Predicting Glioblastoma Prognosis Networks Using Weighted Gene Co-Expression Network Analysis on TCGA Data
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

1 AGING Supplementary Table 2
SUPPLEMENTARY TABLES Supplementary Table 1. Details of the eight domain chains of KIAA0101. Serial IDENTITY MAX IN COMP- INTERFACE ID POSITION RESOLUTION EXPERIMENT TYPE number START STOP SCORE IDENTITY LEX WITH CAVITY A 4D2G_D 52 - 69 52 69 100 100 2.65 Å PCNA X-RAY DIFFRACTION √ B 4D2G_E 52 - 69 52 69 100 100 2.65 Å PCNA X-RAY DIFFRACTION √ C 6EHT_D 52 - 71 52 71 100 100 3.2Å PCNA X-RAY DIFFRACTION √ D 6EHT_E 52 - 71 52 71 100 100 3.2Å PCNA X-RAY DIFFRACTION √ E 6GWS_D 41-72 41 72 100 100 3.2Å PCNA X-RAY DIFFRACTION √ F 6GWS_E 41-72 41 72 100 100 2.9Å PCNA X-RAY DIFFRACTION √ G 6GWS_F 41-72 41 72 100 100 2.9Å PCNA X-RAY DIFFRACTION √ H 6IIW_B 2-11 2 11 100 100 1.699Å UHRF1 X-RAY DIFFRACTION √ www.aging-us.com 1 AGING Supplementary Table 2. Significantly enriched gene ontology (GO) annotations (cellular components) of KIAA0101 in lung adenocarcinoma (LinkedOmics). Leading Description FDR Leading Edge Gene EdgeNum RAD51, SPC25, CCNB1, BIRC5, NCAPG, ZWINT, MAD2L1, SKA3, NUF2, BUB1B, CENPA, SKA1, AURKB, NEK2, CENPW, HJURP, NDC80, CDCA5, NCAPH, BUB1, ZWILCH, CENPK, KIF2C, AURKA, CENPN, TOP2A, CENPM, PLK1, ERCC6L, CDT1, CHEK1, SPAG5, CENPH, condensed 66 0 SPC24, NUP37, BLM, CENPE, BUB3, CDK2, FANCD2, CENPO, CENPF, BRCA1, DSN1, chromosome MKI67, NCAPG2, H2AFX, HMGB2, SUV39H1, CBX3, TUBG1, KNTC1, PPP1CC, SMC2, BANF1, NCAPD2, SKA2, NUP107, BRCA2, NUP85, ITGB3BP, SYCE2, TOPBP1, DMC1, SMC4, INCENP. RAD51, OIP5, CDK1, SPC25, CCNB1, BIRC5, NCAPG, ZWINT, MAD2L1, SKA3, NUF2, BUB1B, CENPA, SKA1, AURKB, NEK2, ESCO2, CENPW, HJURP, TTK, NDC80, CDCA5, BUB1, ZWILCH, CENPK, KIF2C, AURKA, DSCC1, CENPN, CDCA8, CENPM, PLK1, MCM6, ERCC6L, CDT1, HELLS, CHEK1, SPAG5, CENPH, PCNA, SPC24, CENPI, NUP37, FEN1, chromosomal 94 0 CENPL, BLM, KIF18A, CENPE, MCM4, BUB3, SUV39H2, MCM2, CDK2, PIF1, DNA2, region CENPO, CENPF, CHEK2, DSN1, H2AFX, MCM7, SUV39H1, MTBP, CBX3, RECQL4, KNTC1, PPP1CC, CENPP, CENPQ, PTGES3, NCAPD2, DYNLL1, SKA2, HAT1, NUP107, MCM5, MCM3, MSH2, BRCA2, NUP85, SSB, ITGB3BP, DMC1, INCENP, THOC3, XPO1, APEX1, XRCC5, KIF22, DCLRE1A, SEH1L, XRCC3, NSMCE2, RAD21. -

A Homozygous MRPL24 Mutation Causes a Complex Movement Disorder and Affects the Mitoribosome Assembly T
Neurobiology of Disease 141 (2020) 104880 Contents lists available at ScienceDirect Neurobiology of Disease journal homepage: www.elsevier.com/locate/ynbdi A homozygous MRPL24 mutation causes a complex movement disorder and affects the mitoribosome assembly T Michela Di Nottiaa,1, Maria Marcheseb,1, Daniela Verrignia, Christian Daniel Muttic, Alessandra Torracoa, Romina Olivad, Erika Fernandez-Vizarrac, Federica Moranib, Giulia Trania, Teresa Rizzaa, Daniele Ghezzie,f, Anna Ardissoneg,h, Claudia Nestib, Gessica Vascoi, Massimo Zevianic, Michal Minczukc, Enrico Bertinia, Filippo Maria Santorellib,2, ⁎ Rosalba Carrozzoa, ,2 a Unit of Muscular and Neurodegenerative Disorders, Laboratory of Molecular Medicine, Bambino Gesù Children's Hospital, IRCCS, Rome, Italy b Molecular Medicine & Neurogenetics, IRCCS Fondazione Stella Maris, Pisa, Italy c MRC