The Role of ING5 in Maintaining Stemness of Brain Tumor Initiating Cells
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Expression Pattern and Level of ING5 Protein in Normal and Cancer Tissues
ONCOLOGY LETTERS 17: 63-68, 2019 Expression pattern and level of ING5 protein in normal and cancer tissues XUE-FENG YANG1, DAO-FU SHEN1, SHUANG ZHAO1, TIAN-REN REN2, YANG GAO1, SHUAI SHI1, JI-CHENG WU1, HONG-ZHI SUN1 and HUA-CHUAN ZHENG1,3 1Cancer Center and Key Laboratory of Brain and Spinal Cord Injury of Liaoning Province, The First Affiliated Hospital of Jinzhou Medical University, Jinzhou, Liaoning 121001;2 Jilin Province Forestry Bureau, Linjiang, Jilin 134600; 3Institute of Life Sciences, Jinzhou Medical University, Jinzhou, Liaoning 121001, P.R. China Received February 21, 2016; Accepted February 13, 2017 DOI: 10.3892/ol.2018.9581 Abstract. Inhibitor of growth family 5 (ING5) functions as may be involved in cell regeneration and be associated with a type-II tumor suppressor gene and exerts an important role colorectal carcinogenesis. in DNA repair, apoptotic induction, proliferative inhibition, chromatin remodeling and the invasion process. In the present Introduction study, immunohistochemistry was performed to characterize the expression profile of ING5 protein on a tissue microarray Inhibitor of growth family 5 (ING5) is a member of the ING containing mouse and human normal tissues, and human protein family and functions as a type-II tumor suppressor cancer tissues, including hepatocellular (n=62), renal clear cell gene. Human ING5 is mapped to chromosome 2q37.3 and (n=62), pancreatic (n=62), esophageal squamous cell (n=45), encodes a 28-kDa protein with 240 amino acids (1). As indi- cervical squamous cell (n=31), breast (n=144), gastric (n=196), cated in Fig. 1, ING5 protein consists of a number of domains, colorectal (n=96), endometrial (n=96) and lung carcinoma including leucine zipper like (LZL), novel conserved region (n=192). -

The Tumor Suppressor ING5 Is a Dimeric, Bivalent Recognition Molecule of the Histone H3k4me3 Mark
Article The Tumor Suppressor ING5 Is a Dimeric, Bivalent Recognition Molecule of the Histone H3K4me3 Mark Georgina Ormaza 1,†, Jhon A. Rodríguez 1,†, Alain Ibáñez de Opakua 1, Nekane Merino 1, Maider Villate 1, Irantzu Gorroño 1, Miriam Rábano 1, Ignacio Palmero 2, Marta Vilaseca 3, Robert Kypta 1,4, María d.M. Vivanco 1, Adriana L. Rojas 1 and Francisco J. Blanco 1,5 1 - CIC bioGUNE, Parque Tecnológico de Bizkaia, 48160 Derio, Spain 2 - Instituto de Investigaciones Biomédicas “Alberto Sols”, CSIC-UAM, 28029 Madrid, Spain 3 - Institute for Research in Biomedicine, 08028 Barcelona, Spain 4 - Department of Surgery and Cancer, Imperial College London, London, W12 0NN, UK 5 - IKERBASQUE, Basque Foundation for Science, 48011 Bilbao, Spain Correspondence to Francisco J. Blanco: CIC bioGUNE, Parque Tecnológico de Bizkaia, 48160 Derio, Spain. [email protected] https://doi.org/10.1016/j.jmb.2019.04.018 Edited by M. Guss Abstract The INhibitor of Growth (ING) family of tumor suppressors regulates the transcriptional state of chromatin by recruiting remodeling complexes to sites with histone H3 trimethylated at lysine 4 (H3K4me3). This modification is recognized by the plant homeodomain (PHD) present at the C-terminus of the five ING proteins. ING5 facilitates histone H3 acetylation by the HBO1 complex, and also H4 acetylation by the MOZ/ MORF complex. We show that ING5 forms homodimers through its N-terminal domain, which folds independently into an elongated coiled-coil structure. The central region of ING5, which contains the nuclear localization sequence, is flexible and disordered, but it binds dsDNA with micromolar affinity. NMR analysis of the full-length protein reveals that the two PHD fingers of the dimer are chemically equivalent and independent of the rest of the molecule, and they bind H3K4me3 in the same way as the isolated PHD. -

