Metagenomic Analysis of Rumen Microbial Population in Dairy Heifers Fed a High Grain Diet Supplemented with Dicarboxylic Acids O
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Bacterial Communities of the Upper Respiratory Tract of Turkeys
www.nature.com/scientificreports OPEN Bacterial communities of the upper respiratory tract of turkeys Olimpia Kursa1*, Grzegorz Tomczyk1, Anna Sawicka‑Durkalec1, Aleksandra Giza2 & Magdalena Słomiany‑Szwarc2 The respiratory tracts of turkeys play important roles in the overall health and performance of the birds. Understanding the bacterial communities present in the respiratory tracts of turkeys can be helpful to better understand the interactions between commensal or symbiotic microorganisms and other pathogenic bacteria or viral infections. The aim of this study was the characterization of the bacterial communities of upper respiratory tracks in commercial turkeys using NGS sequencing by the amplifcation of 16S rRNA gene with primers designed for hypervariable regions V3 and V4 (MiSeq, Illumina). From 10 phyla identifed in upper respiratory tract in turkeys, the most dominated phyla were Firmicutes and Proteobacteria. Diferences in composition of bacterial diversity were found at the family and genus level. At the genus level, the turkey sequences present in respiratory tract represent 144 established bacteria. Several respiratory pathogens that contribute to the development of infections in the respiratory system of birds were identifed, including the presence of Ornithobacterium and Mycoplasma OTUs. These results obtained in this study supply information about bacterial composition and diversity of the turkey upper respiratory tract. Knowledge about bacteria present in the respiratory tract and the roles they can play in infections can be useful in controlling, diagnosing and treating commercial turkey focks. Next-generation sequencing has resulted in a marked increase in culture-independent studies characterizing the microbiome of humans and animals1–6. Much of these works have been focused on the gut microbiome of humans and other production animals 7–11. -

Global Metagenomic Survey Reveals a New Bacterial Candidate Phylum in Geothermal Springs
ARTICLE Received 13 Aug 2015 | Accepted 7 Dec 2015 | Published 27 Jan 2016 DOI: 10.1038/ncomms10476 OPEN Global metagenomic survey reveals a new bacterial candidate phylum in geothermal springs Emiley A. Eloe-Fadrosh1, David Paez-Espino1, Jessica Jarett1, Peter F. Dunfield2, Brian P. Hedlund3, Anne E. Dekas4, Stephen E. Grasby5, Allyson L. Brady6, Hailiang Dong7, Brandon R. Briggs8, Wen-Jun Li9, Danielle Goudeau1, Rex Malmstrom1, Amrita Pati1, Jennifer Pett-Ridge4, Edward M. Rubin1,10, Tanja Woyke1, Nikos C. Kyrpides1 & Natalia N. Ivanova1 Analysis of the increasing wealth of metagenomic data collected from diverse environments can lead to the discovery of novel branches on the tree of life. Here we analyse 5.2 Tb of metagenomic data collected globally to discover a novel bacterial phylum (‘Candidatus Kryptonia’) found exclusively in high-temperature pH-neutral geothermal springs. This lineage had remained hidden as a taxonomic ‘blind spot’ because of mismatches in the primers commonly used for ribosomal gene surveys. Genome reconstruction from metagenomic data combined with single-cell genomics results in several high-quality genomes representing four genera from the new phylum. Metabolic reconstruction indicates a heterotrophic lifestyle with conspicuous nutritional deficiencies, suggesting the need for metabolic complementarity with other microbes. Co-occurrence patterns identifies a number of putative partners, including an uncultured Armatimonadetes lineage. The discovery of Kryptonia within previously studied geothermal springs underscores the importance of globally sampled metagenomic data in detection of microbial novelty, and highlights the extraordinary diversity of microbial life still awaiting discovery. 1 Department of Energy Joint Genome Institute, Walnut Creek, California 94598, USA. 2 Department of Biological Sciences, University of Calgary, Calgary, Alberta T2N 1N4, Canada. -