Mitochondrial Biology Unit, University of Cambridge, Cambridge, UK d Department of Sciences and Technologies, University Parthenope of Naples, Naples, Italy e Unit of Medical Genetics and Neurogenetics, Fondazione IRCCS Istituto Neurologico Carlo Besta, Milan, Italy f Department of Pathophysiology and Transplantation, University of Milan, Milan, Italy g Child Neurology Unit, Fondazione IRCCS Istituto Neurologico Carlo Besta, Milan, Italy h Department of Molecular and Translational Medicine DIMET, University of Milan-Bicocca, Milan, Italy i Department of Neurosciences, IRCCS Bambino Gesù Children Hospital, Rome, Italy ARTICLE INFO ABSTRACT Keywords: Mitochondrial ribosomal protein large 24 (MRPL24) is 1 of the 82 protein components of mitochondrial ribo- Mitochondrial disorders somes, playing an essential role in the mitochondrial translation process. Movement disorder We report here on a baby girl with cerebellar atrophy, choreoathetosis of limbs and face, intellectual dis- MRPL24 ability and a combined defect of complexes I and IV in muscle biopsy, caused by a homozygous missense mu- Mitoribosomes tation identified in MRPL24. -

The Pennsylvania State University the Graduate School Eberly
The Pennsylvania State University The Graduate School Eberly College of Science REGULATION OF MITOCHONDRIAL TRANSLATION AND OXIDATIVE PHOSPHORYLATION THROUGH REVERSIBLE ACETYLATION A Dissertation in Biochemistry, Microbiology and Molecular Biology by Hüseyin Çimen 2012 Hüseyin Çimen Submitted in Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy August 2012 The Dissertation of Hüseyin Çimen was reviewed and approved* by the following: Emine C. Koc Assistant Professor of Biochemistry and Molecular Biology Dissertation Co-adviser Co-chair of Committee Hasan Koc Assistant Professor of Natural Sciences Dissertation Co-adviser Co-chair of Committee Craig E. Cameron Paul Berg Professor of Biochemistry and Molecular Biology Associate Department Head for Research and Graduate Education Joseph C. Reese Professor of Biochemistry and Molecular Biology Teh-hui Kao Professor of Biochemistry and Molecular Biology Tae-Hee Lee Assistant Professor of Chemistry and the Huck Institute of the Life Sciences Craig E. Cameron Paul Berg Professor of Biochemistry and Molecular Biology Associate Department Head of the Department of Biochemistry and Molecular Biology iii ABSTRACT In a eukaryotic cell, mitochondria provide energy in the form of ATP through oxidative phosphorylation (OXPHOS), which consists of five electron transport chain complexes embedded in the inner membrane of mitochondria. Human mitochondria have their own genome and transcription/translation system to synthesize mitochondrially encoded thirteen proteins of respiratory chain complexes. We investigated how acetylation of ribosomal proteins regulates translation and energy production in mitochondria since reversible acetylation of mitochondrial proteins was found to be critical for maintaining energy homeostasis. We identified mitochondrial ribosomal protein L10 (MRPL10) as the major acetylated ribosomal protein in mammalian mitochondria with two-dimensional gel electrophoresis followed by tandem mass spectrometry and immunoblotting analyses. -

