Curriculum Vitae
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Mast Cells Promote Seasonal White Adipose Beiging in Humans
Diabetes Volume 66, May 2017 1237 Mast Cells Promote Seasonal White Adipose Beiging in Humans Brian S. Finlin,1 Beibei Zhu,1 Amy L. Confides,2 Philip M. Westgate,3 Brianna D. Harfmann,1 Esther E. Dupont-Versteegden,2 and Philip A. Kern1 Diabetes 2017;66:1237–1246 | DOI: 10.2337/db16-1057 Human subcutaneous (SC) white adipose tissue (WAT) localized to the neck and thorax of humans (4–8), and in a increases the expression of beige adipocyte genes in the process known as beiging (9), UCP1-positive adipocytes winter. Studies in rodents suggest that a number of form in subcutaneous (SC) white adipose tissue (WAT) immune mediators are important in the beiging response. (10). Beige adipocytes have unique developmental origins, We studied the seasonal beiging response in SC WAT gene signatures, and functional properties, including being from lean humans. We measured the gene expression of highly inducible to increase UCP1 in response to catechol- various immune cell markers and performed multivariate amines (9,11–13). Although questions exist about whether analysis of the gene expression data to identify genes beige fat can make a meaningful contribution to energy OBESITY STUDIES that predict UCP1. Interleukin (IL)-4 and, unexpectedly, expenditure in humans (reviewed in Porter et al. [14]), the mast cell marker CPA3 predicted UCP1 gene expres- the induction of beige fat in rodent models is associated sion. Therefore, we investigated the effects of mast with increased energy expenditure and improved glucose cells on UCP1 induction by adipocytes. TIB64 mast cells homeostasis (13). responded to cold by releasing histamine and IL-4, and this medium stimulated UCP1 expression and lipolysis by Activation of the sympathetic nervous system by cold 3T3-L1 adipocytes. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Human Lectins, Their Carbohydrate Affinities and Where to Find Them
biomolecules Review Human Lectins, Their Carbohydrate Affinities and Where to Review HumanFind Them Lectins, Their Carbohydrate Affinities and Where to FindCláudia ThemD. Raposo 1,*, André B. Canelas 2 and M. Teresa Barros 1 1, 2 1 Cláudia D. Raposo * , Andr1 é LAQVB. Canelas‐Requimte,and Department M. Teresa of Chemistry, Barros NOVA School of Science and Technology, Universidade NOVA de Lisboa, 2829‐516 Caparica, Portugal; [email protected] 12 GlanbiaLAQV-Requimte,‐AgriChemWhey, Department Lisheen of Chemistry, Mine, Killoran, NOVA Moyne, School E41 of ScienceR622 Co. and Tipperary, Technology, Ireland; canelas‐ [email protected] NOVA de Lisboa, 2829-516 Caparica, Portugal; [email protected] 2* Correspondence:Glanbia-AgriChemWhey, [email protected]; Lisheen Mine, Tel.: Killoran, +351‐212948550 Moyne, E41 R622 Tipperary, Ireland; [email protected] * Correspondence: [email protected]; Tel.: +351-212948550 Abstract: Lectins are a class of proteins responsible for several biological roles such as cell‐cell in‐ Abstract:teractions,Lectins signaling are pathways, a class of and proteins several responsible innate immune for several responses biological against roles pathogens. such as Since cell-cell lec‐ interactions,tins are able signalingto bind to pathways, carbohydrates, and several they can innate be a immuneviable target responses for targeted against drug pathogens. delivery Since sys‐ lectinstems. In are fact, able several to bind lectins to carbohydrates, were approved they by canFood be and a viable Drug targetAdministration for targeted for drugthat purpose. delivery systems.Information In fact, about several specific lectins carbohydrate were approved recognition by Food by andlectin Drug receptors Administration was gathered for that herein, purpose. plus Informationthe specific organs about specific where those carbohydrate lectins can recognition be found by within lectin the receptors human was body. -