The Mysterious Orphans of Mycoplasmataceae
The mysterious orphans of Mycoplasmataceae Tatiana V. Tatarinova1,2*, Inna Lysnyansky3, Yuri V. Nikolsky4,5,6, and Alexander Bolshoy7* 1 Children’s Hospital Los Angeles, Keck School of Medicine, University of Southern California, Los Angeles, 90027, California, USA 2 Spatial Science Institute, University of Southern California, Los Angeles, 90089, California, USA 3 Mycoplasma Unit, Division of Avian and Aquatic Diseases, Kimron Veterinary Institute, POB 12, Beit Dagan, 50250, Israel 4 School of Systems Biology, George Mason University, 10900 University Blvd, MSN 5B3, Manassas, VA 20110, USA 5 Biomedical Cluster, Skolkovo Foundation, 4 Lugovaya str., Skolkovo Innovation Centre, Mozhajskij region, Moscow, 143026, Russian Federation 6 Vavilov Institute of General Genetics, Moscow, Russian Federation 7 Department of Evolutionary and Environmental Biology and Institute of Evolution, University of Haifa, Israel 1,2 [email protected] 3 [email protected] 4-6 [email protected] 7 [email protected] 1 Abstract Background: The length of a protein sequence is largely determined by its function, i.e. each functional group is associated with an optimal size. However, comparative genomics revealed that proteins’ length may be affected by additional factors. In 2002 it was shown that in bacterium Escherichia coli and the archaeon Archaeoglobus fulgidus, protein sequences with no homologs are, on average, shorter than those with homologs [1]. Most experts now agree that the length distributions are distinctly different between protein sequences with and without homologs in bacterial and archaeal genomes. In this study, we examine this postulate by a comprehensive analysis of all annotated prokaryotic genomes and focusing on certain exceptions. -

MIB–MIP Is a Mycoplasma System That Captures and Cleaves Immunoglobulin G
MIB–MIP is a mycoplasma system that captures and cleaves immunoglobulin G Yonathan Arfia,b,1, Laetitia Minderc,d, Carmelo Di Primoe,f,g, Aline Le Royh,i,j, Christine Ebelh,i,j, Laurent Coquetk, Stephane Claveroll, Sanjay Vasheem, Joerg Joresn,o, Alain Blancharda,b, and Pascal Sirand-Pugneta,b aINRA (Institut National de la Recherche Agronomique), UMR 1332 Biologie du Fruit et Pathologie, F-33882 Villenave d’Ornon, France; bUniversity of Bordeaux, UMR 1332 Biologie du Fruit et Pathologie, F-33882 Villenave d’Ornon, France; cInstitut Européen de Chimie et Biologie, UMS 3033, University of Bordeaux, 33607 Pessac, France; dInstitut Bergonié, SIRIC BRIO, 33076 Bordeaux, France; eINSERM U1212, ARN Regulation Naturelle et Artificielle, 33607 Pessac, France; fCNRS UMR 5320, ARN Regulation Naturelle et Artificielle, 33607 Pessac, France; gInstitut Européen de Chimie et Biologie, University of Bordeaux, 33607 Pessac, France; hInstitut de Biologie Structurale, University of Grenoble Alpes, F-38044 Grenoble, France; iCNRS, Institut de Biologie Structurale, F-38044 Grenoble, France; jCEA, Institut de Biologie Structurale, F-38044 Grenoble, France; kCNRS UMR 6270, Plateforme PISSARO, Institute for Research and Innovation in Biomedicine - Normandie Rouen, Normandie Université, F-76821 Mont-Saint-Aignan, France; lProteome Platform, Functional Genomic Center of Bordeaux, University of Bordeaux, F-33076 Bordeaux Cedex, France; mJ. Craig Venter Institute, Rockville, MD 20850; nInternational Livestock Research Institute, 00100 Nairobi, Kenya; and oInstitute of Veterinary Bacteriology, University of Bern, CH-3001 Bern, Switzerland Edited by Roy Curtiss III, University of Florida, Gainesville, FL, and approved March 30, 2016 (received for review January 12, 2016) Mycoplasmas are “minimal” bacteria able to infect humans, wildlife, introduced into naive herds (8). -

