BPS Bioscience
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Mast Cells Promote Seasonal White Adipose Beiging in Humans
Diabetes Volume 66, May 2017 1237 Mast Cells Promote Seasonal White Adipose Beiging in Humans Brian S. Finlin,1 Beibei Zhu,1 Amy L. Confides,2 Philip M. Westgate,3 Brianna D. Harfmann,1 Esther E. Dupont-Versteegden,2 and Philip A. Kern1 Diabetes 2017;66:1237–1246 | DOI: 10.2337/db16-1057 Human subcutaneous (SC) white adipose tissue (WAT) localized to the neck and thorax of humans (4–8), and in a increases the expression of beige adipocyte genes in the process known as beiging (9), UCP1-positive adipocytes winter. Studies in rodents suggest that a number of form in subcutaneous (SC) white adipose tissue (WAT) immune mediators are important in the beiging response. (10). Beige adipocytes have unique developmental origins, We studied the seasonal beiging response in SC WAT gene signatures, and functional properties, including being from lean humans. We measured the gene expression of highly inducible to increase UCP1 in response to catechol- various immune cell markers and performed multivariate amines (9,11–13). Although questions exist about whether analysis of the gene expression data to identify genes beige fat can make a meaningful contribution to energy OBESITY STUDIES that predict UCP1. Interleukin (IL)-4 and, unexpectedly, expenditure in humans (reviewed in Porter et al. [14]), the mast cell marker CPA3 predicted UCP1 gene expres- the induction of beige fat in rodent models is associated sion. Therefore, we investigated the effects of mast with increased energy expenditure and improved glucose cells on UCP1 induction by adipocytes. TIB64 mast cells homeostasis (13). responded to cold by releasing histamine and IL-4, and this medium stimulated UCP1 expression and lipolysis by Activation of the sympathetic nervous system by cold 3T3-L1 adipocytes. -

Role of Negative Regulation of Immune Signaling Pathways in Neutrophil Function
Role of negative regulation of immune signaling pathways in neutrophil function Veronica Azcutia *, Charles A. Parkos *, Jennifer C. Brazil * *Department of Pathology, University of Michigan, Ann Arbor, MI 48109 USA. Summary statement: Review on how PMN functions are negatively regulated by immune signaling pathways. Running title: Negative regulation of PMN function. Send correspondence to V.A and J.C.B, and the Editorial and Production Office information to V.A.: *Veronica Azcutia, Ph.D. Department of Pathology, University of Michigan. Biomedical Science Research Building (BSRB),109 Zina Pitcher Place, Ann Arbor, MI 48109 USA. This is the author manuscript accepted for publication and has undergone full peer review but has not been through the copyediting, typesetting, pagination and proofreading process, which may lead to differences between this version and the Version of Record. Please cite this article as doi: 10.1002/JLB.3MIR0917-374R. This article is protected by copyright. All rights reserved. Tel: (734)-936-1856, Fax: (734)-615-2331, e-Mail: [email protected] *Jennifer C. Brazil, Ph.D. Department of Pathology, University of Michigan. Biomedical Science Research Building (BSRB),109 Zina Pitcher Place, Ann Arbor, MI 48109 USA. Tel: (734)-936-1856, Fax: (734)-615-2331, e-Mail: [email protected] Key words: neutrophils, ITIM, inflammation. Total character count: 43,451; 2 Figures: Figure 1 and 2 are in color; 89 references; 144 words in Abstract; 12 words in summary statement. Abbreviations A(A2)AR = adenosine receptor BM = bone marrow CEACAM = carcinoembryonic antigen-related cell adhesion molecule Csk = C-terminal Scr kinase fMLF = formyl-methionyl-leucyl phenylalanine peptide GAP = GTPase activating proteins GEF = guanine nucleotide exchange factor G-CSF = Granulocyte colony stimulating factor G-CSFR = Granulocyte colony stimulating factor receptor GPCR = G protein coupled receptor GRK = G protein coupled receptor kinase IBD = Inflammatory bowel disease ICAM-1 = Intracellular Adhesion molecule-1 2 This article is protected by copyright. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Gene Expression Within the Periaqueductal Gray Is Linked to Vocal Behavior and Early- Onset Parkinsonism in Pink1 Knockout Rats Cynthia A