Pancancer IO360 Human Vapril2018
Gene Name Official Full Gene name Alias/Prev Symbols Previous Name(s) Alias Symbol(s) Alias Name(s) A2M alpha-2-macroglobulin FWP007,S863-7,CPAMD5 ABCF1 ATP binding cassette subfamily F member 1 ABC50 ATP-binding cassette, sub-family F (GCN20),EST123147 member 1 ACVR1C activin A receptor type 1C activin A receptor, type IC ALK7,ACVRLK7 ADAM12 ADAM metallopeptidase domain 12 a disintegrin and metalloproteinase domainMCMPMltna,MLTN 12 (meltrin alpha) meltrin alpha ADGRE1 adhesion G protein-coupled receptor E1 TM7LN3,EMR1 egf-like module containing, mucin-like, hormone receptor-like sequence 1,egf-like module containing, mucin-like, hormone receptor-like 1 ADM adrenomedullin AM ADORA2A adenosine A2a receptor ADORA2 RDC8 AKT1 AKT serine/threonine kinase 1 v-akt murine thymoma viral oncogene homologRAC,PKB,PRKBA,AKT 1 ALDOA aldolase, fructose-bisphosphate A aldolase A, fructose-bisphosphate ALDOC aldolase, fructose-bisphosphate C aldolase C, fructose-bisphosphate ANGPT1 angiopoietin 1 KIAA0003,Ang1 ANGPT2 angiopoietin 2 Ang2 ANGPTL4 angiopoietin like 4 angiopoietin-like 4 pp1158,PGAR,ARP4,HFARP,FIAF,NL2fasting-induced adipose factor,hepatic angiopoietin-related protein,PPARG angiopoietin related protein,hepatic fibrinogen/angiopoietin-related protein,peroxisome proliferator-activated receptor (PPAR) gamma induced angiopoietin-related protein,angiopoietin-related protein 4 ANLN anillin actin binding protein anillin (Drosophila Scraps homolog), actin bindingANILLIN,Scraps,scra protein,anillin, actin binding protein (scraps homolog, Drosophila) -

Suppressive Myeloid Cells Are a Hallmark of Severe COVID-19
medRxiv preprint doi: https://doi.org/10.1101/2020.06.03.20119818; this version posted June 5, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. It is made available under a CC-BY-NC-ND 4.0 International license . 1 Suppressive myeloid cells are a hallmark of 2 severe COVID-19 3 Jonas Schulte-Schrepping1*, Nico Reusch1*, Daniela Paclik2*, Kevin Baßler1*, Stephan 4 Schlickeiser3*, Bowen Zhang4*, Benjamin Krämer5*, Tobias Krammer6*, Sophia Brumhard7*, 5 Lorenzo Bonaguro1*, Elena De Domenico8*, Daniel Wendisch7*, Martin Grasshoff4, Theodore S. 6 Kapellos1, Michael Beckstette4, Tal Pecht1, Adem Saglam8, Oliver Dietrich6, Henrik E. Mei9, Axel 7 R. Schulz9, Claudia Conrad7, Désirée Kunkel10, Ehsan Vafadarnejad6, Cheng-Jian Xu4,11, Arik 8 Horne1, Miriam Herbert1, Anna Drews8, Charlotte Thibeault7, Moritz Pfeiffer7, Stefan 9 Hippenstiel7,12, Andreas Hocke7,12, Holger Müller-Redetzky7, Katrin-Moira Heim7, Felix Machleidt7, 10 Alexander Uhrig7, Laure Bousquillon de Jarcy7, Linda Jürgens7, Miriam Stegemann7, Christoph 11 R. Glösenkamp7, Hans-Dieter Volk2,3,13, Christine Goffinet14,15, Jan Raabe5, Kim Melanie Kaiser5, 12 Michael To Vinh5, Gereon Rieke5, Christian Meisel14, Thomas Ulas8, Matthias Becker8, Robert 13 Geffers16, Martin Witzenrath7,12, Christian Drosten14,19, Norbert Suttorp7,12, Christof von Kalle17, 14 Florian Kurth7,18, Kristian Händler8, Joachim L. Schultze1,8,#,$, Anna C Aschenbrenner20,#, Yang 15 Li4,#, -

Impact of High Altitude Therapy on Type‐2 Immune Responses In
DR. TADECH BOONPIYATHAD (Orcid ID : 0000-0001-8690-7647) DR. GE TAN (Orcid ID : 0000-0003-0026-8739) MRS. MÜBECCEL AKDIS (Orcid ID : 0000-0002-9228-4594) Article type : Original Article: Basic and Translational Allergy Immunology Impact of High Altitude Therapy on Type-2 Immune Responses in Asthma Patients Tadech Boonpiyathad1,2,3, Gertruda Capova3,4, Hans-Werner Duchna3,4, Andrew L. Croxford5, Herve Farine5, Anita Dreher1,4, Martine Clozel5, Juliane Schreiber4, Petr Kubena4, Nonhlanhla Lunjani1,3, David Mirer1, Beate Rückert1, Pattraporn 1,3 1 5 1 Article Satitsuksanoa , Ge Tan , Peter M.A. Groenen , Mübeccel Akdis , Daniel S. Strasser5, Ellen D. Renner3,4,6, Cezmi