Supplementary Information for Microbial Electrochemical Systems Outperform Fixed-Bed Biofilters for Cleaning-Up Urban Wastewater
Electronic Supplementary Material (ESI) for Environmental Science: Water Research & Technology. This journal is © The Royal Society of Chemistry 2016 Supplementary information for Microbial Electrochemical Systems outperform fixed-bed biofilters for cleaning-up urban wastewater AUTHORS: Arantxa Aguirre-Sierraa, Tristano Bacchetti De Gregorisb, Antonio Berná, Juan José Salasc, Carlos Aragónc, Abraham Esteve-Núñezab* Fig.1S Total nitrogen (A), ammonia (B) and nitrate (C) influent and effluent average values of the coke and the gravel biofilters. Error bars represent 95% confidence interval. Fig. 2S Influent and effluent COD (A) and BOD5 (B) average values of the hybrid biofilter and the hybrid polarized biofilter. Error bars represent 95% confidence interval. Fig. 3S Redox potential measured in the coke and the gravel biofilters Fig. 4S Rarefaction curves calculated for each sample based on the OTU computations. Fig. 5S Correspondence analysis biplot of classes’ distribution from pyrosequencing analysis. Fig. 6S. Relative abundance of classes of the category ‘other’ at class level. Table 1S Influent pre-treated wastewater and effluents characteristics. Averages ± SD HRT (d) 4.0 3.4 1.7 0.8 0.5 Influent COD (mg L-1) 246 ± 114 330 ± 107 457 ± 92 318 ± 143 393 ± 101 -1 BOD5 (mg L ) 136 ± 86 235 ± 36 268 ± 81 176 ± 127 213 ± 112 TN (mg L-1) 45.0 ± 17.4 60.6 ± 7.5 57.7 ± 3.9 43.7 ± 16.5 54.8 ± 10.1 -1 NH4-N (mg L ) 32.7 ± 18.7 51.6 ± 6.5 49.0 ± 2.3 36.6 ± 15.9 47.0 ± 8.8 -1 NO3-N (mg L ) 2.3 ± 3.6 1.0 ± 1.6 0.8 ± 0.6 1.5 ± 2.0 0.9 ± 0.6 TP (mg -

1587714682 277 2.Pdf
Systematic and Applied Microbiology 42 (2019) 107–116 Contents lists available at ScienceDirect Systematic and Applied Microbiology jou rnal homepage: http://www.elsevier.com/locate/syapm The diverse and extensive plant polysaccharide degradative apparatuses of the rumen and hindgut Prevotella species: A factor in their ubiquity? ∗ Tomazˇ Accetto , Gorazd Avgustinˇ University of Ljubljana, Biotechnical faculty, Animal Science Department, Groblje 3, 1230 Domzale,ˇ Slovenia a r t i c l e i n f o a b s t r a c t Article history: Although the Prevotella are commonly observed in high shares in the mammalian hindgut and rumen Received 2 August 2018 studies using NGS approach, the knowledge on their actual role, though postulated to lie in soluble fibre Received in revised form 2 October 2018 degradation, is scarce. Here we analyse in total 23, more than threefold of hitherto known rumen and Accepted 3 October 2018 hindgut Prevotella species and show that rumen/hindgut Prevotella generally possess extensive reper- toires of polysaccharide utilization loci (PULs) and carbohydrate active enzymes targeting various plant Keywords: polysaccharides. These PUL repertoires separate analysed Prevotella into generalists and specialists yet a Prevotella finer diversity among generalists is evident too, both in range of substrates targeted and in PUL combi- Rumen Hindgut nations targeting the same broad substrate classes. Upon evaluation of the shares of species analysed in this study in rumen metagenomes we found firstly, that they contributed significantly to total Prevotella Polysaccharide utilization locus CAZYme abundance though much of rumen Prevotella diversity may still be unknown. Secondly, the hindgut Pre- Metagenome votella species originally isolated in pigs and humans occasionally dominated among the Prevotella with surprisingly high metagenome read shares and were consistently found in rumen metagenome samples from sites as apart as New Zealand and Scotland. -