Kelm-Nelson and Gammie BMC Genomics (2020) 21:625 https://doi.org/10.1186/s12864-020-07037-4 RESEARCH ARTICLE Open Access Gene expression within the periaqueductal gray is linked to vocal behavior and early- onset parkinsonism in Pink1 knockout rats Cynthia A. Kelm-Nelson1* and Stephen Gammie2 Abstract Background: Parkinson’s disease (PD) is a degenerative disease with early-stage pathology hypothesized to manifest in brainstem regions. Vocal deficits, including soft, monotone speech, result in significant clinical and quality of life issues and are present in 90% of PD patients; yet the underlying pathology mediating these significant voice deficits is unknown. The Pink1−/− rat is a valid model of early-onset PD that presents with analogous vocal communication deficits. Previous work shows abnormal α-synuclein protein aggregation in the periaqueductal gray (PAG), a brain region critical and necessary to the modulation of mammalian vocal behavior. In this study, we used high-throughput RNA sequencing to examine gene expression within the PAG of both male and female Pink1−/− rats as compared to age-matched wildtype controls. We used a bioinformatic approach to (1) test the hypothesis that loss of Pink1 in the PAG will influence the differential expression of genes that interact with Pink1, (2) highlight other key genes that relate to this type of Mendelian PD, and (3) catalog molecular targets that may be important for the production of rat vocalizations. Results: Knockout of the Pink1 gene resulted in differentially expressed genes for both male and female rats that also mapped to human PD datasets. Pathway analysis highlighted several significant metabolic pathways. -

CD Markers Are Routinely Used for the Immunophenotyping of Cells
ptglab.com 1 CD MARKER ANTIBODIES www.ptglab.com Introduction The cluster of differentiation (abbreviated as CD) is a protocol used for the identification and investigation of cell surface molecules. So-called CD markers are routinely used for the immunophenotyping of cells. Despite this use, they are not limited to roles in the immune system and perform a variety of roles in cell differentiation, adhesion, migration, blood clotting, gamete fertilization, amino acid transport and apoptosis, among many others. As such, Proteintech’s mini catalog featuring its antibodies targeting CD markers is applicable to a wide range of research disciplines. PRODUCT FOCUS PECAM1 Platelet endothelial cell adhesion of blood vessels – making up a large portion molecule-1 (PECAM1), also known as cluster of its intracellular junctions. PECAM-1 is also CD Number of differentiation 31 (CD31), is a member of present on the surface of hematopoietic the immunoglobulin gene superfamily of cell cells and immune cells including platelets, CD31 adhesion molecules. It is highly expressed monocytes, neutrophils, natural killer cells, on the surface of the endothelium – the thin megakaryocytes and some types of T-cell. Catalog Number layer of endothelial cells lining the interior 11256-1-AP Type Rabbit Polyclonal Applications ELISA, FC, IF, IHC, IP, WB 16 Publications Immunohistochemical of paraffin-embedded Figure 1: Immunofluorescence staining human hepatocirrhosis using PECAM1, CD31 of PECAM1 (11256-1-AP), Alexa 488 goat antibody (11265-1-AP) at a dilution of 1:50 anti-rabbit (green), and smooth muscle KD/KO Validated (40x objective). alpha-actin (red), courtesy of Nicola Smart. PECAM1: Customer Testimonial Nicola Smart, a cardiovascular researcher “As you can see [the immunostaining] is and a group leader at the University of extremely clean and specific [and] displays Oxford, has said of the PECAM1 antibody strong intercellular junction expression, (11265-1-AP) that it “worked beautifully as expected for a cell adhesion molecule.” on every occasion I’ve tried it.” Proteintech thanks Dr. -