Modes of Interaction of KMT2 Histone H3 Lysine 4 Methyltransferase/COMPASS Complexes with Chromatin
cells Review Modes of Interaction of KMT2 Histone H3 Lysine 4 Methyltransferase/COMPASS Complexes with Chromatin Agnieszka Bochy ´nska,Juliane Lüscher-Firzlaff and Bernhard Lüscher * ID Institute of Biochemistry and Molecular Biology, Medical School, RWTH Aachen University, Pauwelsstrasse 30, 52057 Aachen, Germany; [email protected] (A.B.); jluescher-fi[email protected] (J.L.-F.) * Correspondence: [email protected]; Tel.: +49-241-8088850; Fax: +49-241-8082427 Received: 18 January 2018; Accepted: 27 February 2018; Published: 2 March 2018 Abstract: Regulation of gene expression is achieved by sequence-specific transcriptional regulators, which convey the information that is contained in the sequence of DNA into RNA polymerase activity. This is achieved by the recruitment of transcriptional co-factors. One of the consequences of co-factor recruitment is the control of specific properties of nucleosomes, the basic units of chromatin, and their protein components, the core histones. The main principles are to regulate the position and the characteristics of nucleosomes. The latter includes modulating the composition of core histones and their variants that are integrated into nucleosomes, and the post-translational modification of these histones referred to as histone marks. One of these marks is the methylation of lysine 4 of the core histone H3 (H3K4). While mono-methylation of H3K4 (H3K4me1) is located preferentially at active enhancers, tri-methylation (H3K4me3) is a mark found at open and potentially active promoters. Thus, H3K4 methylation is typically associated with gene transcription. The class 2 lysine methyltransferases (KMTs) are the main enzymes that methylate H3K4. KMT2 enzymes function in complexes that contain a necessary core complex composed of WDR5, RBBP5, ASH2L, and DPY30, the so-called WRAD complex. -

Open Data for Differential Network Analysis in Glioma
International Journal of Molecular Sciences Article Open Data for Differential Network Analysis in Glioma , Claire Jean-Quartier * y , Fleur Jeanquartier y and Andreas Holzinger Holzinger Group HCI-KDD, Institute for Medical Informatics, Statistics and Documentation, Medical University Graz, Auenbruggerplatz 2/V, 8036 Graz, Austria; [email protected] (F.J.); [email protected] (A.H.) * Correspondence: [email protected] These authors contributed equally to this work. y Received: 27 October 2019; Accepted: 3 January 2020; Published: 15 January 2020 Abstract: The complexity of cancer diseases demands bioinformatic techniques and translational research based on big data and personalized medicine. Open data enables researchers to accelerate cancer studies, save resources and foster collaboration. Several tools and programming approaches are available for analyzing data, including annotation, clustering, comparison and extrapolation, merging, enrichment, functional association and statistics. We exploit openly available data via cancer gene expression analysis, we apply refinement as well as enrichment analysis via gene ontology and conclude with graph-based visualization of involved protein interaction networks as a basis for signaling. The different databases allowed for the construction of huge networks or specified ones consisting of high-confidence interactions only. Several genes associated to glioma were isolated via a network analysis from top hub nodes as well as from an outlier analysis. The latter approach highlights a mitogen-activated protein kinase next to a member of histondeacetylases and a protein phosphatase as genes uncommonly associated with glioma. Cluster analysis from top hub nodes lists several identified glioma-associated gene products to function within protein complexes, including epidermal growth factors as well as cell cycle proteins or RAS proto-oncogenes. -
