Histone Target Selection Within Chromatin: an Exemplary Case of Teamwork
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Expression Pattern and Level of ING5 Protein in Normal and Cancer Tissues
ONCOLOGY LETTERS 17: 63-68, 2019 Expression pattern and level of ING5 protein in normal and cancer tissues XUE-FENG YANG1, DAO-FU SHEN1, SHUANG ZHAO1, TIAN-REN REN2, YANG GAO1, SHUAI SHI1, JI-CHENG WU1, HONG-ZHI SUN1 and HUA-CHUAN ZHENG1,3 1Cancer Center and Key Laboratory of Brain and Spinal Cord Injury of Liaoning Province, The First Affiliated Hospital of Jinzhou Medical University, Jinzhou, Liaoning 121001;2 Jilin Province Forestry Bureau, Linjiang, Jilin 134600; 3Institute of Life Sciences, Jinzhou Medical University, Jinzhou, Liaoning 121001, P.R. China Received February 21, 2016; Accepted February 13, 2017 DOI: 10.3892/ol.2018.9581 Abstract. Inhibitor of growth family 5 (ING5) functions as may be involved in cell regeneration and be associated with a type-II tumor suppressor gene and exerts an important role colorectal carcinogenesis. in DNA repair, apoptotic induction, proliferative inhibition, chromatin remodeling and the invasion process. In the present Introduction study, immunohistochemistry was performed to characterize the expression profile of ING5 protein on a tissue microarray Inhibitor of growth family 5 (ING5) is a member of the ING containing mouse and human normal tissues, and human protein family and functions as a type-II tumor suppressor cancer tissues, including hepatocellular (n=62), renal clear cell gene. Human ING5 is mapped to chromosome 2q37.3 and (n=62), pancreatic (n=62), esophageal squamous cell (n=45), encodes a 28-kDa protein with 240 amino acids (1). As indi- cervical squamous cell (n=31), breast (n=144), gastric (n=196), cated in Fig. 1, ING5 protein consists of a number of domains, colorectal (n=96), endometrial (n=96) and lung carcinoma including leucine zipper like (LZL), novel conserved region (n=192). -

The Tumor Suppressor ING5 Is a Dimeric, Bivalent Recognition Molecule of the Histone H3k4me3 Mark
Article The Tumor Suppressor ING5 Is a Dimeric, Bivalent Recognition Molecule of the Histone H3K4me3 Mark Georgina Ormaza 1,†, Jhon A. Rodríguez 1,†, Alain Ibáñez de Opakua 1, Nekane Merino 1, Maider Villate 1, Irantzu Gorroño 1, Miriam Rábano 1, Ignacio Palmero 2, Marta Vilaseca 3, Robert Kypta 1,4, María d.M. Vivanco 1, Adriana L. Rojas 1 and Francisco J. Blanco 1,5 1 - CIC bioGUNE, Parque Tecnológico de Bizkaia, 48160 Derio, Spain 2 - Instituto de Investigaciones Biomédicas “Alberto Sols”, CSIC-UAM, 28029 Madrid, Spain 3 - Institute for Research in Biomedicine, 08028 Barcelona, Spain 4 - Department of Surgery and Cancer, Imperial College London, London, W12 0NN, UK 5 - IKERBASQUE, Basque Foundation for Science, 48011 Bilbao, Spain Correspondence to Francisco J. Blanco: CIC bioGUNE, Parque Tecnológico de Bizkaia, 48160 Derio, Spain. [email protected] https://doi.org/10.1016/j.jmb.2019.04.018 Edited by M. Guss Abstract The INhibitor of Growth (ING) family of tumor suppressors regulates the transcriptional state of chromatin by recruiting remodeling complexes to sites with histone H3 trimethylated at lysine 4 (H3K4me3). This modification is recognized by the plant homeodomain (PHD) present at the C-terminus of the five ING proteins. ING5 facilitates histone H3 acetylation by the HBO1 complex, and also H4 acetylation by the MOZ/ MORF complex. We show that ING5 forms homodimers through its N-terminal domain, which folds independently into an elongated coiled-coil structure. The central region of ING5, which contains the nuclear localization sequence, is flexible and disordered, but it binds dsDNA with micromolar affinity. NMR analysis of the full-length protein reveals that the two PHD fingers of the dimer are chemically equivalent and independent of the rest of the molecule, and they bind H3K4me3 in the same way as the isolated PHD. -