CLPP Coordinates Mitoribosomal Assembly Through the Regulation of ERAL1 Levels
Article CLPP coordinates mitoribosomal assembly through the regulation of ERAL1 levels Karolina Szczepanowska1,2,†, Priyanka Maiti1,2,†, Alexandra Kukat1,2, Eduard Hofsetz1,2, Hendrik Nolte1,3, Katharina Senft1,2, Christina Becker1,2, Benedetta Ruzzenente4, Hue-Tran Hornig-Do1,5, Rolf Wibom6, Rudolf J Wiesner1,5, Marcus Krüger1,3 & Aleksandra Trifunovic1,2,* Abstract the elimination of specific proteins when they are no longer needed. The irreversible nature of this process demands rigorous selectivity Despite being one of the most studied proteases in bacteria, very in the choice of substrates and the degradation timing. In prokary- little is known about the role of ClpXP in mitochondria. We now otes, regulated protein degradation is governed by energy-dependent present evidence that mammalian CLPP has an essential role in proteases, of which ClpXP is probably the best studied as it contri- determining the rate of mitochondrial protein synthesis by regu- butes to a large fraction of protein turnover in bacteria (Sauer & lating the level of mitoribosome assembly. Through a proteomic Baker, 2011). ClpXP is a highly conserved proteasome-like machin- approach and the use of a catalytically inactive CLPP, we produced ery present in all bacterial species and is also found in the mitochon- the first comprehensive list of possible mammalian ClpXP dria and chloroplasts of eukaryotic cells (Sauer & Baker, 2011). In substrates involved in the regulation of mitochondrial translation, mammals, ClpXP protein substrates are recognized and unfolded by oxidative phosphorylation, and a number of metabolic pathways. the hexameric AAA+ protein CLPX that delivers them to the CLPP We further show that the defect in mitoribosomal assembly is a protease for degradation (Sauer & Baker, 2011). -

Identification of Gene Expression Pattern Related to Breast Cancer Survival Using Integrated TCGA Datasets and Genomic Tools
Hindawi Publishing Corporation BioMed Research International Volume 2015, Article ID 878546, 10 pages http://dx.doi.org/10.1155/2015/878546 Research Article Identification of Gene Expression Pattern Related to Breast Cancer Survival Using Integrated TCGA Datasets and Genomic Tools Zhenzhen Huang,1 Huilong Duan,1 and Haomin Li2,3 1 College of Biomedical Engineering and Instrument Science, Zhejiang University, Zhouyiqing Building No. 510, Yuquan Campus, Hangzhou 310027, China 2TheChildren’sHospital,ZhejiangUniversity,ZhouyiqingBuildingNo.510,YuquanCampus,Hangzhou310003,China 3The Institute of Translational Medicine, Zhejiang University, Zhouyiqing Building No. 510, Yuquan Campus, Hangzhou 310029, China Correspondence should be addressed to Haomin Li; [email protected] Received 3 July 2015; Revised 14 September 2015; Accepted 28 September 2015 Academic Editor: S´ılvia A. Sousa Copyright © 2015 Zhenzhen Huang et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Several large-scale human cancer genomics projects such as TCGA offered huge genomic and clinical data for researchers to obtain meaningful genomics alterations which intervene in the development and metastasis of the tumor. A web-based TCGA data analysis platform called TCGA4U was developed in this study. TCGA4U provides a visualization solution for this study to illustrate the relationship of these genomics alternations with clinical data. A whole genome screening of the survival related gene expression patterns in breast cancer was studied. The gene list that impacts the breast cancer patient survival was divided into two patterns. Gene list of each of these patterns was separately analyzed on DAVID. -

