Ligand Promiscuity in the Tryptophan Repressor – from Structural Understanding Towards Rational Design
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Serotonin Functioning and Adolescents' Alcohol
Development and Psychopathology, 2017, page 1 of 21 # Cambridge University Press 2017 doi:10.1017/S095457941700058X Serotonin functioning and adolescents’ alcohol use: A genetically informed study examining mechanisms of risk FRANCES L. WANG,a LAURIE CHASSIN,a JOHN E. BATES,b DANIELLE DICK,c JENNIFER E. LANSFORD,d e d GREGORY S. PETTIT, AND KENNETH A. DODGE aArizona State University; bIndiana University Bloomington; cVirginia Commonwealth University; dDuke University; and eAuburn University Abstract The current study used data from two longitudinal samples to test whether self-regulation, depressive symptoms, and aggression/antisociality were mediators in the relation between a polygenic score indexing serotonin (5-HT) functioning and alcohol use in adolescence. The results from an independent genome-wide association study of 5-hydroxyindoleacetic acid in the cerebrospinal fluid were used to create 5-HT polygenic risk scores. Adolescents and/or parents reported on adolescents’ self-regulation (Time 1), depressive symptoms (Time 2), aggression/antisociality (Time 2), and alcohol use (Time 3). The results showed that 5-HT polygenic risk did not predict self-regulation. However, adolescents with higher levels of 5-HT polygenic risk showed greater depression and aggression/antisociality. Adolescents’ aggression/antisociality mediated the relation between 5-HT polygenic risk and later alcohol use. Deficits in self- regulation also predicted depression and aggression/antisociality, and indirectly predicted alcohol use through aggression/antisociality. Pathways to alcohol use were especially salient for males from families with low parental education in one of the two samples. The results provide insights into the longitudinal mechanisms underlying the relation between 5-HT functioning and alcohol use (i.e., earlier aggression/antisociality). -

16.1 | Regulation of Gene Expression
436 Chapter 16 | Gene Expression 16.1 | Regulation of Gene Expression By the end of this section, you will be able to do the following: • Discuss why every cell does not express all of its genes all of the time • Describe how prokaryotic gene regulation occurs at the transcriptional level • Discuss how eukaryotic gene regulation occurs at the epigenetic, transcriptional, post-transcriptional, translational, and post-translational levels For a cell to function properly, necessary proteins must be synthesized at the proper time and place. All cells control or regulate the synthesis of proteins from information encoded in their DNA. The process of turning on a gene to produce RNA and protein is called gene expression. Whether in a simple unicellular organism or a complex multi-cellular organism, each cell controls when and how its genes are expressed. For this to occur, there must be internal chemical mechanisms that control when a gene is expressed to make RNA and protein, how much of the protein is made, and when it is time to stop making that protein because it is no longer needed. The regulation of gene expression conserves energy and space. It would require a significant amount of energy for an organism to express every gene at all times, so it is more energy efficient to turn on the genes only when they are required. In addition, only expressing a subset of genes in each cell saves space because DNA must be unwound from its tightly coiled structure to transcribe and translate the DNA. Cells would have to be enormous if every protein were expressed in every cell all the time. -

Formation of Heterodimers Between Wild Type and Mutant Trp Aporepressor Polypeptides of Eschem'chia Coli Thomas J
PROTEINS: Structure, Function, and Genetics 4:173-181 (1988) Formation of Heterodimers Between Wild Type and Mutant trp Aporepressor Polypeptides of Eschem'chia coli Thomas J. Graddis,' Lisa S. Klig,' Charles Yanofsky,' and Dale L. Oxender' 'Department of Biological Chemistry, University of Michigan, Ann Arbor, Michigan 48109 and 'Department of Biological Sciences, Stanford University, Stanford, California 94305 ABSTRACT Availability of the three-dimen- tophan.1-6 The trp aporepressor is activated by the sional structure of the trp repressor of Escherichia binding of two molecules of its corepressor. Once ac- coli and a large group of repressor mutants has per- tivated, trp repressor binds to the operators of the trp, mitted the identification and analysis of mutants aroH, and trpR operons, regulating transcription ini- with substitutions of the amino acid residues that tiati~n.~,'The aroH and trp operons encode biosyn- form the tryptophan binding pocket. Mutant apore- thetic enzymes, whereas the trpR operon encodes the pressors selected for study were overproduced using trp aporepressor. The trpR gene has been cloned and a multicopy expression plasmid. Equilibrium di- its nucleotide sequence determined.3 The aporepres- alysis with ''C-tryptophan and purified mutant and sor and repressor have been purified and the crystal wild type aporepressors was employed to determine structures of both have been solved at high resolu- tryptophan binding constants. The results obtained ti~n.~-"The trp aporepressor is a dimer of identical indicate that replacement of threonine 44 by methi- 107 residue polypeptides." The existence of the three- onine (TM44) or arginine 84 by histidine (RH84) low- dimensional structures, and the availability of many ers the affinity for tryptophan approximately two- repressor mutants,2i12has facilitated this analysis of and four-fold, respectively. -

