Final Report on the Evaluation 2015
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

WO 2019/079361 Al 25 April 2019 (25.04.2019) W 1P O PCT
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization I International Bureau (10) International Publication Number (43) International Publication Date WO 2019/079361 Al 25 April 2019 (25.04.2019) W 1P O PCT (51) International Patent Classification: CA, CH, CL, CN, CO, CR, CU, CZ, DE, DJ, DK, DM, DO, C12Q 1/68 (2018.01) A61P 31/18 (2006.01) DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, HN, C12Q 1/70 (2006.01) HR, HU, ID, IL, IN, IR, IS, JO, JP, KE, KG, KH, KN, KP, KR, KW, KZ, LA, LC, LK, LR, LS, LU, LY, MA, MD, ME, (21) International Application Number: MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, PCT/US2018/056167 OM, PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SA, (22) International Filing Date: SC, SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, 16 October 2018 (16. 10.2018) TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. (25) Filing Language: English (84) Designated States (unless otherwise indicated, for every kind of regional protection available): ARIPO (BW, GH, (26) Publication Language: English GM, KE, LR, LS, MW, MZ, NA, RW, SD, SL, ST, SZ, TZ, (30) Priority Data: UG, ZM, ZW), Eurasian (AM, AZ, BY, KG, KZ, RU, TJ, 62/573,025 16 October 2017 (16. 10.2017) US TM), European (AL, AT, BE, BG, CH, CY, CZ, DE, DK, EE, ES, FI, FR, GB, GR, HR, HU, ΓΕ , IS, IT, LT, LU, LV, (71) Applicant: MASSACHUSETTS INSTITUTE OF MC, MK, MT, NL, NO, PL, PT, RO, RS, SE, SI, SK, SM, TECHNOLOGY [US/US]; 77 Massachusetts Avenue, TR), OAPI (BF, BJ, CF, CG, CI, CM, GA, GN, GQ, GW, Cambridge, Massachusetts 02139 (US). -
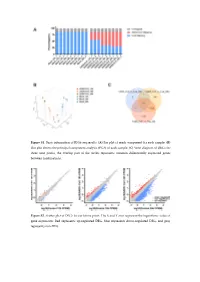
Figure S1. Basic Information of RNA-Seq Results. (A) Bar Plot of Reads Component for Each Sample
Figure S1. Basic information of RNA-seq results. (A) Bar plot of reads component for each sample. (B) Dot plot shows the principal component analysis (PCA) of each sample. (C) Venn diagram of DEGs for three time points, the overlap part of the circles represents common differentially expressed genes between combinations. Figure S2. Scatter plot of DEGs for each time point. The X and Y axes represent the logarithmic value of gene expression. Red represents up-regulated DEG, blue represents down-regulated DEG, and gray represents non-DEG. Table S1. Primers used for quantitative real-time PCR analysis of DEGs. Gene Primer Sequence Forward 5’-CTACGAGTGGATGGTCAAGAGC-3’ FOXO1 Reverse 5’-CCAGTTCCTTCATTCTGCACACG-3’ Forward 5’-GACGTCCGGCATCAGAGAAA-3’ IRS2 Reverse 5’-TCCACGGCTAATCGTCACAG-3’ Forward 5’-CACAACCAGGACCTCACACC-3’ IRS1 Reverse 5’-CTTGGCACGATAGAGAGCGT-3’ Forward 5’-AGGATACCACTCCCAACAGACCT-3’ IL6 Reverse 5’-CAAGTGCATCATCGTTGTTCATAC-3’ Forward 5’-TCACGTTGTACGCAGCTACC-3’ CCL5 Reverse 5’-CAGTCCTCTTACAGCCTTTGG-3’ Forward 5’-CTGTGCAGCCGCAGTGCCTACC-3’ BMP7 Reverse 5’-ATCCCTCCCCACCCCACCATCT-3’ Forward 5’-CTCTCCCCCTCGACTTCTGA-3’ BCL2 Reverse 5’-AGTCACGCGGAACACTTGAT-3’ Forward 5’-CTGTCGAACACAGTGGTACCTG-3’ FGF7 Reverse 5’-CCAACTGCCACTGTCCTGATTTC-3’ Forward 5’-GGGAGCCAAAAGGGTCATCA-3’ GAPDH Reverse 5’-CGTGGACTGTGGTCATGAGT-3’ Supplementary material: Differentially expressed genes log2(SADS-CoV_12h/ Qvalue (SADS-CoV _12h/ Gene Symbol Control_12h) Control_12h) PTGER4 -1.03693 6.79E-04 TMEM72 -3.08132 3.66E-04 IFIT2 -1.02918 2.11E-07 FRAT2 -1.09282 4.66E-05 -

