Identification of Differentially Expressed Genes Using An
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genetic Variations in the PSMA6 and PSMC6 Proteasome Genes Are Associated with Multiple Sclerosis and Response to Interferon‑Β Therapy in Latvians
EXPERIMENTAL AND THERAPEUTIC MEDICINE 21: 478, 2021 Genetic variations in the PSMA6 and PSMC6 proteasome genes are associated with multiple sclerosis and response to interferon‑β therapy in Latvians NATALIA PARAMONOVA1, JOLANTA KALNINA1, KRISTINE DOKANE1, KRISTINE DISLERE1, ILVA TRAPINA1, TATJANA SJAKSTE1 and NIKOLAJS SJAKSTE1,2 1Genomics and Bioinformatics, Institute of Biology of The University of Latvia; 2Department of Medical Biochemistry of The University of Latvia, LV‑1004 Riga, Latvia Received July 8, 2020; Accepted December 8, 2020 DOI: 10.3892/etm.2021.9909 Abstract. Several polymorphisms in genes related to the Introduction ubiquitin‑proteasome system exhibit an association with pathogenesis and prognosis of various human autoimmune Multiple sclerosis (MS) is a lifelong demyelinating disease of diseases. Our previous study reported the association the central nervous system. The clinical onset of MS tends to between multiple sclerosis (MS) and the PSMA3‑rs2348071 be between the second and fourth decade of life. Similarly to polymorphism in the Latvian population. The current study other autoimmune diseases, women are affected 3‑4 times more aimed to evaluate the PSMA6 and PSMC6 genetic variations, frequently than men (1). About 10% of MS patients experience their interaction between each other and with the rs2348071, a primary progressive MS form characterized by the progres‑ on the susceptibility to MS risk and response to therapy in sion of neurological disability from the onset. In about 90% the Latvian population. PSMA6‑rs2277460, ‑rs1048990 and of MS patients, the disease undergoes the relapse‑remitting PSMC6‑rs2295826, ‑rs2295827 were genotyped in the MS MS course (RRMS); in most of these patients, the condition case/control study and analysed in terms of genotype‑protein acquires secondary progressive course (SPMS) (2). -

Genome-Wide Association Study Identifies Loci for Arterial Stiffness
www.nature.com/scientificreports OPEN Genome-wide association study identifes loci for arterial stifness index in 127,121 UK Biobank Received: 1 February 2019 Accepted: 5 June 2019 participants Published: xx xx xxxx Kenneth Fung1, Julia Ramírez 2, Helen R. Warren2,3, Nay Aung 1, Aaron M. Lee1, Evan Tzanis2,3, Stefen E. Petersen 1,3 & Patricia B. Munroe2,3 Arterial stifness index (ASI) is a non-invasive measure of arterial stifness using infra-red fnger sensors (photoplethysmography). It is a well-suited measure for large populations as it is relatively inexpensive to perform, and data can be acquired within seconds. These features raise interest in using ASI as a tool to estimate cardiovascular disease risk as prior work demonstrates increased arterial stifness is associated with elevated systolic blood pressure, and ASI is predictive of cardiovascular disease and mortality. We conducted genome-wide association studies (GWASs) for ASI in 127,121 UK Biobank participants of European-ancestry. Our primary analyses identifed variants at four loci reaching genome-wide signifcance (P < 5 × 10−8): TEX41 (rs1006923; P = 5.3 × 10−12), FOXO1 (rs7331212; P = 2.2 × 10−11), C1orf21 (rs1930290, P = 1.1 × 10−8) and MRVI1 (rs10840457, P = 3.4 × 10−8). Gene- based testing revealed three signifcant genes, the most signifcant gene was COL4A2 (P = 1.41 × 10−8) encoding type IV collagen. Other candidate genes at associated loci were also involved in smooth muscle tone regulation. Our fndings provide new information for understanding the development of arterial stifness. Arterial stifness measures have been reported as independent markers of vascular ageing1,2, hypertension3,4, car- diovascular disease (CVD)5,6 and mortality6,7. -