Download Validation Data
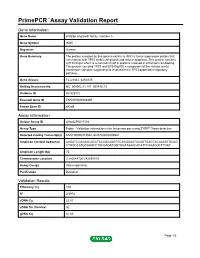
PrimePCR™Assay Validation Report Gene Information Gene Name inhibitor of growth family, member 5 Gene Symbol ING5 Organism Human Gene Summary The protein encoded by this gene is similar to ING1 a tumor suppressor protein that can interact with TP53 inhibit cell growth and induce apoptosis. This protein contains a PHD-finger which is a common motif in proteins involved in chromatin remodeling. This protein can bind TP53 and EP300/p300 a component of the histone acetyl transferase complex suggesting its involvement in TP53-dependent regulatory pathway. Gene Aliases FLJ23842, p28ING5 RefSeq Accession No. NC_000002.11, NT_005416.13 UniGene ID Hs.529172 Ensembl Gene ID ENSG00000168395 Entrez Gene ID 84289 Assay Information Unique Assay ID qHsaCIP0031594 Assay Type Probe - Validation information is for the primer pair using SYBR® Green detection Detected Coding Transcript(s) ENST00000313552, ENST00000406941 Amplicon Context Sequence AAGATCCAGAACGCCTACAGCAAGTGCAAGGAATACAGTGACGACAAAGTGCAG CTGGCCATGCAGACCTACGAGATGGTGGATAAACACATTCGAAGGCTTGAT Amplicon Length (bp) 75 Chromosome Location 2:242648720-242650818 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 100 R2 0.9978 cDNA Cq 22.07 cDNA Tm (Celsius) 82 gDNA Cq 41.86 Page 1/5 PrimePCR™Assay Validation Report Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report ING5, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak -

Ing1 Functions in DNA Demethylation by Directing Gadd45a to H3k4me3
Downloaded from genesdev.cshlp.org on September 30, 2021 - Published by Cold Spring Harbor Laboratory Press Ing1 functions in DNA demethylation by directing Gadd45a to H3K4me3 Andrea Scha¨fer,1,3 Emil Karaulanov,1 Ulrike Stapf,1 Gabi Do¨ derlein,2 and Christof Niehrs1,2,3 1Institute of Molecular Biology, 55128 Mainz, Germany; 2Division of Molecular Embryology, DKFZ-ZMBH Alliance, German Cancer Research Center, 69120 Heidelberg, Germany Active DNA demethylation regulates epigenetic gene activation in numerous processes, but how the target site specificity of DNA demethylation is determined and what factors are involved are still poorly understood. Here we show that the tumor suppressor inhibitor of growth protein 1 (Ing1) is required for targeting active DNA demethylation. Ing1 functions by recruiting the regulator of DNA demethylation growth arrest and DNA damage protein 45a (Gadd45a) to histone H3 trimethylated at Lys 4 (H3K4me3). We show that reduced H3K4 methylation impairs recruitment of Gadd45a/Ing1 and gene-specific DNA demethylation. Our results indicate that histone methylation directs DNA demethylation. [Keywords: p33Ing1b; Gadd45a; Ing1; DNA demethylation; H3K4me3; chromatin] Supplemental material is available for this article. Received January 9, 2012; revised version accepted December 26, 2012. DNA methylation at 5-methylcytosine (5mC) of CpGs 1994). Gadd45 proteins are multifunctional and regulate is a common epigenetic mark in metazoa, which plays a range of cellular processes, including DNA repair, pro- important roles in regulating gene expression, genomic liferation, apoptosis, and differentiation (Fornace et al. imprinting, X-chromosome inactivation, genomic insta- 1988; Kastan et al. 1992; Smith et al. 1996; Carrier et al. bility, embryonic development, and cancer (Costello and 1999; Harkin et al. -

Structure-Function Relationships of Tumor Suppressor ING5