Mir-155: a Novel Target in Allergic Asthma
Int. J. Mol. Sci. 2016, 17, 1773; doi: 10.3390/ijms17101773 S1 of S2 Supplementary Materials: miR-155: A Novel Target in Allergic Asthma Hong Zhou, Junyao Li, Peng Gao, Qi Wang and Jie Zhang Table S1. miR-155 targets identified by TargetScan, miRTarBase and both bioinformatic analyses. Names Total Elements LCORL CHD8 TOMM20 SLC33A1 CHD9 ZNF248 IRF2BP2 DNAJB1 C10orf12 PALLD CARD11 GNAS ZBTB38 RAPH1 ETNK2 MSH6 ARL5B CCDC41 MMP16 RHEB TOMM34 MEF2A RICTOR RAB11FIP2 FAM135A ZBTB18 TMEM33 TCF12 KRAS TM6SF1 DHX40 PICALM MYO10 TCF4 FUBP1 ATP6V1C1 SERTAD2 SH3PXD2A UBQLN2 YWHAZ AGO4 CHAF1A ZNF236 MORC3 MEIS1 WWC1 TAB2 NAA50 PRKAR1A CSNK1G2 PHC2 HBP1 SPRED1 ADAM10 KANSL1 MIDN ZNF644 NFAT5 IL17RB STRN3 MAP3K10 ZSWIM6 DMTF1 ITK PDE3A ZIC3 PELI1 CSNK1A1 ARID2 GSK3B SPIN1 TSPAN14 PTAR1 FOXK1 WEE1 PKN2 TPD52 CARHSP1 MYBL1 WBP1L SAP30L VEZF1 EEF2 FLT1 PHF17 RCOR1 SMAD2 CBFB RORA HIVEP2 CHD7 RAP1B TargetScan 190 SPI1 PEA15 FGF7 RREB1 CBL MYLK S1PR1 TMEM136 PIK3CA NKX3-1 CTLA4 miRTarBase RAB3B SMAD1 ANKFY1 FOS SKIV2L2 SMARCA4 TP53INP1 TSHZ3 PSMG1 FGF2 SKI CPEB4 JARID2 MSI2 SWSAP1 LRRC40 ETS1 COPS3 IKBKE SOCS1 TRIM32 LRRC59 CDC73 RAB5C CAB39 LNX2 NSA2 CDC37 MBNL3 MAFB INPP5D E2F2 PKIA RAB30 CEP41 DET1 UBTD2 C3orf18 BACH1 RAPGEF2 CREBRF SHANK2 PAXBP1 BAG5 KBTBD2 KIF3A HHIP EHD1 HERC4 PALD1 HNRNPA3 N4BP1 PIK3R1 PTPRJ NOVA1 GPM6B CKAP5 TAPT1 CLDN1 SIRT1 SEPT11 COLGALT1 HMGCS1 TLE4 TERF1 ZNF703 FOXO3 KCTD3 APC INADL BCAT1 WNK1 CEBPB TRPS1 CSF1R KDM3A MYO1D RNF123 TADA2B AAK1 RBAK USP8 RCN2 SMAD5 PDE12 ZNF652 MYB SMNDC1 RPTOR PLCE1 KIF26B TNIK RTKN2 ZPLD1 ARRB2 -

A Cross-Sectional Study in 731 Chinese Patients
Genetics Expanding the Phenotypic and Genotypic Landscape of Nonsyndromic High Myopia: A Cross-Sectional Study in 731 Chinese Patients Xue-Bi Cai,1 Yi-Han Zheng,1 De-Fu Chen,1 Fang-Yue Zhou,1 Lu-Qi Xia,1 Xin-Ran Wen,1 Yi-Min Yuan,1 Fang Han,1 Shun-Yu Piao,2 Wenjuan Zhuang,2 Fan Lu,1 Jia Qu,1 A-Yong Yu,1 and Zi-Bing Jin1 1The Eye Hospital, School of Ophthalmology & Optometry, Wenzhou Medical University, National Center for International Research in Regenerative Medicine and Neurogenetics, National Clinical Research Center for Ophthalmology, State Key Laboratory of Ophthalmology, Optometry and Visual Science, Wenzhou, China 2Ningxia Medical University, People’s Hospital of Ningxia Hui Autonomous Region, Yinchuan, China Correspondence: Zi-Bing Jin, The PURPOSE. High myopia (HM) is defined as a refractive error worse than À6.00 diopter (D). This Eye Hospital, Wenzhou Medical Uni- study aims to update the phenotypic and genotypic landscape of nonsyndromic HM and to versity, Wenzhou 325027, China; establish a biological link between the phenotypic traits and genetic deficiencies. [email protected]. A-Yong Yu, The Eye Hospital, Wenz- METHODS. A cross-sectional study involving 731 participants varying in refractive error, axial hou Medical University, Wenzhou length (AL), age, myopic retinopathy, and visual impairment. The phenotypic traits were 325027, China; analyzed by four ophthalmologists while mutational screening was performed in eight [email protected]. autosomal causative genes. Finally, we assessed the clinical relevance of identified mutations Jia Qu, The Eye Hospital, Wenzhou under the guidance of the American College of Medical Genetics and Genomics. -

