Echohealth and the Identification of New Viruses
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

2020 Taxonomic Update for Phylum Negarnaviricota (Riboviria: Orthornavirae), Including the Large Orders Bunyavirales and Mononegavirales
Archives of Virology https://doi.org/10.1007/s00705-020-04731-2 VIROLOGY DIVISION NEWS 2020 taxonomic update for phylum Negarnaviricota (Riboviria: Orthornavirae), including the large orders Bunyavirales and Mononegavirales Jens H. Kuhn1 · Scott Adkins2 · Daniela Alioto3 · Sergey V. Alkhovsky4 · Gaya K. Amarasinghe5 · Simon J. Anthony6,7 · Tatjana Avšič‑Županc8 · María A. Ayllón9,10 · Justin Bahl11 · Anne Balkema‑Buschmann12 · Matthew J. Ballinger13 · Tomáš Bartonička14 · Christopher Basler15 · Sina Bavari16 · Martin Beer17 · Dennis A. Bente18 · Éric Bergeron19 · Brian H. Bird20 · Carol Blair21 · Kim R. Blasdell22 · Steven B. Bradfute23 · Rachel Breyta24 · Thomas Briese25 · Paul A. Brown26 · Ursula J. Buchholz27 · Michael J. Buchmeier28 · Alexander Bukreyev18,29 · Felicity Burt30 · Nihal Buzkan31 · Charles H. Calisher32 · Mengji Cao33,34 · Inmaculada Casas35 · John Chamberlain36 · Kartik Chandran37 · Rémi N. Charrel38 · Biao Chen39 · Michela Chiumenti40 · Il‑Ryong Choi41 · J. Christopher S. Clegg42 · Ian Crozier43 · John V. da Graça44 · Elena Dal Bó45 · Alberto M. R. Dávila46 · Juan Carlos de la Torre47 · Xavier de Lamballerie38 · Rik L. de Swart48 · Patrick L. Di Bello49 · Nicholas Di Paola50 · Francesco Di Serio40 · Ralf G. Dietzgen51 · Michele Digiaro52 · Valerian V. Dolja53 · Olga Dolnik54 · Michael A. Drebot55 · Jan Felix Drexler56 · Ralf Dürrwald57 · Lucie Dufkova58 · William G. Dundon59 · W. Paul Duprex60 · John M. Dye50 · Andrew J. Easton61 · Hideki Ebihara62 · Toufc Elbeaino63 · Koray Ergünay64 · Jorlan Fernandes195 · Anthony R. Fooks65 · Pierre B. H. Formenty66 · Leonie F. Forth17 · Ron A. M. Fouchier48 · Juliana Freitas‑Astúa67 · Selma Gago‑Zachert68,69 · George Fú Gāo70 · María Laura García71 · Adolfo García‑Sastre72 · Aura R. Garrison50 · Aiah Gbakima73 · Tracey Goldstein74 · Jean‑Paul J. Gonzalez75,76 · Anthony Grifths77 · Martin H. Groschup12 · Stephan Günther78 · Alexandro Guterres195 · Roy A. -