Transcriptomic and Proteomic Landscape of Mitochondrial
TOOLS AND RESOURCES Transcriptomic and proteomic landscape of mitochondrial dysfunction reveals secondary coenzyme Q deficiency in mammals Inge Ku¨ hl1,2†*, Maria Miranda1†, Ilian Atanassov3, Irina Kuznetsova4,5, Yvonne Hinze3, Arnaud Mourier6, Aleksandra Filipovska4,5, Nils-Go¨ ran Larsson1,7* 1Department of Mitochondrial Biology, Max Planck Institute for Biology of Ageing, Cologne, Germany; 2Department of Cell Biology, Institute of Integrative Biology of the Cell (I2BC) UMR9198, CEA, CNRS, Univ. Paris-Sud, Universite´ Paris-Saclay, Gif- sur-Yvette, France; 3Proteomics Core Facility, Max Planck Institute for Biology of Ageing, Cologne, Germany; 4Harry Perkins Institute of Medical Research, The University of Western Australia, Nedlands, Australia; 5School of Molecular Sciences, The University of Western Australia, Crawley, Australia; 6The Centre National de la Recherche Scientifique, Institut de Biochimie et Ge´ne´tique Cellulaires, Universite´ de Bordeaux, Bordeaux, France; 7Department of Medical Biochemistry and Biophysics, Karolinska Institutet, Stockholm, Sweden Abstract Dysfunction of the oxidative phosphorylation (OXPHOS) system is a major cause of human disease and the cellular consequences are highly complex. Here, we present comparative *For correspondence: analyses of mitochondrial proteomes, cellular transcriptomes and targeted metabolomics of five [email protected] knockout mouse strains deficient in essential factors required for mitochondrial DNA gene (IKu¨ ); expression, leading to OXPHOS dysfunction. Moreover, -

Distinct Genetic Basis of Closely Related Toad Tadpoles Respectively Adapted to High Altitude and Karst Caves
G C A T T A C G G C A T genes Article Plateau Grass and Greenhouse Flower? Distinct Genetic Basis of Closely Related Toad Tadpoles Respectively Adapted to High Altitude and Karst Caves 1,2, 1, 1,2 1,2 1, Liming Chang y, Wei Zhu y, Shengchao Shi , Meihua Zhang , Jianping Jiang *, Cheng Li 1, Feng Xie 1 and Bin Wang 1,* 1 CAS Key Laboratory of Mountain Ecological Restoration and Bioresource Utilization & Ecological Restoration and Biodiversity Conservation Key Laboratory of Sichuan Province, Chengdu Institute of Biology, Chinese Academy of Sciences, Chengdu 610041, China; [email protected] (L.C.); [email protected] (W.Z.); [email protected] (S.S.); [email protected] (M.Z.); [email protected] (C.L.); [email protected] (F.X.) 2 University of Chinese Academy of Sciences, Beijing 100049, China * Correspondence: [email protected] (B.W.); [email protected] (J.J.) These authors have contributed equally to this work. y Received: 1 December 2019; Accepted: 19 January 2020; Published: 22 January 2020 Abstract: Genetic adaptation to extremes is