Tryptophan and 5-Hydroxytryptophan for Depression (Review)
Cochrane Database of Systematic Reviews Tryptophan and 5-Hydroxytryptophan for depression (Review) Shaw KA, Turner J, Del Mar C Shaw KA, Turner J, Del Mar C. Tryptophan and 5-Hydroxytryptophan for depression. Cochrane Database of Systematic Reviews 2002, Issue 1. Art. No.: CD003198. DOI: 10.1002/14651858.CD003198. www.cochranelibrary.com Tryptophan and 5-Hydroxytryptophan for depression (Review) Copyright © 2010 The Cochrane Collaboration. Published by John Wiley & Sons, Ltd. TABLE OF CONTENTS HEADER....................................... 1 ABSTRACT ...................................... 1 PLAINLANGUAGESUMMARY . 2 BACKGROUND .................................... 2 OBJECTIVES ..................................... 3 METHODS ...................................... 3 RESULTS....................................... 4 DISCUSSION ..................................... 4 AUTHORS’CONCLUSIONS . 5 ACKNOWLEDGEMENTS . 5 REFERENCES ..................................... 6 CHARACTERISTICSOFSTUDIES . 10 DATAANDANALYSES. 15 Analysis 1.1. Comparison 1 L-Tryptophan and 5-HTP versus placebo for the treatment of depression, Outcome 1 Numbers ofresponders................................... 15 Analysis 2.1. Comparison 2 Side-effects of L-Tryptophan and 5-HTP versus placebo, Outcome 1 Numbers with side- effects. .................................... 16 FEEDBACK...................................... 16 WHAT’SNEW..................................... 16 HISTORY....................................... 17 CONTRIBUTIONSOFAUTHORS . 17 DECLARATIONSOFINTEREST . 17 SOURCESOFSUPPORT -

Treating Smoking Dependence in Depressed Alcoholics
Treating Smoking Dependence in Depressed Alcoholics Nassima Ait-Daoud, M.D.; Wendy J. Lynch, Ph.D.; J. Kim Penberthy, Ph.D.; Alison B. Breland, Ph.D.; Gabrielle R. Marzani-Nissen, M.D.; and Bankole A. Johnson, D.Sc., M.D., Ph.D. Alcoholism and nicotine dependence share many neurobiological underpinnings; the presence of one drug can cause a person to crave the other. Depressive illness can complicate comorbid alcohol and nicotine dependence by exacerbating the negative affect encountered during attempts to abstain from one or both drugs. Given the morbidity and mortality associated with cigarette smoking, it is imperative to identify treatments to promote smoking cessation and address comorbid psychiatric conditions contemporaneously. Pharmacotherapeutic options demonstrating varying degrees of efficacy and promise in preclinical and clinical studies include nicotine replacement therapy (NRT), selective serotonin reuptake inhibitors (SSRIs), bupropion, varenicline, tricyclic antidepressants, and bupropion plus NRT. Topiramate has shown potential for promoting smoking cessation in alcoholics, although its safety in depressed patients has not been fully explored. The efficacy of medications for treating nicotine dependence is generally enhanced by the inclusion of behavioral interventions such as cognitive behavioral therapy. When group cohesion and social support are stressed, success rates increase among depressed smokers undergoing smoking cessation treatment. Additional treatment strategies targeting dually dependent individuals with -
L‐Tryptophan As the Origin of Psilocybe Natural Products
Minireviews ChemPlusChem doi.org/10.1002/cplu.202000581 1 2 3 Taking Different Roads: l-Tryptophan as the Origin of 4 5 Psilocybe Natural Products 6 [a] [b] [b] [a] 7 Claudius Lenz, Alexander Sherwood, Robert Kargbo, and Dirk Hoffmeister* 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 ChemPlusChem 2021, 86, 28–35 28 © 2020 The Authors. ChemPlusChem published by Wiley-VCH GmbH Wiley VCH Mittwoch, 30.12.2020 2101 / 181786 [S. 28/35] 1 Minireviews ChemPlusChem doi.org/10.1002/cplu.202000581 1 Psychotropic fungi of the genus Psilocybe, colloquially referred highlighted. Psilocybin and its congeners, the heterogeneous 2 to as „magic mushrooms”, are best known for their l- blue-colored psilocyl oligomers, alongside β-carbolines and 3 tryptophan-derived major natural product, psilocybin. Yet, N,N-dimethyl-l-tryptophan, are presented as well as current 4 recent research has revealed a more diverse secondary knowledge on their biosynthesis is provided. The multidiscipli- 5 metabolism that originates from this amino acid. In this nary character of natural product research is demonstrated, and 6 minireview, the focus is laid on l-tryptophan and the various pharmacological, medicinal, ecological, biochemical, and evolu- 7 Psilocybe natural products and their metabolic routes are tionary aspects are included. 8 9 1. Introduction In this minireview, we focus on the biosynthetic routes of L- 10 tryptophan-derived natural products in Psilocybe mushrooms. -