Modulating Antiviral Response Via CRISPR–Cas Systems
viruses Review Immunity and Viral Infections: Modulating Antiviral Response via CRISPR–Cas Systems Sergey Brezgin 1,2,3,† , Anastasiya Kostyusheva 1,†, Ekaterina Bayurova 4 , Elena Volchkova 5, Vladimir Gegechkori 6 , Ilya Gordeychuk 4,7, Dieter Glebe 8 , Dmitry Kostyushev 1,3,*,‡ and Vladimir Chulanov 1,3,5,‡ 1 National Medical Research Center of Tuberculosis and Infectious Diseases, Ministry of Health, 127994 Moscow, Russia; [email protected] (S.B.); [email protected] (A.K.); [email protected] (V.C.) 2 Institute of Immunology, Federal Medical Biological Agency, 115522 Moscow, Russia 3 Scientific Center for Genetics and Life Sciences, Division of Biotechnology, Sirius University of Science and Technology, 354340 Sochi, Russia 4 Chumakov Federal Scientific Center for Research and Development of Immune-and-Biological Products of Russian Academy of Sciences, 108819 Moscow, Russia; [email protected] (E.B.); [email protected] (I.G.) 5 Department of Infectious Diseases, Sechenov University, 119991 Moscow, Russia; [email protected] 6 Department of Pharmaceutical and Toxicological Chemistry, Sechenov University, 119991 Moscow, Russia; [email protected] 7 Department of Organization and Technology of Immunobiological Drugs, Sechenov University, 119991 Moscow, Russia 8 National Reference Center for Hepatitis B Viruses and Hepatitis D Viruses, Institute of Medical Virology, Justus Liebig University of Giessen, 35392 Giessen, Germany; [email protected] * Correspondence: [email protected] † Co-first authors. Citation: Brezgin, S.; Kostyusheva, ‡ Co-senior authors. A.; Bayurova, E.; Volchkova, E.; Gegechkori, V.; Gordeychuk, I.; Glebe, Abstract: Viral infections cause a variety of acute and chronic human diseases, sometimes resulting D.; Kostyushev, D.; Chulanov, V. Immunity and Viral Infections: in small local outbreaks, or in some cases spreading across the globe and leading to global pandemics. -

Supplementary Materials
Supplementary materials Supplementary Table S1: MGNC compound library Ingredien Molecule Caco- Mol ID MW AlogP OB (%) BBB DL FASA- HL t Name Name 2 shengdi MOL012254 campesterol 400.8 7.63 37.58 1.34 0.98 0.7 0.21 20.2 shengdi MOL000519 coniferin 314.4 3.16 31.11 0.42 -0.2 0.3 0.27 74.6 beta- shengdi MOL000359 414.8 8.08 36.91 1.32 0.99 0.8 0.23 20.2 sitosterol pachymic shengdi MOL000289 528.9 6.54 33.63 0.1 -0.6 0.8 0 9.27 acid Poricoic acid shengdi MOL000291 484.7 5.64 30.52 -0.08 -0.9 0.8 0 8.67 B Chrysanthem shengdi MOL004492 585 8.24 38.72 0.51 -1 0.6 0.3 17.5 axanthin 20- shengdi MOL011455 Hexadecano 418.6 1.91 32.7 -0.24 -0.4 0.7 0.29 104 ylingenol huanglian MOL001454 berberine 336.4 3.45 36.86 1.24 0.57 0.8 0.19 6.57 huanglian MOL013352 Obacunone 454.6 2.68 43.29 0.01 -0.4 0.8 0.31 -13 huanglian MOL002894 berberrubine 322.4 3.2 35.74 1.07 0.17 0.7 0.24 6.46 huanglian MOL002897 epiberberine 336.4 3.45 43.09 1.17 0.4 0.8 0.19 6.1 huanglian MOL002903 (R)-Canadine 339.4 3.4 55.37 1.04 0.57 0.8 0.2 6.41 huanglian MOL002904 Berlambine 351.4 2.49 36.68 0.97 0.17 0.8 0.28 7.33 Corchorosid huanglian MOL002907 404.6 1.34 105 -0.91 -1.3 0.8 0.29 6.68 e A_qt Magnogrand huanglian MOL000622 266.4 1.18 63.71 0.02 -0.2 0.2 0.3 3.17 iolide huanglian MOL000762 Palmidin A 510.5 4.52 35.36 -0.38 -1.5 0.7 0.39 33.2 huanglian MOL000785 palmatine 352.4 3.65 64.6 1.33 0.37 0.7 0.13 2.25 huanglian MOL000098 quercetin 302.3 1.5 46.43 0.05 -0.8 0.3 0.38 14.4 huanglian MOL001458 coptisine 320.3 3.25 30.67 1.21 0.32 0.9 0.26 9.33 huanglian MOL002668 Worenine -