Structure-function relationships of tumor suppressor ING5 Georgina Ormaza Hernández Tesis Doctoral, 2017 STRUCTURE-FUNCTION RELATIONSHIPS OF TUMOR SUPPRESSOR ING5 Departamento de Bioquímica y Biología Molecular Facultad de Ciencia y Tecnología Universidad del País Vasco/Euskal Herriko Unibertsitatea Memoria presentada para optar al grado de Doctor por Georgina Ormaza Hernández Tesis dirigida por Francisco J. Blanco Realizada en CIC bioGUNE (c)2017 GEORGINA ORMAZA HERNANDEZ AGRADECIMIENTOS Por fin ha llegado el momento… Parece que fue ayer cuando empecé mi andadura por este camino lleno de duros esfuerzos y quebraderos de cabeza, pero también de alegrías, compañerismo y superación. Es con esto último con lo que me quedo y lo que me llevo y, es por ello, que no podía irme sin agradecerlo. Quería agradecer a Paco la oportunidad de realizar mi tesis con él en el CICbioGUNE. Muchas gracias, Paco, por enseñarme a aprender, a desarrollar la visión crítica, por confiar en mí y en mi trabajo y, sobretodo, por esa cercanía y esa forma de hacer siempre bien las cosas. Por supuesto, tengo que agradecer el no haber trabajado sola. Detrás de todo mi trabajo, se encuentran grandes compañeros y amigos con los que he tenido el placer de compartir poyata. Gracias a Neka y Maider, por enseñarme todo lo que sé y ser mis “solucionadoras de vida” en momentos extremos, a Jhon Alex por su legado ING5, a Alain por su ayuda en todos los campos, pero sobretodo, en RMN y en el procesado “rápido” gracias a sus scripts maravillosos, y a Amaia la última en incorporarse, el aire fresco del grupo que enseguida se ha convertido en una buena amiga, tanto dentro como fuera del Biogune. -

Yeast Genes Illuminate Human Cancer Gene Functions
Oncogene (2007) 26, 5373–5384 & 2007 Nature Publishing Group All rights reserved 0950-9232/07 $30.00 www.nature.com/onc REVIEW MYST opportunities for growth control: yeast genes illuminate human cancer gene functions A Lafon, CS Chang, EM Scott, SJ Jacobson and L Pillus Section of Molecular Biology, Division of Biological Sciences, UCSD Moores Cancer Center, University of California, San Diego, La Jolla, CA, USA The MYST family of histone acetyltransferases (HATs) genes were identified with potential roles in chromatin- was initially defined by human genes with disease mediated gene control, even modest degrees of similarity connections and by yeast genes identified for their role to the acetyl-CoA-binding regions were viewed with in epigenetic transcriptional silencing. Since then, many special interest as candidate HATs. new MYST genes have been discovered through genetic Such interest was particularly the case when mutants and genomic approaches. Characterization of the com- of SAS2 (something about silencing) were discovered as plexes through which MYST proteins act, regions of enhancers of mutations in the epigenetic transcriptional the genome to which they are targeted and biological silencer factor Sir1 (Reifsnyder et al., 1996) and consequences when they are disrupted, all deepen the suppressors of defects in cis-regulatory sequences for connections of MYST proteins to development, growth silent chromatin (Reifsnyder et al., 1996; Ehrenhofer- control and human cancers. Many of the insights into Murray et al., 1997). The observation that SAS2 and the MYST family function have come from studies in model closely related SAS3 yeast genes shared similarity to organisms. Herein, we review functions of two of the acetyl-CoA-binding domains, and even more significant founding MYST genes, yeast SAS2 and SAS3, and the similarity to two human genes, MOZ (Borrow et al., essential yeast MYST ESA1. -

The 2Q37-Deletion Syndrome: an Update of the Clinical Spectrum Including Overweight, Brachydactyly and Behavioural Features in 14 New Patients