Characterizing and Evaluating the Zoonotic Potential of Novel Viruses Discovered in Vampire Bats
viruses Article Characterizing and Evaluating the Zoonotic Potential of Novel Viruses Discovered in Vampire Bats Laura M. Bergner 1,2,* , Nardus Mollentze 1,2 , Richard J. Orton 2 , Carlos Tello 3,4, Alice Broos 2, Roman Biek 1 and Daniel G. Streicker 1,2 1 Institute of Biodiversity, Animal Health and Comparative Medicine, College of Medical, Veterinary and Life Sciences, University of Glasgow, Glasgow G12 8QQ, UK; [email protected] (N.M.); [email protected] (R.B.); [email protected] (D.G.S.) 2 MRC–University of Glasgow Centre for Virus Research, Glasgow G61 1QH, UK; [email protected] (R.J.O.); [email protected] (A.B.) 3 Association for the Conservation and Development of Natural Resources, Lima 15037, Peru; [email protected] 4 Yunkawasi, Lima 15049, Peru * Correspondence: [email protected] Abstract: The contemporary surge in metagenomic sequencing has transformed knowledge of viral diversity in wildlife. However, evaluating which newly discovered viruses pose sufficient risk of infecting humans to merit detailed laboratory characterization and surveillance remains largely speculative. Machine learning algorithms have been developed to address this imbalance by ranking the relative likelihood of human infection based on viral genome sequences, but are not yet routinely Citation: Bergner, L.M.; Mollentze, applied to viruses at the time of their discovery. Here, we characterized viral genomes detected N.; Orton, R.J.; Tello, C.; Broos, A.; through metagenomic sequencing of feces and saliva from common vampire bats (Desmodus rotundus) Biek, R.; Streicker, D.G. and used these data as a case study in evaluating zoonotic potential using molecular sequencing Characterizing and Evaluating the data. -

Viral Zoonotic Encephalitis: Australian Bat Lyssavirus and Hendra
Viral zoonotic encephalitis: Australian Bat Lyssavirus and Hendra Bev Paterson Hunter© by Medicalauthor Research Institute University of Newcastle Australia ESCMID OnlineEmail: [email protected] Library Encephalitis in Australia Causes substantial morbidity and mortality Herpes simplex virus is the most commonly identified causative pathogen 70% of adult encephalitis hospitalisations no pathogen identified 57% of deaths no pathogen identified (Reference: Huppatz et al. CDI,© 2009; by Huppatzauthor et al. EID, 2009) ESCMID Online Lecture Library Viral zoonotic encphalitis Several recently emerged or resurging pathogens are known to cause an encephalitis syndrome Vectorborne and transmitted flaviviruses – MVEV, WNEV-KUN and JEV © by author Bat-borne viruses – Australian Bat Lyssavirus (ABLV) and Hendra virus ESCMID Online Lecture Library Impact of climatic conditions © by author ESCMID Online Lecture Library Australian Bat Lyssavirus Australia has no endemic rabies ABLV is a member of the family Rhabdoviridae, genus Lyssavirus ABLV is very closely related to rabies (genotype 7 of the Lyssavirus genus) Reservoir is bats © by author Two deaths from ABLV ESCMID Online Lecture Library Human exposure © by author ESCMID Online Lecture Library Epidemiology of human disease Two fatal human cases – coastal Qld – encephalitis indistinguishable from classic rabies 1996 – 39yr female, a few weeks after scratches from bat 1998 – 37yr female, 27 months after bite from bat © by author References (Samaratunga -

Everything You Always Wanted to Know About Rabies Virus ♣♣♣♣♣♣♣♣♣♣♣♣♣♣♣♣♣♣♣♣♣ (But Were Afraid to Ask) Benjamin M
ANNUAL REVIEWS Further Click here to view this article's online features: t%PXOMPBEmHVSFTBT115TMJEFT t/BWJHBUFMJOLFESFGFSFODFT t%PXOMPBEDJUBUJPOT Everything You Always Wanted t&YQMPSFSFMBUFEBSUJDMFT t4FBSDILFZXPSET to Know About Rabies Virus (But Were Afraid to Ask) Benjamin M. Davis,1 Glenn F. Rall,2 and Matthias J. Schnell1,2,3 1Department of Microbiology and Immunology and 3Jefferson Vaccine Center, Sidney Kimmel Medical College, Thomas Jefferson University, Philadelphia, Pennsylvania, 19107; email: [email protected] 2Fox Chase Cancer Center, Philadelphia, Pennsylvania 19111 Annu. Rev. Virol. 2015. 2:451–71 Keywords First published online as a Review in Advance on rabies virus, lyssaviruses, neurotropic virus, neuroinvasive virus, viral June 24, 2015 transport The Annual Review of Virology is online at virology.annualreviews.org Abstract This article’s doi: The cultural impact of rabies, the fatal neurological disease caused by in- 10.1146/annurev-virology-100114-055157 fection with rabies virus, registers throughout recorded history. Although Copyright c 2015 by Annual Reviews. ⃝ rabies has been the subject of large-scale public health interventions, chiefly All rights reserved through vaccination efforts, the disease continues to take the lives of about 40,000–70,000 people per year, roughly 40% of whom are children. Most of Access provided by Thomas Jefferson University on 11/13/15. For personal use only. Annual Review of Virology 2015.2:451-471. Downloaded from www.annualreviews.org these deaths occur in resource-poor countries, where lack of infrastructure prevents timely reporting and postexposure prophylaxis and the ubiquity of domestic and wild animal hosts makes eradication unlikely. Moreover, al- though the disease is rarer than other human infections such as influenza, the prognosis following a bite from a rabid animal is poor: There is cur- rently no effective treatment that will save the life of a symptomatic rabies patient. -