Plasma Cells in Vitro Generation of Long-Lived Human
Downloaded from http://www.jimmunol.org/ by guest on September 24, 2021 is online at: average * The Journal of Immunology , 32 of which you can access for free at: 2012; 189:5773-5785; Prepublished online 16 from submission to initial decision 4 weeks from acceptance to publication November 2012; doi: 10.4049/jimmunol.1103720 http://www.jimmunol.org/content/189/12/5773 In Vitro Generation of Long-lived Human Plasma Cells Mario Cocco, Sophie Stephenson, Matthew A. Care, Darren Newton, Nicholas A. Barnes, Adam Davison, Andy Rawstron, David R. Westhead, Gina M. Doody and Reuben M. Tooze J Immunol cites 65 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription http://www.jimmunol.org/content/suppl/2012/11/16/jimmunol.110372 0.DC1 This article http://www.jimmunol.org/content/189/12/5773.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2012 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 24, 2021. The Journal of Immunology In Vitro Generation of Long-lived Human Plasma Cells Mario Cocco,*,1 Sophie Stephenson,*,1 Matthew A. -

Cysteine‑Rich 61‑Associated Gene Expression Profile Alterations in Human Glioma Cells
MOLECULAR MEDICINE REPORTS 16: 5561-5567, 2017 Cysteine‑rich 61‑associated gene expression profile alterations in human glioma cells RUI WANG1, BO WEI2, JUN WEI3, YU TIAN2 and CHAO DU2 Departments of 1Radiology, 2Neurosurgery and 3Science and Education Section, China‑Japan Union Hospital of Jilin University, Changchun, Jilin 130033, P.R. China Received January 28, 2016; Accepted February 20, 2017 DOI: 10.3892/mmr.2017.7216 Abstract. The present study aimed to investigate gene be critical for maintaining the role of CYR61 during cancer expression profile alterations associated with cysteine‑rich 61 progression. (CYR61) expression in human glioma cells. The GSE29384 dataset, downloaded from the Gene Expression Omnibus, Introduction includes three LN229 human glioma cell samples expressing CYR61 induced by doxycycline (Dox group), and three Cysteine‑rich 61 (CYR61) is a secreted, cysteine‑rich, control samples not exposed to doxycycline (Nodox group). heparin‑binding protein (1) involved in a variety of cellular Differentially expressed genes (DEGs) between the Dox and functions including adhesion, migration and proliferation (2). Nodox groups were identified with cutoffs of |log2 fold change Previously, Xie et al (3) reported that CYR61 was overexpressed (FC)|>0.5 and P<0.05. Gene ontology and Kyoto Encyclopedia in 66 primary gliomas compared with healthy brain samples, of Genes and Genomes pathway enrichment analyses for and that CYR61 expression was significantly correlated with DEGs were performed. Protein‑protein interaction (PPI) tumor grade and patient survival (3). CYR61‑overexpressing network and module analyses were performed to identify glioma cells were observed to have an increased proliferation the most important genes. -

"The Genecards Suite: from Gene Data Mining to Disease Genome Sequence Analyses". In: Current Protocols in Bioinformat