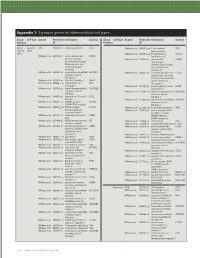
Technical Note, Appendix: an Analysis of Blood Processing Methods to Prepare Samples for Genechip® Expression Profiling (Pdf, 1
Appendix 1: Signature genes for different blood cell types. Blood Cell Type Source Probe Set Description Symbol Blood Cell Type Source Probe Set Description Symbol Fraction ID Fraction ID Mono- Lympho- GSK 203547_at CD4 antigen (p55) CD4 Whitney et al. 209813_x_at T cell receptor TRG nuclear cytes gamma locus cells Whitney et al. 209995_s_at T-cell leukemia/ TCL1A Whitney et al. 203104_at colony stimulating CSF1R lymphoma 1A factor 1 receptor, Whitney et al. 210164_at granzyme B GZMB formerly McDonough (granzyme 2, feline sarcoma viral cytotoxic T-lymphocyte- (v-fms) oncogene associated serine homolog esterase 1) Whitney et al. 203290_at major histocompatibility HLA-DQA1 Whitney et al. 210321_at similar to granzyme B CTLA1 complex, class II, (granzyme 2, cytotoxic DQ alpha 1 T-lymphocyte-associated Whitney et al. 203413_at NEL-like 2 (chicken) NELL2 serine esterase 1) Whitney et al. 203828_s_at natural killer cell NK4 (H. sapiens) transcript 4 Whitney et al. 212827_at immunoglobulin heavy IGHM Whitney et al. 203932_at major histocompatibility HLA-DMB constant mu complex, class II, Whitney et al. 212998_x_at major histocompatibility HLA-DQB1 DM beta complex, class II, Whitney et al. 204655_at chemokine (C-C motif) CCL5 DQ beta 1 ligand 5 Whitney et al. 212999_x_at major histocompatibility HLA-DQB Whitney et al. 204661_at CDW52 antigen CDW52 complex, class II, (CAMPATH-1 antigen) DQ beta 1 Whitney et al. 205049_s_at CD79A antigen CD79A Whitney et al. 213193_x_at T cell receptor beta locus TRB (immunoglobulin- Whitney et al. 213425_at Homo sapiens cDNA associated alpha) FLJ11441 fis, clone Whitney et al. 205291_at interleukin 2 receptor, IL2RB HEMBA1001323, beta mRNA sequence Whitney et al. -

Supplementary Material DNA Methylation in Inflammatory Pathways Modifies the Association Between BMI and Adult-Onset Non- Atopic
Supplementary Material DNA Methylation in Inflammatory Pathways Modifies the Association between BMI and Adult-Onset Non- Atopic Asthma Ayoung Jeong 1,2, Medea Imboden 1,2, Akram Ghantous 3, Alexei Novoloaca 3, Anne-Elie Carsin 4,5,6, Manolis Kogevinas 4,5,6, Christian Schindler 1,2, Gianfranco Lovison 7, Zdenko Herceg 3, Cyrille Cuenin 3, Roel Vermeulen 8, Deborah Jarvis 9, André F. S. Amaral 9, Florian Kronenberg 10, Paolo Vineis 11,12 and Nicole Probst-Hensch 1,2,* 1 Swiss Tropical and Public Health Institute, 4051 Basel, Switzerland; [email protected] (A.J.); [email protected] (M.I.); [email protected] (C.S.) 2 Department of Public Health, University of Basel, 4001 Basel, Switzerland 3 International Agency for Research on Cancer, 69372 Lyon, France; [email protected] (A.G.); [email protected] (A.N.); [email protected] (Z.H.); [email protected] (C.C.) 4 ISGlobal, Barcelona Institute for Global Health, 08003 Barcelona, Spain; [email protected] (A.