Bat Rabies and Other Lyssavirus Infections
Prepared by the USGS National Wildlife Health Center Bat Rabies and Other Lyssavirus Infections Circular 1329 U.S. Department of the Interior U.S. Geological Survey Front cover photo (D.G. Constantine) A Townsend’s big-eared bat. Bat Rabies and Other Lyssavirus Infections By Denny G. Constantine Edited by David S. Blehert Circular 1329 U.S. Department of the Interior U.S. Geological Survey U.S. Department of the Interior KEN SALAZAR, Secretary U.S. Geological Survey Suzette M. Kimball, Acting Director U.S. Geological Survey, Reston, Virginia: 2009 For more information on the USGS—the Federal source for science about the Earth, its natural and living resources, natural hazards, and the environment, visit http://www.usgs.gov or call 1–888–ASK–USGS For an overview of USGS information products, including maps, imagery, and publications, visit http://www.usgs.gov/pubprod To order this and other USGS information products, visit http://store.usgs.gov Any use of trade, product, or firm names is for descriptive purposes only and does not imply endorsement by the U.S. Government. Although this report is in the public domain, permission must be secured from the individual copyright owners to reproduce any copyrighted materials contained within this report. Suggested citation: Constantine, D.G., 2009, Bat rabies and other lyssavirus infections: Reston, Va., U.S. Geological Survey Circular 1329, 68 p. Library of Congress Cataloging-in-Publication Data Constantine, Denny G., 1925– Bat rabies and other lyssavirus infections / by Denny G. Constantine. p. cm. - - (Geological circular ; 1329) ISBN 978–1–4113–2259–2 1. -

Finfish Diseases
SECTION 2 - FINFISH DISEASES Basic Anatomy of a Typical Bony Fish 48 SECTION 2 - FINFISH DISEASES F. 1 GENERAL TECHNIQUES 50 F.1.1 Gross Observations 50 F.1.1.1 Behaviour 50 F.1.1.2 Surface Observations 50 F.1.1.2.1 Skin and Fins 50 F.1.1.2.2 Gills 51 F.1.1.2.3 Body 52 F.1.1.3 Internal Observations 52 F.1.1.3.1 Body Cavity and Muscle 52 F.1.1.3.2 Organs 52 F.1.2 Environmental Parameters 53 F.1.3 General Procedures 53 F.1.3.1 Pre-Collection Preparation 53 F.1.3.2 Background Information 54 F.1.3.3 Sample Collection for Health Surveillance 54 F.1.3.4 Sample Collection for Disease Diagnosis 54 F.1.3.5 Live Specimen Collection for Shipping 55 F.1.3.6 Dead or Tissue Specimen Collection for Shipping 55 F.1.3.7 Preservation of Tissue Samples 56 F.1.3.8 Shipping Preserved Samples 56 F.1.4 Record-Keeping 57 F.1.4.1 Gross Observations 57 F.1.4.2 Environmental Observations 57 F.1.4.3 Stocking Records 57 F.1.5 References 57 VIRAL DISEASES OF FINFISH F.2 Epizootic Haematopoietic Necrosis (EHN) 59 F.3 Infectious Haematopoietic Necrosis (IHN) 62 F.4 Oncorhynchus masou Virus (OMV) 65 F.5 Infectious Pancreatic Necrosis (IPN) 68 F.6 Viral Encephalopathy and Retinopathy (VER) 72 F.7 Spring Viraemia of Carp (SVC) 76 F.8 Viral Haemorrhagic Septicaemia (VHS) 79 F.9 Lymphocystis 82 BACTERIAL DISEASE OF FINFISH F.10 Bacterial Kidney Disease (BKD) 86 FUNGUS ASSOCIATED DISEASE FINFISH F.11 Epizootic Ulcerative Syndrome (EUS) 90 ANNEXES F.AI OIE Reference Laboratories for Finfish Diseases 95 F.AII List of Regional Resource Experts for Finfish 98 Diseases in Asia-Pacific F.AIII List of Useful Diagnostic Manuals/Guides to 105 Finfish Diseases in Asia-Pacific 49 F.1 GENERAL TECHNIQUES infectious disease agent and should be sampled immediately. -

