25 huanglian MOL000098 quercetin 302.3 1.5 46.43 0.05 -0.8 0.3 0.38 14.4 huanglian MOL001458 coptisine 320.3 3.25 30.67 1.21 0.32 0.9 0.26 9.33 huanglian MOL002668 Worenine -

Literature Mining Sustains and Enhances Knowledge Discovery from Omic Studies
LITERATURE MINING SUSTAINS AND ENHANCES KNOWLEDGE DISCOVERY FROM OMIC STUDIES by Rick Matthew Jordan B.S. Biology, University of Pittsburgh, 1996 M.S. Molecular Biology/Biotechnology, East Carolina University, 2001 M.S. Biomedical Informatics, University of Pittsburgh, 2005 Submitted to the Graduate Faculty of School of Medicine in partial fulfillment of the requirements for the degree of Doctor of Philosophy University of Pittsburgh 2016 UNIVERSITY OF PITTSBURGH SCHOOL OF MEDICINE This dissertation was presented by Rick Matthew Jordan It was defended on December 2, 2015 and approved by Shyam Visweswaran, M.D., Ph.D., Associate Professor Rebecca Jacobson, M.D., M.S., Professor Songjian Lu, Ph.D., Assistant Professor Dissertation Advisor: Vanathi Gopalakrishnan, Ph.D., Associate Professor ii Copyright © by Rick Matthew Jordan 2016 iii LITERATURE MINING SUSTAINS AND ENHANCES KNOWLEDGE DISCOVERY FROM OMIC STUDIES Rick Matthew Jordan, M.S. University of Pittsburgh, 2016 Genomic, proteomic and other experimentally generated data from studies of biological systems aiming to discover disease biomarkers are currently analyzed without sufficient supporting evidence from the literature due to complexities associated with automated processing. Extracting prior knowledge about markers associated with biological sample types and disease states from the literature is tedious, and little research has been performed to understand how to use this knowledge to inform the generation of classification models from ‘omic’ data. Using pathway analysis methods to better understand the underlying biology of complex diseases such as breast and lung cancers is state-of-the-art. However, the problem of how to combine literature- mining evidence with pathway analysis evidence is an open problem in biomedical informatics research. -

Interactome Analyses Revealed That the U1 Snrnp Machinery Overlaps Extensively with the RNAP II Machinery and Contains Multiple
www.nature.com/scientificreports OPEN Interactome analyses revealed that the U1 snRNP machinery overlaps extensively with the RNAP II Received: 12 April 2018 Accepted: 24 May 2018 machinery and contains multiple Published: xx xx xxxx ALS/SMA-causative proteins Binkai Chi1, Jeremy D. O’Connell1,2, Tomohiro Yamazaki1, Jaya Gangopadhyay1, Steven P. Gygi1 & Robin Reed1 Mutations in multiple RNA/DNA binding proteins cause Amyotrophic Lateral Sclerosis (ALS). Included among these are the three members of the FET family (FUS, EWSR1 and TAF15) and the structurally similar MATR3. Here, we characterized the interactomes of these four proteins, revealing that they largely have unique interactors, but share in common an association with U1 snRNP. The latter observation led us to analyze the interactome of the U1 snRNP machinery. Surprisingly, this analysis revealed the interactome contains ~220 components, and of these, >200 are shared with the RNA polymerase II (RNAP II) machinery. Among the shared components are multiple ALS and Spinal muscular Atrophy (SMA)-causative proteins and numerous discrete complexes, including the SMN complex, transcription factor complexes, and RNA processing complexes. Together, our data indicate that the RNAP II/U1 snRNP machinery functions in a wide variety of molecular pathways, and these pathways are candidates for playing roles in ALS/SMA pathogenesis. Te neurodegenerative disease Amyotrophic Lateral Sclerosis (ALS) has no known treatment, and elucidation of disease mechanisms is urgently needed. Tis problem has been especially daunting, as mutations in greater than 30 genes are ALS-causative, and these genes function in numerous cellular pathways1. Tese include mitophagy, autophagy, cytoskeletal dynamics, vesicle transport, DNA damage repair, RNA dysfunction, apoptosis, and pro- tein aggregation2–6. -

An Investigation of 53BP1 in Multiple Myeloma
University of Calgary PRISM: University of Calgary's Digital Repository Graduate Studies The Vault: Electronic Theses and Dissertations 2017 An Investigation of 53BP1 in Multiple Myeloma Simms, Justin Simms, J. (2017). An Investigation of 53BP1 in Multiple Myeloma (Unpublished master's thesis). University of Calgary, Calgary, AB. doi:10.11575/PRISM/24911 http://hdl.handle.net/11023/4002 master thesis University of Calgary graduate students retain copyright ownership and moral rights for their thesis. You may use this material in any way that is permitted by the Copyright Act or through licensing that has been assigned to the document. For uses that are not allowable under copyright legislation or licensing, you are required to seek permission. Downloaded from PRISM: https://prism.ucalgary.ca UNIVERSITY OF CALGARY An Investigation of 53BP1 in Multiple Myeloma by Justin Andrew Simms A THESIS SUBMITTED TO THE FACULTY OF GRADUATE STUDIES IN PARTIAL FULFILMENT OF THE REQUIREMENTS FOR THE DEGREE OF MASTER OF SCIENCE GRADUATE PROGRAM IN MEDICAL SCIENCE CALGARY, ALBERTA JULY, 2017 © Justin Andrew Simms 2017 Abstract Currently, multiple myeloma is the second most common hematological malignancy, and as of yet it remains incurable. Although many therapeutic advances have been made in the recent past, there is still room for improvement in the treatment of myeloma. Here, using CRISPR/Cas9 genome editing, we show that a therapeutic combination of proteasome inhibition in combination with PARP inhibition that is in clinical trials depends on the DNA damage response protein 53BP1. These findings contribute to the understanding of the mechanisms behind potential resistance to the combination of proteasome inhibitors with PAPR inhibitors.