Subcloning, Expression, and Enzymatic Study of PRMT5
Georgia State University ScholarWorks @ Georgia State University Biology Theses Department of Biology Summer 7-12-2010 Subcloning, Expression, and Enzymatic Study of PRMT5 Ran Guo Georgia State University Follow this and additional works at: https://scholarworks.gsu.edu/biology_theses Part of the Biology Commons Recommended Citation Guo, Ran, "Subcloning, Expression, and Enzymatic Study of PRMT5." Thesis, Georgia State University, 2010. https://scholarworks.gsu.edu/biology_theses/26 This Thesis is brought to you for free and open access by the Department of Biology at ScholarWorks @ Georgia State University. It has been accepted for inclusion in Biology Theses by an authorized administrator of ScholarWorks @ Georgia State University. For more information, please contact [email protected]. SUBCLONING, EXPRESSION, AND ENZYMATIC STUDY OF PRMT5 by RAN GUO Under the Direction of Yujun George Zheng ABSTRACT Protein arginine methyltransferases (PRMTs) mediate the transfer of methyl groups to arginine residues in histone and non-histone proteins. PRMT5 is an important member of PRMTs which symmetrically dimethylates arginine 8 in histone H3 (H3R8) and arginine 3 in histone H4 (H4R3). PRMT5 was reported to inhibit some tumor suppressors in leukemia and lymphoma cells and regulate p53 gene, through affecting the promoter of p53. Through methylation of H4R3, PRMT5 can recruit DNA-methyltransferase 3A (DNMT3A) which regulates gene transcription. All the above suggest that PRMT5 has an important function of suppressing cell apoptosis and is a potential anticancer target. Currently, the enzymatic activities of PRMT5 are not clearly understood. In our study, we improved the protein expression methodology and greatly enhanced the yield and quality of the recombinant PRMT5. -

Modes of Interaction of KMT2 Histone H3 Lysine 4 Methyltransferase/COMPASS Complexes with Chromatin
cells Review Modes of Interaction of KMT2 Histone H3 Lysine 4 Methyltransferase/COMPASS Complexes with Chromatin Agnieszka Bochy ´nska,Juliane Lüscher-Firzlaff and Bernhard Lüscher * ID Institute of Biochemistry and Molecular Biology, Medical School, RWTH Aachen University, Pauwelsstrasse 30, 52057 Aachen, Germany; [email protected] (A.B.); jluescher-fi[email protected] (J.L.-F.) * Correspondence: [email protected]; Tel.: +49-241-8088850; Fax: +49-241-8082427 Received: 18 January 2018; Accepted: 27 February 2018; Published: 2 March 2018 Abstract: Regulation of gene expression is achieved by sequence-specific transcriptional regulators, which convey the information that is contained in the sequence of DNA into RNA polymerase activity. This is achieved by the recruitment of transcriptional co-factors. One of the consequences of co-factor recruitment is the control of specific properties of nucleosomes, the basic units of chromatin, and their protein components, the core histones. The main principles are to regulate the position and the characteristics of nucleosomes. The latter includes modulating the composition of core histones and their variants that are integrated into nucleosomes, and the post-translational modification of these histones referred to as histone marks. One of these marks is the methylation of lysine 4 of the core histone H3 (H3K4). While mono-methylation of H3K4 (H3K4me1) is located preferentially at active enhancers, tri-methylation (H3K4me3) is a mark found at open and potentially active promoters. Thus, H3K4 methylation is typically associated with gene transcription. The class 2 lysine methyltransferases (KMTs) are the main enzymes that methylate H3K4. KMT2 enzymes function in complexes that contain a necessary core complex composed of WDR5, RBBP5, ASH2L, and DPY30, the so-called WRAD complex. -

Inferring Causal Relationships Among Different Histone Modifications and Gene Expression
Downloaded from genome.cshlp.org on September 28, 2021 - Published by Cold Spring Harbor Laboratory Press Inferring Causal Relationships among Different Histone Modifications and Gene Expression Hong Yu*, Shanshan Zhu*, Bing Zhou*, Huiling Xue and Jing-Dong J. Han Chinese Academy of Sciences Key Laboratory of Molecular Developmental Biology, Center for Molecular Systems Biology, Institute of Genetics and Developmental Biology, Chinese Academy of Sciences, Datun Road, Beijing, 100101, China * These authors contributed equally to this work. Correspondence should be addressed to J.-D.J.H ([email protected]). Running title: histone modification and gene expression network Keywords: histone modification, gene expression, Bayesian network, causal relationship, epigenetics 1 Downloaded from genome.cshlp.org on September 28, 2021 - Published by Cold Spring Harbor Laboratory Press Abstract Histone modifications are major epigenetic factors regulating gene expression. They play important roles in maintaining stem cell pluripotency and in cancer pathogenesis. Different modifications may combine to form complex ‘histone codes’. Recent high throughput technologies, such as ‘ChIP-chip’ and ‘ChIP-seq’, have generated high resolution maps for many histone modifications on the human genome. Here we use these maps to build a Bayesian network to infer causal and combinatorial relationships among histone modifications and gene expression. A pilot network derived by the same method among polycomb group (PcG) genes and H3K27 trimethylation is accurately supported by current literature. Our unbiased network model among histone modifications is also well supported by cross validation results. It not only confirmed already known relationships, such as those of H3K27me3 to gene silencing, H3K4me3 to gene activation, and the effect of bivalent modification of both H3K4me3 and H3K27me3, but also identified many other relationships that may predict new epigenetic interactions important in epigenetic gene regulation. -