Clinical, Molecular, and Immune Analysis of Dabrafenib-Trametinib
Supplementary Online Content Chen G, McQuade JL, Panka DJ, et al. Clinical, molecular and immune analysis of dabrafenib-trametinib combination treatment for metastatic melanoma that progressed during BRAF inhibitor monotherapy: a phase 2 clinical trial. JAMA Oncology. Published online April 28, 2016. doi:10.1001/jamaoncol.2016.0509. eMethods. eReferences. eTable 1. Clinical efficacy eTable 2. Adverse events eTable 3. Correlation of baseline patient characteristics with treatment outcomes eTable 4. Patient responses and baseline IHC results eFigure 1. Kaplan-Meier analysis of overall survival eFigure 2. Correlation between IHC and RNAseq results eFigure 3. pPRAS40 expression and PFS eFigure 4. Baseline and treatment-induced changes in immune infiltrates eFigure 5. PD-L1 expression eTable 5. Nonsynonymous mutations detected by WES in baseline tumors This supplementary material has been provided by the authors to give readers additional information about their work. © 2016 American Medical Association. All rights reserved. Downloaded From: https://jamanetwork.com/ on 09/30/2021 eMethods Whole exome sequencing Whole exome capture libraries for both tumor and normal samples were constructed using 100ng genomic DNA input and following the protocol as described by Fisher et al.,3 with the following adapter modification: Illumina paired end adapters were replaced with palindromic forked adapters with unique 8 base index sequences embedded within the adapter. In-solution hybrid selection was performed using the Illumina Rapid Capture Exome enrichment kit with 38Mb target territory (29Mb baited). The targeted region includes 98.3% of the intervals in the Refseq exome database. Dual-indexed libraries were pooled into groups of up to 96 samples prior to hybridization. -

Immune Profiling of Premalignant Lesions in Patients with Lynch Syndrome
Supplementary Online Content Chang K, Taggart MW, Reyes-Uribe L, et al. Immune profiling of premalignant lesions in patients with Lynch syndrome. JAMA Oncol. Published online April 16, 2018. doi:10.1001/jamaoncol.2018.1482 eTable 1. Clinical characteristics of FAP and LS patients eTable 2. Pathology characteristics of FAP and LS polyps eTable 3. Immune profile of LS polyps compared to FAP eTable 4. Immune profile of FAP polyps LS polyps, and LS carcinomas eTable 5. Gene set enrichment analysis of FAP and LS polyps, LS tumor eTable 6. Mutational rate analysis in FAP, LS and LS hypermutated polyps, and Nonhyper and hypermutated Stage I-II TCGA CRC using RNAseq data eTable 7. Immune profile of LS polyps classified by mutational rate (hyper- versus non- hypermutant) eTable 8. Predicted Class I and II HLA genotype of FAP and LS normal mucosa samples eTable 9. List of Predicted neoantigens eTable 10. Pathways analysis of MHC I and II neoantigens eTable 11. Germline and somatic mutation detected in FAP and LS polyps. Mutational data derived from RNA-seq analysis eFigure 1. Immune profile of Lynch syndrome premalignancy and carcinomas eFigure 2. Correlation of DNA and RNA mutation rate in samples from TCGA eFigure 3. Mutation Spectrum of FAP and LS polyps eFigure 4. Immunohistochemical staining of CD4 and FOXP3 eFigure 5. MHC Class I and II neoantigens in Lynch syndrome and FAP premalignancy, and LS carcinomas eMaterials and Methods eReferences © 2018 American Medical Association. All rights reserved. Downloaded From: https://jamanetwork.com/ on 09/24/2021 This supplementary material has been provided by the authors to give readers additional information about their work. -