The GeneCards Suite: From Gene Data UNIT 1.30 Mining to Disease Genome Sequence Analyses Gil Stelzer,1,5 Naomi Rosen,1,5 Inbar Plaschkes,1,2 Shahar Zimmerman,1 Michal Twik,1 Simon Fishilevich,1 Tsippi Iny Stein,1 Ron Nudel,1 Iris Lieder,2 Yaron Mazor,2 Sergey Kaplan,2 Dvir Dahary,2,4 David Warshawsky,3 Yaron Guan-Golan,3 Asher Kohn,3 Noa Rappaport,1 Marilyn Safran,1 and Doron Lancet1,6 1Department of Molecular Genetics, Weizmann Institute of Science, Rehovot, Israel 2LifeMap Sciences Ltd., Tel Aviv, Israel 3LifeMap Sciences Inc., Marshfield, Massachusetts 4Toldot Genetics Ltd., Hod Hasharon, Israel 5These authors contributed equally to the paper 6Corresponding author GeneCards, the human gene compendium, enables researchers to effectively navigate and inter-relate the wide universe of human genes, diseases, variants, proteins, cells, and biological pathways. Our recently launched Version 4 has a revamped infrastructure facilitating faster data updates, better-targeted data queries, and friendlier user experience. It also provides a stronger foundation for the GeneCards suite of companion databases and analysis tools. Improved data unification includes gene-disease links via MalaCards and merged biological pathways via PathCards, as well as drug information and proteome expression. VarElect, another suite member, is a phenotype prioritizer for next-generation sequencing, leveraging the GeneCards and MalaCards knowledgebase. It au- tomatically infers direct and indirect scored associations between hundreds or even thousands of variant-containing genes and disease phenotype terms. Var- Elect’s capabilities, either independently or within TGex, our comprehensive variant analysis pipeline, help prepare for the challenge of clinical projects that involve thousands of exome/genome NGS analyses. -

Rnaseq Reveals the Contribution of Interferon Stimulated Genes to the Increased Host Defense and Decreased PPR Viral Replication in Cattle
viruses Article RNAseq Reveals the Contribution of Interferon Stimulated Genes to the Increased Host Defense and Decreased PPR Viral Replication in Cattle 1, 1, Krishnaswamy Gopalan Tirumurugaan y , Rahul Mohanchandra Pawar y, Gopal Dhinakar Raj 2,* , Arthanari Thangavelu 3, John A. Hammond 4 and Satya Parida 4,* 1 Department of Animal Biotechnology, Madras Veterinary College, Tamil Nadu Veterinary and Animal Sciences University, Chennai 600007, India; [email protected] (K.G.T.); [email protected] (R.M.P.) 2 Centre for Animal Health Studies, Tamil Nadu Veterinary and Animal Sciences University, Chennai 600051, India 3 Department of Veterinary Microbiology, Madras Veterinary College, Tamil Nadu Veterinary and Animal Sciences University, Chennai 600007, India; [email protected] 4 The Pirbright Institute, Ash Road, Pirbright, Surrey GU24 0NF, UK; [email protected] * Correspondence: [email protected] (G.D.R.); [email protected] (S.P.) These authors contributed equally. y Received: 7 March 2020; Accepted: 16 April 2020; Published: 20 April 2020 Abstract: Peste des petits ruminants virus (PPRV) is known to replicate in a wide variety of ruminants causing very species-specific clinical symptoms. Small ruminants (goats and sheep) are susceptible to disease while domesticated cattle and buffalo are dead-end hosts and do not display clinical symptoms. Understanding the host factors that influence differential pathogenesis and disease susceptibility could help the development of better diagnostics and control measures. To study this, we generated transcriptome data from goat and cattle peripheral blood mononuclear cells (PBMC) experimentally infected with PPRV in-vitro. After identifying differentially expressed genes, we further analyzed these immune related pathway genes using the Search Tool for the Retrieval of Interacting Genes/Proteins (STRING) and selected candidate genes were validated using in-vitro experiments. -