a fascinating topic. Nevertheless, few studies have explored the genetic adaptation of closely related species respectively inhabiting distinct extremes. With deep transcriptome sequencing, we attempt to detect the genetic architectures of tadpoles of five closely related toad species adapted to the Tibetan Plateau, middle-altitude mountains and karst caves. Molecular evolution analyses indicated that not only the number of fast evolving genes (FEGs), but also the functioning coverage of FEGs, increased with elevation. Enrichment analyses correspondingly revealed that the highland species had most of the FEGs involved in high-elevation adaptation, for example, amino acid substitutions of XRCC6 in its binding domains might improve the capacity of DNA repair of the toad. -
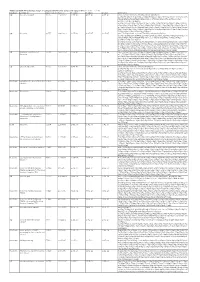
Additional Tables.Xlsx
Additional Table 5 Enriched pathways of upregulated DEGs after spinal cord injury in WT (P < 0.05, q < 0.05) Database Description Ratio of DEGs Ratio of P -value P adjust q value gene name GO ribosomal subunit 87/755 214/12262 1.03E-49 4.61E-47 4.20E-47 Gm6576/Gm10036/Gm5786/Mrps28/Gm9493/Rpl13- ps3/Rpl7l1/Mrpl10/Mrpl44/Rps20/Mrpl51/Mrps11/Mpv17l2/Mrps7/Gm10020/Gm10073 /Mrpl2/Mrpl52/Rpl22/Mrpl4/Mrps24/Zcchc17/Rpl28/Mrpl34/Rpl29/Mrps36/Rpl9- ps6/Hba-a2/Hba-a1/Rpl23a- ps3/Rpl15/Rack1/Mrps18a/Mrpl28/Rpl10a/Mrpl12/Rpl18/Rpl35a/Mrpl41/Mrps12/Rpl36 a/Rps9/Rps18/Rps5/Rps27a/Rps24/Rplp2/Rpl14/Mrpl30/Rps6/Rpl6/Rps3/Rps4x/Rpsa/R pl7/Rpl27a/Rps23/Rps10/Mrps21/Rpl36/Rpl30/Rps17/Mrpl42/Rpl35/Rpl27/Rplp0/Rps28 /Rpl31/Rpl39/Rps13/Rpl13/Rps11/Rpl8/Rps3a1/Fau/Rpl9/Rpl3/Rpl26/Rps15/Rpl13a/Uba 52/Rpl24/Rpl19/Rps14/Rpl4/Rps27/Rplp1 GO ribosome 93/755 248/12262 1.67E-49 4.61E-47 4.20E-47 Gm6576/Pnpt1/Gm10036/Gm5786/Mrps28/Gm9493/Rpl13- ps3/Rpl7l1/Mrpl10/Mrpl44/Rps20/Mrpl51/Mrps11/Mpv17l2/Mrps7/Gm10020/Gm10073 /Mrpl2/Mrpl52/Rpl22/Mrpl4/Mrps24/Zcchc17/Rpl28/Mrpl34/Rpl29/Mrps36/Rpl9- ps6/Hba-a2/Hba-a1/Rpl23a- ps3/Rpl15/Rack1/Eif3h/Mrps18a/Mrpl28/Mrps33/Rpl10a/Mrpl12/Rpl18/Rpl35a/Btf3/Mrp l41/Mrps12/Rpl36a/Rps9/Rps18/Rps5/Rps27a/Rps24/Rplp2/Rpl14/Mrpl30/Rps6/Rpl6/R ps3/Rps4x/Rpsa/Rpl7/Rpl27a/Rps23/Rps10/Mrps21/Rpl36/Rpl30/Rps17/Mrpl42/Rpl35/R pl27/Rplp0/Rps28/Rpl31/Rpl39/Rps13/Rpl13/Rps11/Rpl8/Rps3a1/Fau/Rpl9/Rpl3/Rpl26/ Rps15/Rpl13a/Uba52/Rpl41/Rpl24/Rpl19/Rps14/Rpl4/Ndufa7/Rps27/Rplp1 GO structural constituent of 76/733 169/11958 1.65E-47 1.16E-44 1.12E-44 Gm6576/Gm10036/Gm5786/Rpl7l1/Mrpl10/Rps20/Mrpl51/Mrps11/Mrps7/Gm10020/G -

MRPS7 Rabbit Pab
Leader in Biomolecular Solutions for Life Science MRPS7 Rabbit pAb Catalog No.: A16272 Basic Information Background Catalog No. Mammalian mitochondrial ribosomal proteins are encoded by nuclear genes and help in A16272 protein synthesis within the mitochondrion. Mitochondrial ribosomes (mitoribosomes) consist of a small 28S subunit and a large 39S subunit. They have an estimated 75% Observed MW protein to rRNA composition compared to prokaryotic ribosomes, where this ratio is 28kDa reversed. Another difference between mammalian mitoribosomes and prokaryotic ribosomes is that the latter contain a 5S rRNA. Among different species, the proteins Calculated MW comprising the mitoribosome differ greatly in sequence, and sometimes in