Supplemental Solier
Supplementary Figure 1. Importance of Exon numbers for transcript downregulation by CPT Numbers of down-regulated genes for four groups of comparable size genes, differing only by the number of exons. Supplementary Figure 2. CPT up-regulates the p53 signaling pathway genes A, List of the GO categories for the up-regulated genes in CPT-treated HCT116 cells (p<0.05). In bold: GO category also present for the genes that are up-regulated in CPT- treated MCF7 cells. B, List of the up-regulated genes in both CPT-treated HCT116 cells and CPT-treated MCF7 cells (CPT 4 h). C, RT-PCR showing the effect of CPT on JUN and H2AFJ transcripts. Control cells were exposed to DMSO. β2 microglobulin (β2) mRNA was used as control. Supplementary Figure 3. Down-regulation of RNA degradation-related genes after CPT treatment A, “RNA degradation” pathway from KEGG. The genes with “red stars” were down- regulated genes after CPT treatment. B, Affy Exon array data for the “CNOT” genes. The log2 difference for the “CNOT” genes expression depending on CPT treatment was normalized to the untreated controls. C, RT-PCR showing the effect of CPT on “CNOT” genes down-regulation. HCT116 cells were treated with CPT (10 µM, 20 h) and CNOT6L, CNOT2, CNOT4 and CNOT6 mRNA were analysed by RT-PCR. Control cells were exposed to DMSO. β2 microglobulin (β2) mRNA was used as control. D, CNOT6L down-regulation after CPT treatment. CNOT6L transcript was analysed by Q- PCR. Supplementary Figure 4. Down-regulation of ubiquitin-related genes after CPT treatment A, “Ubiquitin-mediated proteolysis” pathway from KEGG. -

W O 2019/079360 a L 25 April 2019 (25.04.2019) W 1P O PCT
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization I International Bureau (10) International Publication Number (43) International Publication Date W O 2019/079360 A l 25 April 2019 (25.04.2019) W 1P O PCT (51) International Patent Classification: (72) Inventors; and G01N 33/48 (2006.01) G01N 33/53 (2006.01) (71) Applicants: KEAN, Leslie [US/US]; c/o 818 Stewart St, Suite 603, Seattle, Washington 98101 (US). COLONNA, (21) International Application Number: Lucrezia [US/US]; c/o 818 Stewart St, Suite 603, Seattle, PCT/US2018/056166 Washington 98101 (US). CARROLL, Shaina [US/US]; (22) International Filing Date: c/o 77 Massachusetts Avenue, Cambridge, Massachusetts 16 October 2018 (16. 10.2018) 02139 (US). (25) Filing Language: English (72) Inventors: SHALEK, Alexander K.; c/o 77 Massachu¬ setts Avenue, Cambridge, Massachusetts 02139 (US). ZIE- (26) Publication Language: English GLER, Carly; c/o 77 Massachusetts Avenue, Cambridge, (30) Priority Data: Massachusetts 02139 (US). 62/573,015 16 October 2017 (16. 10.2017) US (74) Agent: SCHER, Michael B. et al.; Johnson, Marcou & (71) Applicants: MASSACHUSETTS INSTITUTE OF Isaacs, LLC, P.O. Bo 691, Hoschton, Georgia 30548 (US). TECHNOLOGY [US/US]; 77 Massachusetts Av¬ (81) Designated States (unless otherwise indicated, for every enue, Cambridge, Massachusetts 02139 (US). SEAT¬ kind of national protection available): AE, AG, AL, AM, TLE CHILDREN'S HOSPITAL DBA SEATTLE AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, BZ, CHILDREN'S RESEARCH INSTITUTE [US/US]; 818 CA, CH, CL, CN, CO, CR, CU, CZ, DE, DJ, DK, DM, DO, Stewart St, Suite 603, Seattle, Washington 98101 (US). -

Autocrine IFN Signaling Inducing Profibrotic Fibroblast Responses By
Downloaded from http://www.jimmunol.org/ by guest on September 23, 2021 Inducing is online at: average * The Journal of Immunology , 11 of which you can access for free at: 2013; 191:2956-2966; Prepublished online 16 from submission to initial decision 4 weeks from acceptance to publication August 2013; doi: 10.4049/jimmunol.1300376 http://www.jimmunol.org/content/191/6/2956 A Synthetic TLR3 Ligand Mitigates Profibrotic Fibroblast Responses by Autocrine IFN Signaling Feng Fang, Kohtaro Ooka, Xiaoyong Sun, Ruchi Shah, Swati Bhattacharyya, Jun Wei and John Varga J Immunol cites 49 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html http://www.jimmunol.org/content/suppl/2013/08/20/jimmunol.130037 6.DC1 This article http://www.jimmunol.org/content/191/6/2956.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 23, 2021. The Journal of Immunology A Synthetic TLR3 Ligand Mitigates Profibrotic Fibroblast Responses by Inducing Autocrine IFN Signaling Feng Fang,* Kohtaro Ooka,* Xiaoyong Sun,† Ruchi Shah,* Swati Bhattacharyya,* Jun Wei,* and John Varga* Activation of TLR3 by exogenous microbial ligands or endogenous injury-associated ligands leads to production of type I IFN. -