Impaired Iloprost-Induced Platelet Inhibition and Phosphoproteome Changes in Patients with Confirmed Pseudohypoparathyroidism Ty
www.nature.com/scientificreports OPEN Impaired iloprost‑induced platelet inhibition and phosphoproteome changes in patients with confrmed pseudohypoparathyroidism type Ia, linked to genetic mutations in GNAS Frauke Swieringa1,2, Fiorella A. Solari1,9, Oliver Pagel1,9, Florian Beck1, Jingnan Huang1,2, Marion A. H. Feijge2, Kerstin Jurk4, Irene M. L. W. Körver‑Keularts5, Nadine J. A. Mattheij2, Jörg Faber3, Joachim Pohlenz3, Alexandra Russo3, Connie T. R. M. Stumpel5,6, Dirk E. Schrander7, Barbara Zieger8, Paola E. J. van der Meijden2, René P. Zahedi1, Albert Sickmann1 & Johan W. M. Heemskerk2* Patients diagnosed with pseudohypoparathyroidism type Ia (PHP Ia) sufer from hormonal resistance and abnormal postural features, in a condition classifed as Albright hereditary osteodystrophy (AHO) syndrome. This syndrome is linked to a maternally inherited mutation in the GNAS complex locus, encoding for the GTPase subunit Gsα. Here, we investigated how platelet phenotype and omics analysis can assist in the often difcult diagnosis. By coupling to the IP receptor, Gsα induces platelet inhibition via adenylyl cyclase and cAMP-dependent protein kinase A (PKA). In platelets from seven patients with suspected AHO, one of the largest cohorts examined, we studied the PKA-induced phenotypic changes. Five patients with a confrmed GNAS mutation, displayed impairments in Gsα- dependent VASP phosphorylation, aggregation, and microfuidic thrombus formation. Analysis of the platelet phosphoproteome revealed 2,516 phosphorylation sites, of which 453 were regulated by Gsα-PKA. Common changes in the patients were: (1) a joint panel of upregulated and downregulated phosphopeptides; (2) overall PKA dependency of the upregulated phosphopeptides; (3) links to key platelet function pathways. In one patient with GNAS mutation, diagnosed as non‑AHO, the changes in platelet phosphoproteome were reversed. -

Download Validation Data
PrimePCR™Assay Validation Report Gene Information Gene Name interferon, alpha-inducible protein 6 Gene Symbol IFI6 Organism Human Gene Summary This gene was first identified as one of the many genes induced by interferon. The encoded protein may play a critical role in the regulation of apoptosis. A minisatellite that consists of 26 repeats of a 12 nucleotide repeating element resembling the mammalian splice donor consensus sequence begins near the end of the second exon. Alternatively spliced transcript variants that encode different isoforms by using the two downstream repeat units as splice donor sites have been described. Gene Aliases 6-16, FAM14C, G1P3, IFI-6-16, IFI616 RefSeq Accession No. NC_000001.10, NT_004610.19 UniGene ID Hs.523847 Ensembl Gene ID ENSG00000126709 Entrez Gene ID 2537 Assay Information Unique Assay ID qHsaCID0015543 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000361157, ENST00000339145, ENST00000362020 Amplicon Context Sequence CTCATCCTCCTCACTATCGAGATACTTGTGGGTGGCGTAGCCCATCAGGGCACC AATATTACCTATGACGACGCTGCTGCCACCAGCCCCGAGGCTCTGCAGCGTGGC CACTAGCCCCCCGGCGGGCACGCCGCCCCCATTCAGGATCGCAGACCAGCTCA TCAGCGAGGCAGCCACC Amplicon Length (bp) 148 Chromosome Location 1:27992898-27994825 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 97 R2 0.9994 cDNA Cq Target not expressed in universal RNA cDNA Tm (Celsius) Target not expressed in universal RNA Page 1/5 PrimePCR™Assay Validation Report gDNA Cq Target not expressed in universal RNA Specificity (%) No cross reactivity detected -
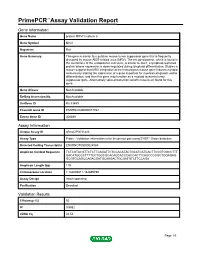