Risk Groups: Viruses (C) 1988, American Biological Safety Association
Rev.: 1.0 Risk Groups: Viruses (c) 1988, American Biological Safety Association BL RG RG RG RG RG LCDC-96 Belgium-97 ID Name Viral group Comments BMBL-93 CDC NIH rDNA-97 EU-96 Australia-95 HP AP (Canada) Annex VIII Flaviviridae/ Flavivirus (Grp 2 Absettarov, TBE 4 4 4 implied 3 3 4 + B Arbovirus) Acute haemorrhagic taxonomy 2, Enterovirus 3 conjunctivitis virus Picornaviridae 2 + different 70 (AHC) Adenovirus 4 Adenoviridae 2 2 (incl animal) 2 2 + (human,all types) 5 Aino X-Arboviruses 6 Akabane X-Arboviruses 7 Alastrim Poxviridae Restricted 4 4, Foot-and- 8 Aphthovirus Picornaviridae 2 mouth disease + viruses 9 Araguari X-Arboviruses (feces of children 10 Astroviridae Astroviridae 2 2 + + and lambs) Avian leukosis virus 11 Viral vector/Animal retrovirus 1 3 (wild strain) + (ALV) 3, (Rous 12 Avian sarcoma virus Viral vector/Animal retrovirus 1 sarcoma virus, + RSV wild strain) 13 Baculovirus Viral vector/Animal virus 1 + Togaviridae/ Alphavirus (Grp 14 Barmah Forest 2 A Arbovirus) 15 Batama X-Arboviruses 16 Batken X-Arboviruses Togaviridae/ Alphavirus (Grp 17 Bebaru virus 2 2 2 2 + A Arbovirus) 18 Bhanja X-Arboviruses 19 Bimbo X-Arboviruses Blood-borne hepatitis 20 viruses not yet Unclassified viruses 2 implied 2 implied 3 (**)D 3 + identified 21 Bluetongue X-Arboviruses 22 Bobaya X-Arboviruses 23 Bobia X-Arboviruses Bovine 24 immunodeficiency Viral vector/Animal retrovirus 3 (wild strain) + virus (BIV) 3, Bovine Bovine leukemia 25 Viral vector/Animal retrovirus 1 lymphosarcoma + virus (BLV) virus wild strain Bovine papilloma Papovavirus/ -
Supporting Information
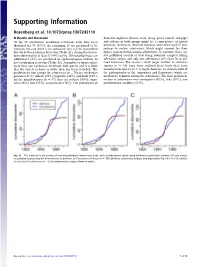
Supporting Information Rosenberg et al. 10.1073/pnas.1307243110 SI Results and Discussion domestic ungulates (horses, cows, sheep, goats, camels, and pigs) Of the 83 arboviruses, nonhuman vertebrate hosts have been and rodents in both groups might be a consequence of spatial identified for 70 (84%); the remaining 13 are presumed to be proximity to humans. Sentinel monkeys were often used in pro- zoonoses because there is no indication they can be transmitted cedures to isolate arboviruses, which might account for their directly between humans by vectors (Table S1). Animal hosts have higher representation among arboviruses. In contrast, there are been identified for at least 57 (44%) of the 130 nonarboviruses; an few published records of bats being routinely sampled during additional 5 (8%) are presumed on epidemiological evidence to arbovirus studies, and only two arboviruses (3%) have been iso- have nonhuman reservoirs (Table S1). A number of viruses infect lated from bats. The reason a much larger number of arbovirus more than one nonhuman vertebrate host species and it is likely species (n = 16) have been isolated from birds than have that the variety of hosts is wider than has been recorded. The nonarbovirus species (n = 1) might, however, be characteristic of predominant host groups for arboviruses (n = 70) are nonhuman the pathogenicity of the togaviruses and flaviviruses, which are primates (31%), rodents (29%), ungulates (26%), and birds (23%); much more common among the arboviruses. The most prominent for the nonarboviruses (n = 57), they are rodents (30%), ungu- vectors of arboviruses were mosquitoes (67%), ticks (19%), and lates (26%), bats (23%), and primates (16%). -