biochemical 28kDa properties, which prevents easy recognition by sequence homology. This gene encodes a 28S subunit protein. In the prokaryotic ribosome, the comparable protein is thought to Category play an essential role in organizing the 3' domain of the 16 S rRNA in the vicinity of the P- and A-sites. Pseudogenes corresponding to this gene are found on chromosomes 8p Primary antibody and 12p. Applications WB Cross-Reactivity Human, Mouse, Rat Recommended Dilutions Immunogen Information WB 1:500 - 1:2000 Gene ID Swiss Prot 51081 Q9Y2R9 Immunogen Recombinant fusion protein containing a sequence corresponding to amino acids 110-190 of human MRPS7 (NP_057055.2). Synonyms MRPS7;MRP-S;MRP-S7;RP-S7;RPMS7;S7mt;bMRP27a Contact Product Information www.abclonal.com Source Isotype Purification Rabbit IgG Affinity purification Storage Store at -20℃. Avoid freeze / thaw cycles. Buffer: PBS with 0.02% sodium azide,50% glycerol,pH7.3. Validation Data Western blot analysis of extracts of various cell lines, using MRPS7 antibody (A16272) at 1:1000 dilution. -

Impaired Mitochondrial Biogenesis in Adipose Tissue in Acquired Obesity
Diabetes Volume 64, September 2015 3135 Sini Heinonen,1 Jana Buzkova,2 Maheswary Muniandy,1 Risto Kaksonen,1,3 Miina Ollikainen,4 Khadeeja Ismail,4 Antti Hakkarainen,5 Jesse Lundbom,5,6 Nina Lundbom,5 Katriina Vuolteenaho,7 Eeva Moilanen,7 Jaakko Kaprio,8,9,10 Aila Rissanen,1,11 Anu Suomalainen,2,12 and Kirsi H. Pietiläinen1,8,13 Impaired Mitochondrial Biogenesis in Adipose Tissue in Acquired Obesity Diabetes 2015;64:3135–3145 | DOI: 10.2337/db14-1937 Low mitochondrial number and activity have been sug- and OXPHOS proteins in SAT are downregulated in ac- gested as underlying factors in obesity, type 2 diabetes, quired obesity, and are associated with metabolic distur- and metabolic syndrome. However, the stage at which bances already at the preclinical stage. mitochondrial dysfunction manifests in adipose tissue after the onset of obesity remains unknown. Here we examined subcutaneous adipose tissue (SAT) samples Adipocytes are important contributors to energy balance – from healthy monozygotic twin pairs, 22.8 36.2 years of and metabolic homeostasis. Within these highly dynamic D > 2 age, who were discordant ( BMI 3kg/m, mean length cells, mitochondria are at the center of energy metabo- OBESITY STUDIES 6 n of discordance 6.3 0.3 years, = 26) and concordant lism, using carbohydrates, lipids, and proteins to produce D < 2 n ( BMI 3kg/m, = 14) for body weight, and assessed ATP and metabolites for growth, as well as contributing to their detailed mitochondrial metabolic characteristics: adipocyte differentiation and maturation (1). Mitochon- mitochondrial-related transcriptomes with dysregulated dria possess their own multicopy genome, a 16.6-kb cir- pathways, mitochondrial DNA (mtDNA) amount, mtDNA- cular mitochondrial DNA (mtDNA) that encodes two encoded transcripts, and mitochondrial oxidative phos- phorylation (OXPHOS) protein levels. -

Comprehensive Mitochondrial Nuclear Gene Panel