Correlation Between the Oral Microbiome and Brain Resting State Connectivity in Smokers
bioRxiv preprint doi: https://doi.org/10.1101/444612; this version posted October 16, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Correlation between the oral microbiome and brain resting state connectivity in smokers Dongdong Lin1, Kent Hutchison2, Salvador Portillo3, Victor Vegara1, Jarod Ellingson2, Jingyu Liu1,3, Amanda Carroll-Portillo3,* ,Vince D. Calhoun1,3,* 1The Mind Research Network, Albuquerque, New Mexico, 87106 2University of Colorado Boulder, Boulder, CO 3University of New Mexico, Department of Electrical and Computer Engineering, Albuquerque, New Mexico, 87106 * authors contributed equally to the work. Abstract Recent studies have shown a critical role for the gastrointestinal microbiome in brain and behavior via a complex gut–microbiome–brain axis, however, the influence of the oral microbiome in neurological processes is much less studied, especially in response to the stimuli in the oral microenvironment such as smoking. Additionally, given the complex structural and functional networks in brain system, our knowledge about the relationship between microbiome and brain functions on specific brain circuits is still very limited. In this pilot work, we leverage next generation microbial sequencing with functional MRI techniques to enable the delineation of microbiome-brain network links as well as their relations to cigarette smoking. Thirty smokers and 30 age- and sex- matched non-smokers were recruited for measuring both microbial community and brain functional networks. Statistical analyses were performed to demonstrate the influence of smoking on: the taxonomy and abundance of the constituents within the oral microbial community, brain functional network connectivity, and associations between microbial shifts and the brain signaling network. -

Characterization of Antibiotic Resistance Genes in the Species of the Rumen Microbiota
ARTICLE https://doi.org/10.1038/s41467-019-13118-0 OPEN Characterization of antibiotic resistance genes in the species of the rumen microbiota Yasmin Neves Vieira Sabino1, Mateus Ferreira Santana1, Linda Boniface Oyama2, Fernanda Godoy Santos2, Ana Júlia Silva Moreira1, Sharon Ann Huws2* & Hilário Cuquetto Mantovani 1* Infections caused by multidrug resistant bacteria represent a therapeutic challenge both in clinical settings and in livestock production, but the prevalence of antibiotic resistance genes 1234567890():,; among the species of bacteria that colonize the gastrointestinal tract of ruminants is not well characterized. Here, we investigate the resistome of 435 ruminal microbial genomes in silico and confirm representative phenotypes in vitro. We find a high abundance of genes encoding tetracycline resistance and evidence that the tet(W) gene is under positive selective pres- sure. Our findings reveal that tet(W) is located in a novel integrative and conjugative element in several ruminal bacterial genomes. Analyses of rumen microbial metatranscriptomes confirm the expression of the most abundant antibiotic resistance genes. Our data provide insight into antibiotic resistange gene profiles of the main species of ruminal bacteria and reveal the potential role of mobile genetic elements in shaping the resistome of the rumen microbiome, with implications for human and animal health. 1 Departamento de Microbiologia, Universidade Federal de Viçosa, Viçosa, Minas Gerais, Brazil. 2 Institute for Global Food Security, School of Biological -