European Journal of Human Genetics (2013) 21, 602–612 & 2013 Macmillan Publishers Limited All rights reserved 1018-4813/13 www.nature.com/ejhg ARTICLE The 2q37-deletion syndrome: an update of the clinical spectrum including overweight, brachydactyly and behavioural features in 14 new patients Camille Leroy1,2,3, Emilie Landais1,2,4, Sylvain Briault5, Albert David6, Olivier Tassy7, Nicolas Gruchy8, Bruno Delobel9, Marie-Jose´ Gre´goire10, Bruno Leheup3,11, Laurence Taine12, Didier Lacombe12, Marie-Ange Delrue12, Annick Toutain13, Agathe Paubel13, Francine Mugneret14, Christel Thauvin-Robinet3,15, Ste´phanie Arpin13, Cedric Le Caignec6, Philippe Jonveaux3,10, Myle`ne Beri10, Nathalie Leporrier8, Jacques Motte16, Caroline Fiquet17,18, Olivier Brichet16, Monique Mozelle-Nivoix1,3, Pascal Sabouraud16, Nathalie Golovkine19, Nathalie Bednarek20, Dominique Gaillard1,2,3 and Martine Doco-Fenzy*,1,2,3,18 The 2q37 locus is one of the most commonly deleted subtelomeric regions. Such a deletion has been identified in 4100 patients by telomeric fluorescence in situ hybridization (FISH) analysis and, less frequently, by array-based comparative genomic hybridization (array-CGH). A recognizable ‘2q37-deletion syndrome’ or Albright’s hereditary osteodystrophy-like syndrome has been previously described. To better map the deletion and further refine this deletional syndrome, we formed a collaboration with the Association of French Language Cytogeneticists to collect 14 new intellectually deficient patients with a distal or interstitial 2q37 deletion characterized by FISH and array-CGH. Patients exhibited facial dysmorphism (13/14) and brachydactyly (10/14), associated with behavioural problems, autism or autism spectrum disorders of varying severity and overweight or obesity. The deletions in these 14 new patients measured from 2.6 to 8.8 Mb. -

Inhibitor of Growth (ING) Family: an Emerging Molecular Target for Cancer Therapy
Esra Gunduz et.al.: ING Tumor Supressor Family Journal of Hard Tissue Biology 17[1] (2008) p1-10 © 2008 The Hard Tissue Biology Network Printed in Japan, All rights reserved. CODEN-JHTBFF, ISSN 1341-7649 Review Inhibitor of Growth (ING) Family: An Emerging Molecular Target for Cancer Therapy Esra Gunduz1,2), Mehmet Gunduz1,3), Levent Bekir Beder3), Ryo Tamamura1), Hitoshi Nagatsuka1) and Noriyuki Nagai1) 1)Department of 1Oral Pathology and Medicine, Dentistry and Pharmaceutical Sciences, Okayama University, 2-5-1 Shikatacho, Okayama, 700-8525, Japan 2)Molecular Genetics, Graduate School of Medicine, Dentistry and Pharmaceutical Sciences, Okayama University, 2-5-1 Shikatacho, Okayama, 700-8525, Japan 3)Department of Otolaryngology Head and Neck Surgery, Wakayama Medical University, 811-1, Kimiidera, Wakayama, 641-8509, Japan (Accepted for publication, March 20, 2008) Abstract: ING1 gene, the founding member of the ING tumor suppressor family, was originally identified through subtractive hybridization between normal mammary epithelial cells and breast cancer cell lines, and subsequent in vivo selection of genetic suppressor element that displayed oncogenic features. Soon after identification of ING1, four additional members of the ING family (ING2-5) were cloned and all the gene products contain a highly conserved plant homeodomain (PHD) finger motif in the carboxy (C)-terminal end, that plays important role for their function. Furthermore, ING family members contain nuclear localization signals and N-terminal sequences important in the interaction with histone acetyltransferase (HAT) and histone deacetyltransferase (HDAC) that regulate gene promoter activity within chromatin. Although exact functions of ING family genes have not been clarified, the gene products are involved in transcriptional regulation, apoptosis, cell cyle, angiogenesis and DNA repair through p53-dependent and –independent pathways. -