Comprehensive Mitochondrial Nuclear Gene Panel Test Code D4606 Test Summary This panel analyzes 254 genes that have been associated with mitochondrial disorders in nuclear genes. Turn-Around-Time (TAT)* 3 - 5 weeks Acceptable Sample Types Whole Blood (EDTA) (Preferred sample type) DNA, Isolated Dried Blood Spots Saliva Acceptable Billing Types Self (patient) Payment Institutional Billing Commercial Insurance Indications for Testing This test may be appropriate for individuals who have a clinical suspicion of a mitochondrial disorder and/or family history of a mitochondrial disorder. Test Description This panel analyzes 254 genes that have been associated with mitochondrial disorders in nuclear genes. Both sequencing and deletion/duplication (CNV) analysis will be performed on the coding regions of all genes included (unless otherwise marked). All analysis is performed utilizing Next Generation Sequencing (NGS) technology. CNV analysis is designed to detect deletions and duplications of three exons or greater in size. All variants are classified according to ACMG guidelines. Condition Description Mitochondrial disease is a varied group of disorders characterized by impaired energy production. Hallmarks of mitochondrial disease include neurological, cardiovascular, ophthalmologic and gastroenterological features. It is estimated that 1/5,000 individuals has a genetic mitochondrial disease. Genes AARS, AARS2, ABCB11, ABCB4, ABCB7, ABCD4, ACAD9, ACADM, ACADVL, ACO2, ACSF3, ADCK3, ADCK4, AFG3L2, AGK, AGL, AIFM1, ALAS2, ALDOA, ALDOB, ALG1, -
Mitochondrial Ribosomal Proteins: Candidate Genes for Mitochondrial Disease James E
March/April 2004 ⅐ Vol. 6 ⅐ No. 2 review Mitochondrial ribosomal proteins: Candidate genes for mitochondrial disease James E. Sylvester, PhD1, Nathan Fischel-Ghodsian, MD2, Edward B. Mougey, PhD1, and Thomas W. O’Brien, PhD3 Most of the energy requirement for cell growth, differentiation, and development is met by the mitochondria in the form of ATP produced by the process of oxidative phosphorylation. Human mitochondrial DNA encodes a total of 13 proteins, all of which are essential for oxidative phosphorylation. The mRNAs for these proteins are translated on mitochondrial ribosomes. Recently, the genes for human mitochondrial ribosomal proteins (MRPs) have been identified. In this review, we summarize their refined chromosomal location. It is well known that mutations in the mitochondrial translation system, i.e., ribosomal RNA and transfer RNA cause various pathologies. In this review, we suggest possible associations between clinical conditions and MRPs based on coincidence of genetic map data and chromosomal location. These MRPs may be candidate genes for the clinical condition or may act as modifiers of existing known gene mutations (mt-tRNA, mt-rRNA, etc.). Genet Med 2004:6(2):73–80. Key Words: mitochondrial, ribosomal proteins, oxidative phosphorylation, candidate genes, translation Most of the energy requirements for cell growth, differenti- THE MITORIBOSOME ation, and development are met by the mitochondrial ATP Human cells contain two genomes and two protein synthe- produced by the process of oxidative phosphorylation. Mito- sizing (translation) systems. The first is the nuclear genome of chondrial DNA encodes 13 essential proteins of this oxidative 3 ϫ 109 bp that has 30,000 to 40,000 genes coding a much phosphorylation system.