-E.C.); [email protected] (M.K.) 5 Universitat Pompeu Fabra (UPF), 08002 Barcelona, Spain 6 CIBER Epidemiología y Salud Pública (CIBERESP), 08005 Barcelona, Spain 7 Department of Economics, Business and Statistics, University of Palermo, 90128 Palermo, Italy; [email protected] 8 Environmental Epidemiology Division, Utrecht University, Institute for Risk Assessment Sciences, 3584CM Utrecht, Netherlands; [email protected] 9 Population Health and Occupational Disease, National Heart and Lung Institute, Imperial College, SW3 6LR London, UK; [email protected] (D.J.); [email protected] (A.F.S.A.) 10 Division of Genetic Epidemiology, Medical University of Innsbruck, 6020 Innsbruck, Austria; [email protected] 11 MRC-PHE Centre for Environment and Health, School of Public Health, Imperial College London, W2 1PG London, UK; [email protected] 12 Italian Institute for Genomic Medicine (IIGM), 10126 Turin, Italy * Correspondence: [email protected]; Tel.: +41-61-284-8378 Int. -

Human Lectins, Their Carbohydrate Affinities and Where to Find Them
biomolecules Review Human Lectins, Their Carbohydrate Affinities and Where to Review HumanFind Them Lectins, Their Carbohydrate Affinities and Where to FindCláudia ThemD. Raposo 1,*, André B. Canelas 2 and M. Teresa Barros 1 1, 2 1 Cláudia D. Raposo * , Andr1 é LAQVB. Canelas‐Requimte,and Department M. Teresa of Chemistry, Barros NOVA School of Science and Technology, Universidade NOVA de Lisboa, 2829‐516 Caparica, Portugal; [email protected] 12 GlanbiaLAQV-Requimte,‐AgriChemWhey, Department Lisheen of Chemistry, Mine, Killoran, NOVA Moyne, School E41 of ScienceR622 Co. and Tipperary, Technology, Ireland; canelas‐ [email protected] NOVA de Lisboa, 2829-516 Caparica, Portugal; [email protected] 2* Correspondence:Glanbia-AgriChemWhey, [email protected]; Lisheen Mine, Tel.: Killoran, +351‐212948550 Moyne, E41 R622 Tipperary, Ireland; [email protected] * Correspondence: [email protected]; Tel.: +351-212948550 Abstract: Lectins are a class of proteins responsible for several biological roles such as cell‐cell in‐ Abstract:teractions,Lectins signaling are pathways, a class of and proteins several responsible innate immune for several responses biological against roles pathogens. such as Since cell-cell lec‐ interactions,tins are able signalingto bind to pathways, carbohydrates, and several they can innate be a immuneviable target responses for targeted against drug pathogens. delivery Since sys‐ lectinstems. In are fact, able several to bind lectins to carbohydrates, were approved they by canFood be and a viable Drug targetAdministration for targeted for drugthat purpose. delivery systems.Information In fact, about several specific lectins carbohydrate were approved recognition by Food by andlectin Drug receptors Administration was gathered for that herein, purpose. plus Informationthe specific organs about specific where those carbohydrate lectins can recognition be found by within lectin the receptors human was body. -

CBA Vs. BL/6 BL/6 Vs. CBA Biological Processes
Table SI. Gene ontology (GO) analysis of genes preferentially expressed by CBA or BL/6 DCs CBA vs. BL/6 BL/6 vs. CBA Biological • Immune Response • Regulation of immune effector Processes • Defense response process o Innate immune response o Regulation of leukocyte • Response to external stimulus mediated immunity . Regulation of lymphocyte mediated immunity • Amino sugar catabolic process o Glucosamine catabolic process . N-acetylglucosamine catabolic process • Chitin metabolic process o Chitin catabolic process • Defense response • Response to wounding o Inflammatory response Cellular • Intrinsic to membrane • Extracellular space Component • Cell surface • External side of plasma membrane Molecular • Molecular transducer activity • Chitinase activity Function o Signal transducer activity • G-protein-coupled receptor binding . Receptor activity o Chemokine receptor binding • Carbohydrate binding • Cytokine activity o Sugar binding o Chemokine activity . Monosaccharide binding • C-C chemokine binding • C-C chemokine receptor activity • Cytokine binding 1 Table SII. Genes preferentially expressed by unstimulated CBA DCs mRNA Access Fold Gene Symbol Number Diff mRNA Description H2-Ea NM 010381 88.23 Histocompatibility 2, class II antigen E alpha Ifi202b NM 008327 32.73 Interferon activated gene 202B Chi3l1 NM 007695 24.14 Chitinase-3-like 1 Ctse NM 007799 23.64 Cathepsin E Cd209a (SIGNR5) NM 133238 18.32 CD209a antigen Colec12 (CL-P1) NM130449 16.95 Collectin sub-family member 12 Retnla (FIZZ1) NM 020509 10.22 resistin like alpha Cd209c -