The Role of ING5 in Maintaining Stemness of Brain Tumor Initiating Cells
University of Calgary PRISM: University of Calgary's Digital Repository Graduate Studies The Vault: Electronic Theses and Dissertations 2016 The Role of ING5 in Maintaining Stemness of Brain Tumor Initiating Cells Wang, Fangwu Jr Wang, F. J. (2016). The Role of ING5 in Maintaining Stemness of Brain Tumor Initiating Cells (Unpublished master's thesis). University of Calgary, Calgary, AB. doi:10.11575/PRISM/28327 http://hdl.handle.net/11023/3233 master thesis University of Calgary graduate students retain copyright ownership and moral rights for their thesis. You may use this material in any way that is permitted by the Copyright Act or through licensing that has been assigned to the document. For uses that are not allowable under copyright legislation or licensing, you are required to seek permission. Downloaded from PRISM: https://prism.ucalgary.ca UNIVERSITY OF CALGARY The Role of ING5 in Maintaining Stemness of Brain Tumor Initiating Cells by Fangwu Wang A THESIS SUBMITTED TO THE FACULTY OF GRADUATE STUDIES IN PARTIAL FULFILMENT OF THE REQUIREMENTS FOR THE DEGREE OF MASTER OF SCIENCE GRADUATE PROGRAM IN BIOCHEMISTRY AND MOLECULAR BIOLOGY CALGARY, ALBERTA AUGUST, 2016 © Fangwu Wang 2016 Abstract Brain tumor initiating cells (BTICs) are believed to account for the recurrence of glioblastomas following treatment. Recent studies have shown that the stemness of BTICs and intratumoral differentiation hierarchy are determined largely on the epigenetic level. The ING family of epigenetic regulators function in diverse growth regulatory, metastasis and chemoresistance pathways, through targeting different histone acetyltransferase (HAT) and histone deacetylase (HDAC) complexes to the H3K4me3 mark to alter histone acetylation. -

Open Data for Differential Network Analysis in Glioma
International Journal of Molecular Sciences Article Open Data for Differential Network Analysis in Glioma , Claire Jean-Quartier * y , Fleur Jeanquartier y and Andreas Holzinger Holzinger Group HCI-KDD, Institute for Medical Informatics, Statistics and Documentation, Medical University Graz, Auenbruggerplatz 2/V, 8036 Graz, Austria; [email protected] (F.J.); [email protected] (A.H.) * Correspondence: [email protected] These authors contributed equally to this work. y Received: 27 October 2019; Accepted: 3 January 2020; Published: 15 January 2020 Abstract: The complexity of cancer diseases demands bioinformatic techniques and translational research based on big data and personalized medicine. Open data enables researchers to accelerate cancer studies, save resources and foster collaboration. Several tools and programming approaches are available for analyzing data, including annotation, clustering, comparison and extrapolation, merging, enrichment, functional association and statistics. We exploit openly available data via cancer gene expression analysis, we apply refinement as well as enrichment analysis via gene ontology and conclude with graph-based visualization of involved protein interaction networks as a basis for signaling. The different databases allowed for the construction of huge networks or specified ones consisting of high-confidence interactions only. Several genes associated to glioma were isolated via a network analysis from top hub nodes as well as from an outlier analysis. The latter approach highlights a mitogen-activated protein kinase next to a member of histondeacetylases and a protein phosphatase as genes uncommonly associated with glioma. Cluster analysis from top hub nodes lists several identified glioma-associated gene products to function within protein complexes, including epidermal growth factors as well as cell cycle proteins or RAS proto-oncogenes. -