A. Akdis1,3 1Swiss Institute of Allergy and Asthma Research, University of Zurich, Davos, Switzerland 2Department of Medicine, Phramongkutklao Hospital, Bangkok, Thailand 3Christine Kühne Center for Allergy Research and Education, Davos, Switzerland 4Hochgebirgsklinik Davos, Davos, Switzerland 5Drug Discovery, Idorsia Pharmaceuticals Ltd., Allschwil, Switzerland 6Chair and Institute of Enviromental Medicine – UNIKA-T, TU Munich and Helmholtz Zentrum Munich, Germany Corresponding author: Cezmi A. Akdis Swiss Institute of Allergy and Asthma Research, Obere Strasse 22, CH-7270, Davos Platz, Switzerland Email: [email protected] Tel.: +41 81 140 0848 Fax: +41 81 410 0840 This article has been accepted for publication and undergone full peer review but has not Accepted been through the copyediting, typesetting, pagination and proofreading process, which may lead to differences between this version and the Version of Record. Please cite this article as doi: 10.1111/all.13967 This article is protected by copyright. All rights reserved. Short title Immune response of asthma treatment in high altitude Abstract Background: Asthma patients present with distinct immunological profiles, with a predominance of type 2 endotype. -

Revealing the Acute Asthma Ignorome: Characterization and Validation of Uninvestigated Gene Networks
www.nature.com/scientificreports OPEN Revealing the acute asthma ignorome: characterization and validation of uninvestigated gene Received: 07 December 2015 Accepted: 01 April 2016 networks Published: 21 April 2016 Michela Riba1,*, Jose Manuel Garcia Manteiga1,*, Berislav Bošnjak2,*, Davide Cittaro1, Pavol Mikolka2,†, Connie Le2,‡, Michelle M. Epstein2,# & Elia Stupka1,# Systems biology provides opportunities to fully understand the genes and pathways in disease pathogenesis. We used literature knowledge and unbiased multiple data meta-analysis paradigms to analyze microarray datasets across different mouse strains and acute allergic asthma models. Our combined gene-driven and pathway-driven strategies generated a stringent signature list totaling 933 genes with 41% (440) asthma-annotated genes and 59% (493) ignorome genes, not previously associated with asthma. Within the list, we identified inflammation, circadian rhythm, lung-specific insult response, stem cell proliferation domains, hubs, peripheral genes, and super-connectors that link the biological domains (Il6, Il1ß, Cd4, Cd44, Stat1, Traf6, Rela, Cadm1, Nr3c1, Prkcd, Vwf, Erbb2). In conclusion, this novel bioinformatics approach will be a powerful strategy for clinical and across species data analysis that allows for the validation of experimental models and might lead to the discovery of novel mechanistic insights in asthma. Allergen exposure causes a complex interaction of cellular and molecular networks leading to allergic asthma in susceptible individuals. Experimental mouse models of allergic asthma are widely used to understand disease pathogenesis and elucidate mechanisms underlying the initiation of allergic asthma1. For example, gene profiling of lung tissue from experimental mice during the initiation of allergic asthma in experiments with different proto- cols2–7 validated well-known genes and identified new genes with roles in disease pathogenesis such as C53, Arg18, Adam89, and Pon17, and dissected pathways activated by Il1310 and Stat611. -

Immune-Proteome Landscape Post-COVID19 1 Title