Histone Target Selection Within Chromatin: an Exemplary Case of Teamwork
Downloaded from genesdev.cshlp.org on September 30, 2021 - Published by Cold Spring Harbor Laboratory Press REVIEW Histone target selection within chromatin: an exemplary case of teamwork Marie-Eve Lalonde, Xue Cheng, and Jacques Coˆ te´ 1 St-Patrick Research Group in Basic Oncology, Laval University Cancer Research Center, Centre de Recherche du CHU de Que´bec-Axe Oncologie, Hoˆ tel-Dieu de Que´bec, Quebec City, Quebec G1R 2J6, Canada Histone modifiers like acetyltransferases, methyltrans- influence chromatin dynamics (Tan et al. 2011; Zentner ferases, and demethylases are critical regulators of most and Henikoff 2013). DNA-based nuclear processes, de facto controlling cell Histone PTMs can be deposited on or removed from cycle progression and cell fate. These enzymes perform chromatin by different enzymes. These ‘‘writers’’ and very precise post-translational modifications on spe- ‘‘erasers’’ of histone marks include different kinases and cific histone residues, which in turn are recognized phosphatases, ubiquitin ligases and deubiquitinases, by different effector modules/proteins. We now have a lysine/arginine methyltransferases, and demethylases. better understanding of how these enzymes exhibit such To date, numerous lysine methyltransferases have been specificity. As they often reside in multisubunit com- identified, and each can either mono-, di-, or trimeth- plexes, they use associated factors to target their sub- ylate and may act on only a specific methylation status strates within chromatin structure and select specific of a residue. Conversely, there are two classes of histone histone mark-bearing nucleosomes. In this review, we lysine demethylases identified: the LSD1 family (found cover the current understanding of how histone modi- only in humans/mammals) and the Jumonjis (Greer and fiers select their histone targets. -

The Roles of ING5 in Gliomas: a Good Marker for Tumorigenesis and a Potential Target for Gene Therapy
www.impactjournals.com/oncotarget/ Oncotarget, 2017, Vol. 8, (No. 34), pp: 56558-56568 Research Paper The roles of ING5 in gliomas: a good marker for tumorigenesis and a potential target for gene therapy Shuang Zhao1, Zhi-Juan Zhao2, Hao-Yu He1, Ji-Cheng Wu1, Xiao-Qing Ding1, Lei Yang1, Ning Jia2, Zhi-Jie Li1 and Hua-Chuan Zheng1 1Department of Experimental Oncology and Animal Center, Shengjing Hospital of China Medical University, Shenyang 110004, China 2The First Affiliated Hospital of Jinzhou Medical University, Jinzhou 121001, China Correspondence to: Hua-Chuan Zheng, email: [email protected] Keywords: glioma, ING5, tumorigenesis, chemotherapy, gene therapy Received: December 16, 2016 Accepted: April 28, 2017 Published: May 11, 2017 Copyright: Zhao et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License 3.0 (CC BY 3.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT To elucidate the anti-tumor effects and molecular mechanisms of ING5 on glioma cells, we overexpressed it in U87 cells, and examined the phenotypes and their relevant molecules. It was found that ING5 overexpression suppressed proliferation, energy metabolism, migration, invasion, and induced G2/M arrest, apoptosis, dedifferentiation, senescence, mesenchymal- epithelial transition and chemoresistance to cisplatin, MG132, paclitaxel and SAHA in U87 cells. There appeared a lower expression of N- cadherin, Twist, Slug, Zeb1, Zeb2, Snail, Ac-H3, Ac-H4, Cdc2, Cdk4 and XIAP, but a higher expression of Claudin 1, Histones 3 and 4, p21, p53, Bax, β-catenin, PI3K, Akt, and p-Akt in ING5 transfectants.