Novel Identified Circular Transcript of RCAN2, Circ-RCAN2, Shows Deviated Expression Pattern in Pig Reperfused Infarcted Myocard
International Journal of Molecular Sciences Article Novel Identified Circular Transcript of RCAN2, circ-RCAN2, Shows Deviated Expression Pattern in Pig Reperfused Infarcted Myocardium and Hypoxic Porcine Cardiac Progenitor Cells In Vitro Julia Mester-Tonczar 1 , Patrick Einzinger 2, Johannes Winkler 1, Nina Kastner 1 , Andreas Spannbauer 1 , Katrin Zlabinger 1 , Denise Traxler 1 , Dominika Lukovic 1, Ena Hasimbegovic 1, Georg Goliasch 1, Noemi Pavo 1 and Mariann Gyöngyösi 1,* 1 Department of Internal Medicine II, Division of Cardiology, Medical University of Vienna, 1090 Vienna, Austria; [email protected] (J.M.-T.); [email protected] (J.W.); [email protected] (N.K.); [email protected] (A.S.); [email protected] (K.Z.); [email protected] (D.T.); [email protected] (D.L.); [email protected] (E.H.); [email protected] (G.G.); [email protected] (N.P.) 2 Institute of Information Systems Engineering, Research Unit of Information and Software Engineering, Vienna University of Technology, 1040 Vienna, Austria; [email protected] * Correspondence: [email protected] Citation: Mester-Tonczar, J.; Abstract: Circular RNAs (circRNAs) are crucial in gene regulatory networks and disease devel- Einzinger, P.; Winkler, J.; Kastner, N.; Spannbauer, A.; Zlabinger, K.; Traxler, opment, yet circRNA expression in myocardial infarction (MI) is poorly understood. Here, we D.; Lukovic, D.; Hasimbegovic, E.; harvested myocardium samples from domestic pigs 3 days after closed-chest reperfused MI or Goliasch, G.; et al. Novel Identified sham surgery. Cardiac circRNAs were identified by RNA-sequencing of rRNA-depleted RNA from Circular Transcript of RCAN2, infarcted and healthy myocardium tissue samples. -

A Meta-Analysis of the Effects of High-LET Ionizing Radiations in Human Gene Expression
Supplementary Materials A Meta-Analysis of the Effects of High-LET Ionizing Radiations in Human Gene Expression Table S1. Statistically significant DEGs (Adj. p-value < 0.01) derived from meta-analysis for samples irradiated with high doses of HZE particles, collected 6-24 h post-IR not common with any other meta- analysis group. This meta-analysis group consists of 3 DEG lists obtained from DGEA, using a total of 11 control and 11 irradiated samples [Data Series: E-MTAB-5761 and E-MTAB-5754]. Ensembl ID Gene Symbol Gene Description Up-Regulated Genes ↑ (2425) ENSG00000000938 FGR FGR proto-oncogene, Src family tyrosine kinase ENSG00000001036 FUCA2 alpha-L-fucosidase 2 ENSG00000001084 GCLC glutamate-cysteine ligase catalytic subunit ENSG00000001631 KRIT1 KRIT1 ankyrin repeat containing ENSG00000002079 MYH16 myosin heavy chain 16 pseudogene ENSG00000002587 HS3ST1 heparan sulfate-glucosamine 3-sulfotransferase 1 ENSG00000003056 M6PR mannose-6-phosphate receptor, cation dependent ENSG00000004059 ARF5 ADP ribosylation factor 5 ENSG00000004777 ARHGAP33 Rho GTPase activating protein 33 ENSG00000004799 PDK4 pyruvate dehydrogenase kinase 4 ENSG00000004848 ARX aristaless related homeobox ENSG00000005022 SLC25A5 solute carrier family 25 member 5 ENSG00000005108 THSD7A thrombospondin type 1 domain containing 7A ENSG00000005194 CIAPIN1 cytokine induced apoptosis inhibitor 1 ENSG00000005381 MPO myeloperoxidase ENSG00000005486 RHBDD2 rhomboid domain containing 2 ENSG00000005884 ITGA3 integrin subunit alpha 3 ENSG00000006016 CRLF1 cytokine receptor like