BMC Genomics Biomed Central
BMC Genomics BioMed Central Research article Open Access Determination of enriched histone modifications in non-genic portions of the human genome Jeffrey A Rosenfeld1,2,3, Zhibin Wang4, Dustin E Schones4, Keji Zhao4, Rob DeSalle3 and Michael Q Zhang*1 Address: 1Cold Spring Harbor Laboratory, Cold Spring Harbor, NY 11724 USA, 2Department of Biology, New York University, New York, NY USA, 3American Museum of Natural History, New York, NY USA and 4Laboratory of Molecular Immunology, National Heart, Lung and Blood Institute, NIH, Bethesda, MD USA Email: Jeffrey A Rosenfeld - [email protected]; Zhibin Wang - [email protected]; Dustin E Schones - [email protected]; Keji Zhao - [email protected]; Rob DeSalle - [email protected]; Michael Q Zhang* - [email protected] * Corresponding author Published: 31 March 2009 Received: 8 September 2008 Accepted: 31 March 2009 BMC Genomics 2009, 10:143 doi:10.1186/1471-2164-10-143 This article is available from: http://www.biomedcentral.com/1471-2164/10/143 © 2009 Rosenfeld et al; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Abstract Background: Chromatin immunoprecipitation followed by high-throughput sequencing (ChIP- seq) has recently been used to identify the modification patterns for the methylation and acetylation of many different histone tails in genes and enhancers. Results: We have extended the analysis of histone modifications to gene deserts, pericentromeres and subtelomeres. Using data from human CD4+ T cells, we have found that each of these non- genic regions has a particular profile of histone modifications that distinguish it from the other non- coding regions. -

Histone Methylation Regulation in Neurodegenerative Disorders
International Journal of Molecular Sciences Review Histone Methylation Regulation in Neurodegenerative Disorders Balapal S. Basavarajappa 1,2,3,4,* and Shivakumar Subbanna 1 1 Division of Analytical Psychopharmacology, Nathan Kline Institute for Psychiatric Research, Orangeburg, NY 10962, USA; [email protected] 2 New York State Psychiatric Institute, New York, NY 10032, USA 3 Department of Psychiatry, College of Physicians & Surgeons, Columbia University, New York, NY 10032, USA 4 New York University Langone Medical Center, Department of Psychiatry, New York, NY 10016, USA * Correspondence: [email protected]; Tel.: +1-845-398-3234; Fax: +1-845-398-5451 Abstract: Advances achieved with molecular biology and genomics technologies have permitted investigators to discover epigenetic mechanisms, such as DNA methylation and histone posttransla- tional modifications, which are critical for gene expression in almost all tissues and in brain health and disease. These advances have influenced much interest in understanding the dysregulation of epigenetic mechanisms in neurodegenerative disorders. Although these disorders diverge in their fundamental causes and pathophysiology, several involve the dysregulation of histone methylation- mediated gene expression. Interestingly, epigenetic remodeling via histone methylation in specific brain regions has been suggested to play a critical function in the neurobiology of psychiatric disor- ders, including that related to neurodegenerative diseases. Prominently, epigenetic dysregulation currently brings considerable interest as an essential player in neurodegenerative disorders, such as Alzheimer’s disease (AD), Parkinson’s disease (PD), Huntington’s disease (HD), Amyotrophic lateral sclerosis (ALS) and drugs of abuse, including alcohol abuse disorder, where it may facilitate connections between genetic and environmental risk factors or directly influence disease-specific Citation: Basavarajappa, B.S.; Subbanna, S. -
Download Validation Data
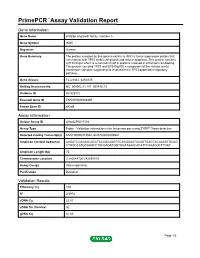
PrimePCR™Assay Validation Report Gene Information Gene Name inhibitor of growth family, member 5 Gene Symbol ING5 Organism Human Gene Summary The protein encoded by this gene is similar to ING1 a tumor suppressor protein that can interact with TP53 inhibit cell growth and induce apoptosis. This protein contains a PHD-finger which is a common motif in proteins involved in chromatin remodeling. This protein can bind TP53 and EP300/p300 a component of the histone acetyl transferase complex suggesting its involvement in TP53-dependent regulatory pathway. Gene Aliases FLJ23842, p28ING5 RefSeq Accession No. NC_000002.11, NT_005416.13 UniGene ID Hs.529172 Ensembl Gene ID ENSG00000168395 Entrez Gene ID 84289 Assay Information Unique Assay ID qHsaCIP0031594 Assay Type Probe - Validation information is for the primer pair using SYBR® Green detection Detected Coding Transcript(s) ENST00000313552, ENST00000406941 Amplicon Context Sequence AAGATCCAGAACGCCTACAGCAAGTGCAAGGAATACAGTGACGACAAAGTGCAG CTGGCCATGCAGACCTACGAGATGGTGGATAAACACATTCGAAGGCTTGAT Amplicon Length (bp) 75 Chromosome Location 2:242648720-242650818 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 100 R2 0.9978 cDNA Cq 22.07 cDNA Tm (Celsius) 82 gDNA Cq 41.86 Page 1/5 PrimePCR™Assay Validation Report Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report ING5, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak -

Stepwise Histone Modifications Are Mediated by Multiple Enzymes That
ARTICLE Received 4 Apr 2013 | Accepted 29 Oct 2013 | Published 26 Nov 2013 DOI: 10.1038/ncomms3841 Stepwise histone modifications are mediated by multiple enzymes that rapidly associate with nascent DNA during replication Svetlana Petruk1, Kathryn L. Black1, Sina K. Kovermann2, Hugh W. Brock2 & Alexander Mazo1 The mechanism of epigenetic inheritance following DNA replication may involve dissociation of chromosomal proteins from parental DNA and reassembly on daughter strands in a specific order. Here we investigated the behaviour of different types of chromosomal proteins using newly developed methods that allow assessment of the assembly of proteins during DNA replication. Unexpectedly, most chromatin-modifying proteins tested, including methylases, demethylases, acetyltransferases and a deacetylase, are found in close proximity to PCNA or associate with short nascent DNA. Histone modifications occur in a temporal order following DNA replication, mediated by complex activities of different enzymes. In contrast, components of several major nucleosome-remodelling complexes are dissociated from parental DNA, and are later recruited to nascent DNA following replication. Epigenetic inheritance of gene expression patterns may require many aspects of chromatin structure to remain in close proximity to the replication complex followed by reassembly on nascent DNA shortly after replication. 1 Department of Biochemistry and Molecular Biology and Kimmel Cancer Center, Thomas Jefferson University, 1020 Locust Street, Philadelphia, Pennsylvania 19107, USA. 2 Department of Zoology, University of British Columbia, 6270 University Boulevard, Vancouver, British Columbia, Canada V6T 1Z4. Correspondence and requests for materials should be addressed to A.M. (email: [email protected]). NATURE COMMUNICATIONS | 4:2841 | DOI: 10.1038/ncomms3841 | www.nature.com/naturecommunications 1 & 2013 Macmillan Publishers Limited. -

Global Histone Modifications in Breast Cancer Correlate with Tumor Phenotypes, Prognostic Factors, and Patient Outcome
Published OnlineFirst April 14, 2009; DOI: 10.1158/0008-5472.CAN-08-3907 Research Article Global Histone Modifications in Breast Cancer Correlate with Tumor Phenotypes, Prognostic Factors, and Patient Outcome Somaia E. Elsheikh,1,7 Andrew R. Green,1 Emad A. Rakha,1 Des G. Powe,1 Rabab A. Ahmed,1,8 Hilary M. Collins,2 Daniele Soria,3 Jonathan M. Garibaldi,3 Claire E. Paish,1 Amr A. Ammar,7 Matthew J. Grainge,4 Graham R. Ball,6 Magdy K. Abdelghany,2,9 Luisa Martinez-Pomares,5 David M. Heery,2 and Ian O. Ellis1 1Department of Histopathology, School of Molecular Medical Sciences, University of Nottingham and Nottingham Universities Hospital Trust, Schools of 2Pharmacy and 3Computer Science, 4Division of Epidemiology and Public Health, and 5School of Molecular Medical Sciences, Institute of Infection, Immunity and Inflammation, University of Nottingham; 6Division of Life Sciences, Nottingham Trent University, Nottingham, United Kingdom; 7Department of Pathology, Faculty of Medicine, Menoufyia University, Menoufyia, Egypt; 8Department of Pathology, Faculty of Medicine, Asuit University, Asuit, Egypt; and 9Department of Pathology, Faculty of Medicine, Suez Canal University, Ismailia, Egypt Abstract Introduction Post-translational histone modifications are known to be Breast cancer is a heterogeneous disease ranging from altered in cancer cells, and loss of selected histone acetylation premalignant hyperproliferation to invasive and metastatic carci- and methylation marks has recently been shown to predict nomas (1). Disease progression is poorly understood but is likely patient outcome in human carcinoma. Immunohistochemistry due to the accumulation of genetic mutations leading to was used to detect a series of histone lysine acetylation widespread changes in gene expression and, in particular, affecting (H3K9ac, H3K18ac, H4K12ac, and H4K16ac), lysine methyla- the expression of tumor suppressors and oncogenes (2).