A Placebo Controlled Investigation of the Effects of Tryptophan Or Placebo on Subjective and Objective Measures of Fatigue
European Journal of Clinical Nutrition (1998) 52, 425±431 ß 1998 Stockton Press. All rights reserved 0954±3007/98 $12.00 http://www.stockton-press.co.uk/ejcn A placebo controlled investigation of the effects of tryptophan or placebo on subjective and objective measures of fatigue A Cunliffe, OA Obeid and J Powell-Tuck Department of Human Nutrition, St Bartholomew's and Royal London School of Medicine and Dentistry, Queen Mary and West®eld College, London E1 2AD Objective: To examine the effect of L-tryptophan administration on subjective and objective measures of fatigue in healthy volunteers. Subjects: Six healthy volunteers (4M:2F) were recruited from staff and students at the College. Setting: Department of Human Nutrition, St. Bartholomews and the Royal London School of Medicine and Dentistry. Design: Subjects were tested for central and peripheral fatigue using a visual analogue scale, ¯icker fusion frequency, grip strength, reaction time and wrist ergometry. In addition, plasma free tryptophan concentrations and Trp:LNAA ratio were determined. Measurements were made before, and at 1, 2, 3 and 4 h after drinking one of two test drinks. The drinks were of either caffeine free diet Coca-Cola (placebo) or caffeine free diet Coca- Cola plus L-tryptophan (30 mg/kg: active drink). Each of the six subjects was tested after placebo and active drink with a one week washout period between test days. Results: Subjective fatigue was signi®cantly increased following tryptophan compared to placebo (P < 0.002), and objective measures of central fatigue were signi®cantly increased by tryptophan compared to placebo (¯icker fusion frequency: P < 0.001; reaction time P < 0.001). -

Alcohol Dependence and Withdrawal Impair Serotonergic Regulation Of
Research Articles: Cellular/Molecular Alcohol dependence and withdrawal impair serotonergic regulation of GABA transmission in the rat central nucleus of the amygdala https://doi.org/10.1523/JNEUROSCI.0733-20.2020 Cite as: J. Neurosci 2020; 10.1523/JNEUROSCI.0733-20.2020 Received: 30 March 2020 Revised: 8 July 2020 Accepted: 14 July 2020 This Early Release article has been peer-reviewed and accepted, but has not been through the composition and copyediting processes. The final version may differ slightly in style or formatting and will contain links to any extended data. Alerts: Sign up at www.jneurosci.org/alerts to receive customized email alerts when the fully formatted version of this article is published. Copyright © 2020 the authors 1 Alcohol dependence and withdrawal impair serotonergic regulation of GABA 2 transmission in the rat central nucleus of the amygdala 3 Abbreviated title: Alcohol dependence impairs CeA regulation by 5-HT 4 Sophia Khom, Sarah A. Wolfe, Reesha R. Patel, Dean Kirson, David M. Hedges, Florence P. 5 Varodayan, Michal Bajo, and Marisa Roberto$ 6 The Scripps Research Institute, Department of Molecular Medicine, 10550 N. Torrey Pines 7 Road, La Jolla CA 92307 8 $To whom correspondence should be addressed: 9 Dr. Marisa Roberto 10 Department of Molecular Medicine 11 The Scripps Research Institute 12 10550 N. Torrey Pines Road, La Jolla, CA 92037 13 Tel: (858) 784-7262 Fax: (858) 784-7405 14 Email: [email protected] 15 16 Number of pages: 30 17 Number of figures: 7 18 Number of tables: 2 19 Number of words (Abstract): 250 20 Number of words (Introduction): 650 21 Number of words (Discussion): 1500 22 23 The authors declare no conflict of interest. -