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name protein MRVI1 isoform b Gene Symbol Mrvi1 Organism Rat Gene Summary This gene is similar to a putative mouse tumor suppressor gene that is frequently disrupted by mouse AIDS-related virus (MRV). The encoded protein, which is found in the membrane of the endoplasmic reticulum, is similar to Jaw1, a lymphoid-restricted protein whose expression is down-regulated during lymphoid differentiation. Studies in mouse suggest that MRV integration at the homologous mouse gene induces myeloid leukemia by altering the expression of a gene important for myeloid cell growth and/or differentiation, and thus this gene may function as a myeloid leukemia tumor suppressor gene. Alternatively spliced transcript variants have been found for this gene. Gene Aliases Not Available RefSeq Accession No. Not Available UniGene ID Rn.33889 Ensembl Gene ID ENSRNOG00000017767 Entrez Gene ID 308899 Assay Information Unique Assay ID qRnoCIP0031443 Assay Type Probe - Validation information is for the primer pair using SYBR® Green detection Detected Coding Transcript(s) ENSRNOT00000024060 Amplicon Context Sequence TCTCGTACGTTCTCTTCAGATTCTCCACATACTGCATCATCACTTCCGTGGCCTTT GACATGCGCTTTTCCTGGCGCACAGCACCCACCACTTCAGCCCGGCTGGAGAG GCGTGCAGCCAGACGATGCAAGACTGCGATGTCTTCCAGA Amplicon Length (bp) 119 Chromosome Location 1:182696811-182699799 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 95 R2 0.9982 cDNA Cq 24.54 Page 1/5 PrimePCR™Assay Validation Report cDNA Tm (Celsius) 88 gDNA Cq Specificity -

Mrvil, a Common MRV Integration Site in BXH2 Myeloid Leukemias, Encodes a Protein with Homology to a Lymphoid-Restricted Membrane Protein Jaw1
Oncogene (1999) 18, 2069 ± 2084 ã 1999 Stockton Press All rights reserved 0950 ± 9232/99 $12.00 http://www.stockton-press.co.uk/onc Mrvil, a common MRV integration site in BXH2 myeloid leukemias, encodes a protein with homology to a lymphoid-restricted membrane protein Jaw1 John D Shaughnessy Jr*,1, David A Largaespada2, Erming Tian1, Colin F Fletcher3, Brian C Cho4, Paresh Vyas5, Nancy A Jenkins3 and Neal G Copeland3 1Division of Hematology and Oncology, Department of Medicine, University of Arkansas for Medical Sciences, Little Rock, Arkansas, AR 72205, USA; 2Department of Laboratory Medicine and Pathology, University of Minnesota Cancer Center, Minneapolis, Minnesota, MN 55455, USA; 3Mammalian Genetics Laboratory, ABL-Basic Research Program, NCI-Frederick Cancer Research and Development Center, Frederick, Maryland, MD 21702, USA; 4Department of Microbiology and Immunology, Thomas Jeerson University Cancer Institute, Philadelphia, Pennsylvania, PA 19107, USA; 5Department of Hematology and Oncology, Children's Hospital, Boston, Massachusetts, USA Ecotropic MuLVs induce myeloid leukemia in BXH2 Introduction mice by insertional mutagenesis of cellular proto- oncogenes or tumor suppressor genes. Disease genes BXH2 mice represent an important model for the can thus be identi®ed by viral tagging as common sites of identi®cation of myeloid leukemia disease genes. Not viral integration in BXH2 leukemias. Previous studies only do these mice have the highest spontaneous showed that a frequent common integration site in BXH2 incidence of myeloid leukemia of any known inbred leukemias is the Nf1 tumor suppressor gene. Unexpect- mouse strain, but the leukemias in these mice are edly, about half of the viral integrations at Nf1 retrovirally-induced and the proviruses in the tumors represented a previously undiscovered defective none- can thus be used as insertional tags to identify the cotropic virus, termed MRV.