Data Sheet on Spiroplasma Citri
Prepared by CABI and EPPO for the EU under Contract 90/399003 Data Sheets on Quarantine Pests Spiroplasma citri The vectors of Spiroplasma citri are individually included in EU Directive 77/93. Since their importance only arises in relation to S. citri, they are covered in this data sheet. IDENTITY • Spiroplasma citri Name: Spiroplasma citri Saglio et al. Taxonomic position: Bacteria: Tenericutes: Mollicutes Common names: Stubborn, little leaf (English) Stubborn (French) Bayer computer code: SPIRCI EU Annex designation: II/A2 • Circulifer tenellus Name: Circulifer tenellus (Baker) Synonyms: Neoaliturus tenellus (Baker) Eutettix tenellus Baker Taxonomic position: Insecta: Hemiptera: Homoptera: Cicadellidae Common names: Beet leafhopper (English) Cicadelle de la betterave (French) Saltahojas de la remolacha (Spanish) Notes on taxonomy and nomenclature: Oman (1970) argues for the retention of Circulifer and Neoaliturus as separate genera; in his concept both tenellus and haematoceps are placed in Circulifer. Nast (1972) treats the two genera as Neoaliturus notwithstanding Oman's (1970) arguments for the retention of the two genera. He includes 17 species in the Palaearctic region of which 14 are recorded from the Mediterranean Basin. The two species, tenellus and haematoceps, are both included in Circulifer by della Giustina (1989), who reports that tenellus is confirmed in Corsica. Bayer computer code: CIRCTE EU Annex designation: II/A2 • Neoaliturus haematoceps Name: Neoaliturus haematoceps (Mulsant & Rey) Synonyms: Circulifer haematoceps (Mulsant & Rey) Jassus haematoceps Mulsant & Rey Taxonomic position: Insecta: Hemiptera: Homoptera: Cicadellidae Bayer computer code: NEOAHA EU Annex designation: II/A2 (under the name Circulifer haematoceps) HOSTS • Spiroplasma citri The principal economic hosts of S. citri are susceptible Citrus spp. -

Downloaded from Genome Website
bioRxiv preprint doi: https://doi.org/10.1101/2020.11.18.388454; this version posted November 19, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Characterization of the first cultured free-living representative of 2 Candidatus Izimaplasma uncovers its unique biology 3 Rikuan Zheng1,2,3,4, Rui Liu1,2,4, Yeqi Shan1,2,3,4, Ruining Cai1,2,3,4, Ge Liu1,2,4, Chaomin Sun1,2,4* 1 4 CAS Key Laboratory of Experimental Marine Biology & Center of Deep Sea 5 Research, Institute of Oceanology, Chinese Academy of Sciences, Qingdao, China 2 6 Laboratory for Marine Biology and Biotechnology, Qingdao National Laboratory 7 for Marine Science and Technology, Qingdao, China 3 8 College of Earth Science, University of Chinese Academy of Sciences, Beijing, 9 China 10 4Center of Ocean Mega-Science, Chinese Academy of Sciences, Qingdao, China 11 12 * Corresponding author 13 Chaomin Sun Tel.: +86 532 82898857; fax: +86 532 82898857. 14 E-mail address: [email protected] 15 16 17 Key words: Candidatus Izimaplasma, uncultivation, biogeochemical cycling, 18 extracellular DNA, in situ, deep sea 19 Running title: Characterization of the first cultured Izimaplasma 20 21 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.11.18.388454; this version posted November 19, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 22 Abstract 23 Candidatus Izimaplasma, an intermediate in the reductive evolution from Firmicutes 24 to Mollicutes, was proposed to represent a novel class of free-living wall-less bacteria 25 within the phylum Tenericutes found in deep-sea methane seeps. -