Hallucinogens: an Update
National Institute on Drug Abuse RESEARCH MONOGRAPH SERIES Hallucinogens: An Update 146 U.S. Department of Health and Human Services • Public Health Service • National Institutes of Health Hallucinogens: An Update Editors: Geraline C. Lin, Ph.D. National Institute on Drug Abuse Richard A. Glennon, Ph.D. Virginia Commonwealth University NIDA Research Monograph 146 1994 U.S. DEPARTMENT OF HEALTH AND HUMAN SERVICES Public Health Service National Institutes of Health National Institute on Drug Abuse 5600 Fishers Lane Rockville, MD 20857 ACKNOWLEDGEMENT This monograph is based on the papers from a technical review on “Hallucinogens: An Update” held on July 13-14, 1992. The review meeting was sponsored by the National Institute on Drug Abuse. COPYRIGHT STATUS The National Institute on Drug Abuse has obtained permission from the copyright holders to reproduce certain previously published material as noted in the text. Further reproduction of this copyrighted material is permitted only as part of a reprinting of the entire publication or chapter. For any other use, the copyright holder’s permission is required. All other material in this volume except quoted passages from copyrighted sources is in the public domain and may be used or reproduced without permission from the Institute or the authors. Citation of the source is appreciated. Opinions expressed in this volume are those of the authors and do not necessarily reflect the opinions or official policy of the National Institute on Drug Abuse or any other part of the U.S. Department of Health and Human Services. The U.S. Government does not endorse or favor any specific commercial product or company. -