Integrated Weighted Gene Co-Expression Network Analysis Identi Ed That TLR2 and CD40 Are Related to Coronary Artery Disease
Integrated weighted gene co-expression network analysis identied that TLR2 and CD40 are related to coronary artery disease Bin Qi Liuzhou People's Hospital Jian-Hong Chen Liuzhou People's Hospital Lin Tao Liuzhou People's Hospital Chuan-Meng Zhu Liuzhou People's Hospital Yong Wang Liuzhou People's Hospital Guo-Xiong Deng The First People's of Nanning Liu Miao ( [email protected] ) LiuZhou People's Hospital https://orcid.org/0000-0001-6642-7005 Research article Keywords: Gene Expression Omnibus, Integrated weighted gene co-expression network analysis (WGCNA), Functional enrichment, Functional validation and prognostic analysis. Posted Date: October 7th, 2020 DOI: https://doi.org/10.21203/rs.3.rs-86115/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/16 Abstract Background: The current research attempted to identify possible hub genes and pathways of coronary artery disease (CAD) and to detect the possible mechanisms. Methods: Array data from GSE90074 were downloaded from the Gene Expression Omnibus (GEO) database. Integrated weighted gene co-expression network analysis (WGCNA) was performed to analyze the gene module and clinical characteristics. Gene Ontology annotation, Disease Ontology and the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses were performed by clusterProler and the DOSE package in R. A protein-protein interaction (PPI) network was established using Cytoscape software, and signicant modules were analyzed using Molecular Complex Detection to identify hub genes. Then, further functional validation of hub genes in other microarrays and population samples was performed, and survival analysis was performed to investigate the prognosis. -

Pancancer IO360 Human Vapril2018
Gene Name Official Full Gene name Alias/Prev Symbols Previous Name(s) Alias Symbol(s) Alias Name(s) A2M alpha-2-macroglobulin FWP007,S863-7,CPAMD5 ABCF1 ATP binding cassette subfamily F member 1 ABC50 ATP-binding cassette, sub-family F (GCN20),EST123147 member 1 ACVR1C activin A receptor type 1C activin A receptor, type IC ALK7,ACVRLK7 ADAM12 ADAM metallopeptidase domain 12 a disintegrin and metalloproteinase domainMCMPMltna,MLTN 12 (meltrin alpha) meltrin alpha ADGRE1 adhesion G protein-coupled receptor E1 TM7LN3,EMR1 egf-like module containing, mucin-like, hormone receptor-like sequence 1,egf-like module containing, mucin-like, hormone receptor-like 1 ADM adrenomedullin AM ADORA2A adenosine A2a receptor ADORA2 RDC8 AKT1 AKT serine/threonine kinase 1 v-akt murine thymoma viral oncogene homologRAC,PKB,PRKBA,AKT 1 ALDOA aldolase, fructose-bisphosphate A aldolase A, fructose-bisphosphate ALDOC aldolase, fructose-bisphosphate C aldolase C, fructose-bisphosphate ANGPT1 angiopoietin 1 KIAA0003,Ang1 ANGPT2 angiopoietin 2 Ang2 ANGPTL4 angiopoietin like 4 angiopoietin-like 4 pp1158,PGAR,ARP4,HFARP,FIAF,NL2fasting-induced adipose factor,hepatic angiopoietin-related protein,PPARG angiopoietin related protein,hepatic fibrinogen/angiopoietin-related protein,peroxisome proliferator-activated receptor (PPAR) gamma induced angiopoietin-related protein,angiopoietin-related protein 4 ANLN anillin actin binding protein anillin (Drosophila Scraps homolog), actin bindingANILLIN,Scraps,scra protein,anillin, actin binding protein (scraps homolog, Drosophila)