Arenaviridae Astroviridae Filoviridae Flaviviridae Hantaviridae
Hantaviridae 0.7 Filoviridae 0.6 Picornaviridae 0.3 Wenling red spikefish hantavirus Rhinovirus C Ahab virus * Possum enterovirus * Aronnax virus * * Wenling minipizza batfish hantavirus Wenling filefish filovirus Norway rat hunnivirus * Wenling yellow goosefish hantavirus Starbuck virus * * Porcine teschovirus European mole nova virus Human Marburg marburgvirus Mosavirus Asturias virus * * * Tortoise picornavirus Egyptian fruit bat Marburg marburgvirus Banded bullfrog picornavirus * Spanish mole uluguru virus Human Sudan ebolavirus * Black spectacled toad picornavirus * Kilimanjaro virus * * * Crab-eating macaque reston ebolavirus Equine rhinitis A virus Imjin virus * Foot and mouth disease virus Dode virus * Angolan free-tailed bat bombali ebolavirus * * Human cosavirus E Seoul orthohantavirus Little free-tailed bat bombali ebolavirus * African bat icavirus A Tigray hantavirus Human Zaire ebolavirus * Saffold virus * Human choclo virus *Little collared fruit bat ebolavirus Peleg virus * Eastern red scorpionfish picornavirus * Reed vole hantavirus Human bundibugyo ebolavirus * * Isla vista hantavirus * Seal picornavirus Human Tai forest ebolavirus Chicken orivirus Paramyxoviridae 0.4 * Duck picornavirus Hepadnaviridae 0.4 Bildad virus Ned virus Tiger rockfish hepatitis B virus Western African lungfish picornavirus * Pacific spadenose shark paramyxovirus * European eel hepatitis B virus Bluegill picornavirus Nemo virus * Carp picornavirus * African cichlid hepatitis B virus Triplecross lizardfish paramyxovirus * * Fathead minnow picornavirus -

Genomics and Structure/Function Studies of Rhabdoviridae Proteins Involved in Replication and Transcription
Genomics and structure/function studies of Rhabdoviridae proteins involved in replication and transcription. R. Assenberg, O Delmas, B Morin, C Graham, X de Lamballerie, C Laubert, B Coutard, J Grimes, J Neyts, R J Owens, et al. To cite this version: R. Assenberg, O Delmas, B Morin, C Graham, X de Lamballerie, et al.. Genomics and struc- ture/function studies of Rhabdoviridae proteins involved in replication and transcription.. Antivi- ral Research, Elsevier Masson, 2010, 87 (2), pp.149-61. 10.1016/j.antiviral.2010.02.322. pasteur- 01492926 HAL Id: pasteur-01492926 https://hal-pasteur.archives-ouvertes.fr/pasteur-01492926 Submitted on 21 Apr 2017 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Distributed under a Creative Commons Attribution - NonCommercial - ShareAlike| 4.0 International License *Manuscript Genomics and structure/function studies of Rhabdoviridae proteins involved in replication and transcription R. Assenberg1, O. Delmas2, B. Morin3, S. C. Graham1, X. De Lamballerie4, C. Laubert5, B. Coutard3, J. M. Grimes1, J. Neyts6, R. J. Owens1, -