Redalyc.Bacterial Diversity in Bovine Rumen by Metagenomic 16S Rdna
Acta Scientiarum. Animal Sciences ISSN: 1806-2636 [email protected] Universidade Estadual de Maringá Brasil Barbetta de Jesus, Raphael; Pine Omori, Wellington; de Macedo Lemos, Eliana Gertrudes; Marcondes de Souza, Jackson Antônio Bacterial diversity in bovine rumen by metagenomic 16S rDNA sequencing and scanning electron microscopy Acta Scientiarum. Animal Sciences, vol. 37, núm. 3, julio-septiembre, 2015, pp. 251-257 Universidade Estadual de Maringá Maringá, Brasil Available in: http://www.redalyc.org/articulo.oa?id=303141017006 How to cite Complete issue Scientific Information System More information about this article Network of Scientific Journals from Latin America, the Caribbean, Spain and Portugal Journal's homepage in redalyc.org Non-profit academic project, developed under the open access initiative Acta Scientiarum http://www.uem.br/acta ISSN printed: 1806-2636 ISSN on-line: 1807-8672 Doi: 10.4025/actascianimsci.v37i3.26535 Bacterial diversity in bovine rumen by metagenomic 16S rDNA sequencing and scanning electron microscopy Raphael Barbetta de Jesus1,2, Wellington Pine Omori1,2, Eliana Gertrudes de Macedo Lemos1,3 and Jackson Antônio Marcondes de Souza1,2* 1Faculdade de Ciências Agrárias e Veterinárias, Universidade Estadual Paulista, Via de Acesso Professor Paulo Donato Castellane, s/n, 14884- 900, Jaboticabal, São Paulo, Brazil. 2Departamento de Biologia Aplicada à Agropecuária, Faculdade de Ciências Agrárias e Veterinárias, Jaboticabal, São Paulo, Brazil. 3Departamento de Tecnologia, Faculdade de Ciências Agrárias e Veterinárias, Jaboticabal, São Paulo, Brazil. *Author for correspondence. E-mail: [email protected] ABSTRACT. The bacterial diversity by 16S rDNA partial sequencing and scanning electron microscope (SEM) of the rumen microbiome was characterized. Three Nellore bovines, cannulated at the rumen, were utilized. -
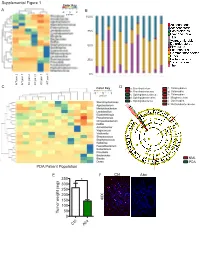
Mental Figure 1 Color Key a -2 0 2 B Z-Score 100%
Supplemental Figure 1 Color Key A -2 0 2 B z-score 100% 75% 50% 25% 0% KC pan 1 WT pan 3 WT KC pan 3 WT pan 2 WT pan 1 WT KC pan 2 C Color Key D a: Brevibacterium f: Chlamydiales b: Brevibacteriaceae g: Chlamydiia -3 0 3 z-score c: Sphingobacteriaceae h: Chlamydiae d: Sphingobacteriales i: Mogibacterium e: Sphingobacteriia j: Oscillospira k: Methylobacteriaceae NML Control Microb.-entrained MΦ PDA PDA Patient Population Control Microb.-entrained MΦ + Myd88i E F Ctrl Abx 350 * 300 250 200 150 40X 100 Tumor weight (mg) 50 0 x Ctrl Ab Supplemental Figure 2 A KC WT B ** * Actinobacteria * ** Bacteroidetes Cyanobacteria Deferribacteres * Firmicutes Proteobacteria % Relative abundance TM7 Others Time(wks) 3 9 13 16 20 24 28 32 36 3 9 13 16 20 24 28 32 36 Alpha Diversity Measure C E 60 KC WT 40 20 B. pseudolongum B. animalis 60 5 KC WT 0 B. adolescentis 40% Rel. abundance 3 9 13 16 20 24 28 32 36 3 9 13 16 20 24 28 32 36 20 Age (weeks) B. pseudolongum B. animalis 5 0 B. adolescentis % Rel. abundance 3 9 13 16 20 24 28 32 36 3 9 13 16 20 24 28 32 36 F Age (weeks) Week 3 Week 9 Week 13 p=0.678 p=0.02 p=0.385 Time(wks) 3 9 24 20 13 16 D 28 32 36 Week 13 KC WT Firmicutes; Ruminococcus Firmicutes; Dehalobacterium Alpha Diversity Measure Firmicutes; Oscillospira Bacteroidates; Odoribacter Axis.2 [12.7%] Actinobacteria; Bifidobacterium Axis.2 [23.8%] Axis.2 [24.7%] Week 16 Bacteroidetes; Bacteroidales Axis.1 [80.8%] Axis.1 [65.4%] Axis.1 [49.6%] Actinobacteria; Bifidobacterium Week 16 Week 20 Week 24 Week 20 p=0.339 p=0.036 p=0.021 Firmicutes; Dehalobacterium