N,N-Dimethyltryptamine Compound Found in the Hallucinogenic Tea Ayahuasca, Regulates Adult Neurogenesis in Vitro and in Vivo Jose A
Morales-Garcia et al. Translational Psychiatry (2020) 10:331 https://doi.org/10.1038/s41398-020-01011-0 Translational Psychiatry ARTICLE Open Access N,N-dimethyltryptamine compound found in the hallucinogenic tea ayahuasca, regulates adult neurogenesis in vitro and in vivo Jose A. Morales-Garcia 1,2,3,4, Javier Calleja-Conde 5, Jose A. Lopez-Moreno 5, Sandra Alonso-Gil1,2, Marina Sanz-SanCristobal1,2, Jordi Riba6 and Ana Perez-Castillo 1,2,4 Abstract N,N-dimethyltryptamine (DMT) is a component of the ayahuasca brew traditionally used for ritual and therapeutic purposes across several South American countries. Here, we have examined, in vitro and vivo, the potential neurogenic effect of DMT. Our results demonstrate that DMT administration activates the main adult neurogenic niche, the subgranular zone of the dentate gyrus of the hippocampus, promoting newly generated neurons in the granular zone. Moreover, these mice performed better, compared to control non-treated animals, in memory tests, which suggest a functional relevance for the DMT-induced new production of neurons in the hippocampus. Interestingly, the neurogenic effect of DMT appears to involve signaling via sigma-1 receptor (S1R) activation since S1R antagonist blocked the neurogenic effect. Taken together, our results demonstrate that DMT treatment activates the subgranular neurogenic niche regulating the proliferation of neural stem cells, the migration of neuroblasts, and promoting the generation of new neurons in the hippocampus, therefore enhancing adult neurogenesis and -
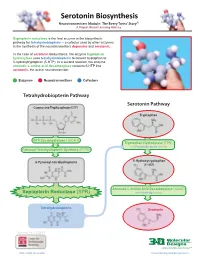
Serotonin Biosynthesis Neurotransmitters Module: the Beery Twins’ Story© a Project-Based Learning Activity
Serotonin Biosynthesis Neurotransmitters Module: The Beery Twins’ Story© A Project-Based Learning Activity Sepiapterin reductase is the final enzyme in the biosynthetic pathway for tetrahydrobiopterin – a cofactor used by other enzymes in the synthesis of the neurotransmitters dopamine and serotonin. In the case of serotonin biosynthesis, the enzyme tryptophan hydroxylase uses tetrahydrobiopterin to convert tryptophan to 5-hydroxytryptophan (5-HTP). In a second reaction, the enzyme aromatic L-amino acid decarboxylase converts 5-HTP into serotonin, the active neurotransmitter. Enzymes Neurotransmitters Cofactors Tetrahydrobiopterin Pathway Serotonin Pathway Guanosine Triphosphate (GTP) Tryptophan GTP Cyclohydrolase I (GCH1) Tryptophan Hydroxylase (TPH) with Tetrahydrobiopterin Cofactor Pyruvoyl-Tetrahydropterin Synthase (PTPS) 6-Pyruvoyl-tetrahydropterin 5-Hydroxytryptophan (5-HTP) Aromatic L-Amino Acid Decarboxylase (AAAD) Sepiapterin Reductase (SPR) with Vitamin B6 Cofactor Tetrahydrobiopterin Serotonin Version 1.4 -12/2015 ...where molecules become real TM http://cbm.msoe.edu www.3dmoleculardesigns.com Serotonin Biosynthesis Model Guide Neurotransmitters Module: The Beery Twins’ Story© A Project-Based Learning Activity TPH AAAD Tryptophan 5-Hydroxytryptophan Serotonin Tryptophan (Trp or W) is one of the 20 standard 5-Hydroxytryptophan, an intermediate The nal step in the serotonin biosynthesis amino acids and is an essential amino acid that molecule in the serotonin biosynthesis pathway requires the removal of the cannot be synthesized by the human body. pathway, is formed by the addition of a carboxylic acid group (COOH) from the Tryptophan is composed of the standard amino hydroxyl (OH) group to the fth carbon of the backbone of 5-hydroxytryptophan to form acid backbone with an indole ring side chain. indole ring of tryptophan. -

Monoamine Biosynthesis Via a Noncanonical Calcium-Activatable Aromatic Amino Acid Decarboxylase in Psilocybin Mushroom
Monoamine Biosynthesis via a Noncanonical Calcium-Activatable Aromatic Amino Acid Decarboxylase in Psilocybin Mushroom The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation Torrens-Spence, Michael Patrick et al. "Monoamine Biosynthesis via a Noncanonical Calcium-Activatable Aromatic Amino Acid Decarboxylase in Psilocybin Mushroom." ACS chemical biology 13 (2018): 3343-3353 © 2018 The Author(s) As Published 10.1021/acschembio.8b00821 Publisher American Chemical Society (ACS) Version Author's final manuscript Citable link https://hdl.handle.net/1721.1/124629 Terms of Use Article is made available in accordance with the publisher's policy and may be subject to US copyright law. Please refer to the publisher's site for terms of use. Articles Cite This: ACS Chem. Biol. XXXX, XXX, XXX−XXX pubs.acs.org/acschemicalbiology Monoamine Biosynthesis via a Noncanonical Calcium-Activatable Aromatic Amino Acid Decarboxylase in Psilocybin Mushroom † ∇ † ‡ § ∇ † † ∥ Michael Patrick Torrens-Spence, , Chun-Ting Liu, , , , Tomaś̌Pluskal, Yin Kwan Chung, , † ‡ and Jing-Ke Weng*, , † Whitehead Institute for Biomedical Research, 455 Main Street, Cambridge, Massachusetts 02142, United States ‡ Department of Biology, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139, United States § Department of Chemistry, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139, United States ∥ Division of Life Science, Hong Kong University of Science & Technology, Clear Water Bay, Hong Kong, China *S Supporting Information ABSTRACT: Aromatic L-amino acid decarboxylases (AAADs) are a phylogenetically diverse group of enzymes responsible for the decarboxylation of aromatic amino acid substrates into their corresponding aromatic arylalkylamines. AAADs have been extensively studied in mammals and plants as they catalyze the first step in the production of neurotransmitters and bioactive phytochemicals, respectively.