Module 4: Negative Strand RNA Viruses Lecture 23: Negative Strand RNA Viruses
NPTEL – Biotechnology – General Virology Module 4: Negative strand RNA viruses Lecture 23: Negative strand RNA viruses Negative strand RNA viruses belong to order Mononegavirales. The viruses in this group have similar genome organization and replication strategies and are probably diverged from a common ancestor (Filoviridae, Paramyxoviridae, Bornaviridae and Rhabdoviridae). They are often associated with emerging infection and havoc to human population (Ebola, Marburg, Nipah and Hendra). Virus contains a negative sense RNA genome which means the polarity of the genome is opposite to that of an mRNA. The negative sense RNA cannot use its genome to synthesize proteins and hence its RNA is not infectious (absence of protein synthesis). Because of the above stated property viruses in this group encode their own polymerase (RNA dependent RNA polymerase [RDRP]). Another unique property about these viruses is about its transcription, first a leader RNA is synthesized, which is followed by sequential transcription of the genes in the 3’ to 5’ order to yield individual mRNAs by a stop-start mechanism guided by the conserved gene-start and gene-end signals. 23.1 Genome features I. Linear non-segmented negative sense RNA genome II. Organization of genome- 3'-Leader-Virion core- Surface proteins-Polymerase- Trailer 5'. III. Helical nucleocapsid contains the RNA dependent RNA polymerase. IV. The leader RNA is neither capped nor polyadenylated and is not functional as mRNA. V. Replication occurs when the polymerase complex ignores the transcription stop signals at the 3’ end of each gene and a full-length positive-sense antigenome is synthesized. VI. Transcription at the gene-start site is not perfect, which leads to a gradient of mRNA abundance that decreases according to the distance from the 3’ end of the genome. -

Function of Host Protein Staufen1 in Rabies Virus Replication
viruses Article Function of Host Protein Staufen1 in Rabies Virus Replication Gaowen Liu †, Congjie Chen †, Ruixian Xu, Ming Yang, Qinqin Han, Binghui Wang, Yuzhu Song *, Xueshan Xia * and Jinyang Zhang * Research Center of Molecular Medicine of Yunnan Province, Faculty of Life Science and Technology, Kunming University of Science and Technology, Kunming 650500, China; [email protected] (G.L.); [email protected] (C.C.); [email protected] (R.X.); [email protected] (M.Y.); [email protected] (Q.H.); [email protected] (B.W.) * Correspondence: [email protected] (Y.S.); [email protected] (X.X.); [email protected] (J.Z.); Tel.: +86-871-65939528 (Y.S. & J.Z.) † These authors contributed equally to this work. Abstract: Rabies virus is a highly neurophilic negative-strand RNA virus with high lethality and remains a huge public health problem in developing countries to date. The double-stranded RNA- binding protein Staufen1 (STAU1) has multiple functions in RNA virus replication, transcription, and translation. However, its function in RABV infection and its mechanism of action are not clear. In this study, we investigated the role of host factor STAU1 in RABV infection of SH-SY-5Y cells. Immunofluorescence, TCID50 titers, confocal microscopy, quantitative real-time PCR and Western blotting were carried out to determine the molecular function and subcellular distribution of STAU1 in these cell lines. Expression of STAU1 in SH-SY-5Y cells was down-regulated by RNA interference or up-regulated by transfection of eukaryotic expression vectors. The results showed that N proficiently colocalized with STAU1 in SH-SY-5Y at 36 h post-infection, and the expression level of STAU1 was also proportional to the time of infection.