medRxiv preprint doi: https://doi.org/10.1101/2021.08.10.21261834; this version posted August 11, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. It is made available under a CC-BY 4.0 International license . Immune-proteome landscape post-COVID19 1 Title: Immuno-proteomic profiling reveals abundant airway CD8 T cells and ongoing 2 epithelial injury in prolonged post-COVID19 respiratory disease 3 4 Authors: Bavithra Vijayakumar1,2,3,+, Karim Boustani1,4,+, Patricia P. Ogger1,+, Artermis 5 Papadaki5,+, James Tonkin1,3, Christopher M. Orton1,3, Poonam Ghai1, Kornelija 6 Suveizdyte1, Richard J. Hewitt1,3, Robert J. Snelgrove1,4, Philip L. Molyneaux1,3, 7 Justin L. Garner2,3, James E. Peters5,*, Pallav L. Shah1,2,3,*, Clare M. Lloyd1,4* and 8 James A. Harker1,4,* 9 10 1National Heart and Lung Institute, Imperial College London, London, UK 11 2Chelsea and Westminster Hospital, London, UK 12 3Royal Brompton and Harefield Hospitals, Guy’s and St Thomas’ NHS Foundation 13 Trust, London, UK 14 4Asthma UK Centre for Allergic Mechanisms of Asthma 15 5Centre for Inflammatory Disease, Dept of Immunology and Inflammation, Imperial 16 College London, Hammersmith Hospital, London, UK 17 +denotes equal contribution 18 *co-senior authors 19 20 Lead contact: Dr James A Harker, [email protected] 21 Sir Alexander Fleming Building, National Heart and Lung Institute, South Kensington 22 Campus, Imperial College London, London, UK. SW7 2AZ. NOTE: This preprint reports new research that has not been certified1 by peer review and should not be used to guide clinical practice. -

N6-Methyladenosine (M6a)-Mediated Messenger RNA Signatures and the Tumor Immune Microenvironment Can Predict the Prognosis of Hepatocellular Carcinoma
59 Original Article Page 1 of 12 N6-methyladenosine (m6A)-mediated messenger RNA signatures and the tumor immune microenvironment can predict the prognosis of hepatocellular carcinoma Shen Shen1,2#, Jingya Yan1,2#, Yize Zhang1,2, Zihui Dong1,2, Jiyuan Xing1,2, Yuting He1 1Gene Hospital of Henan Province, Precision Medicine Center, The First Affiliated Hospital of Zhengzhou University, Zhengzhou, China; 2Department of Infectious Diseases, the First Affiliated Hospital of Zhengzhou University, Zhengzhou, China Contributions: (I) Conception and design: All authors; (II) Administrative support: Y He, S Shen; (III) Provision of study materials or patients: J Yan, Z Dong; (IV) Collection and assembly of data: Y Zhang; (V) Data analysis and interpretation: S Shen, J Xing; (VI) Manuscript writing: All authors; (VII) Final approval of manuscript: All authors. #These authors contributed equally to this work. Correspondence to: Yuting He. Gene Hospital of Henan Province, Precision Medicine Center, The First Affiliated Hospital of Zhengzhou University, No.1 Jianshe Road, Zhengzhou 450052, China. Email: [email protected]. Background: N6-methyladenosine (m6A)-mediated ribonucleic acid (RNA) methylation is considered to be the most significant and abundant epigenetic modification in eukaryotic cells, and plays an essential role in the carcinogenesis and molecular pathogenesis of hepatocellular carcinoma (HCC). However, the relationship between m6A regulation and immune cell infiltration of the tumor immune microenvironment (TIME) has not yet been clarified. We aimed to investigate the roles of m6A RNA gene regulators in HCC immune regulation and prognosis. Methods: The Cancer Genome Atlas (TCGA) database was used, and unsupervised clustering of 21 m6A regulators was performed based on differential gene expression. -
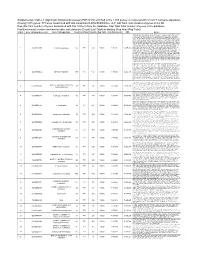
(FDR<0.05) Enriched in the 1015 Genes Co-Clustered with Known T
Supplementary Table 3. Significant biological processes (FDR<0.05) enriched in the 1,015 genes co-clustered with known T cell gene signatures. Among 1,015 genes, 771 were associated with GO annotation in DAVID database v6.7. List Total: total number of genes in my list. Pop Hits: total number of genes associated with this GO term from the database. Pop Total: total number of genes in the database. Fold Enrichment: relative enrichment ratio, calculated by (Count)/(List Total) divided by (Pop Hits)/(Pop Total). Index Gene Ontology Accession Gene Ontology Name Count List Total Pop Hits Pop Total Fold Enrichment FDR Genes AQP9, C1QC, B2M, LILRA1, LILRA2, CLEC4E, S1PR4, LILRA4, IFNG, LILRA6, CLEC4A, VNN1, ERAP2, FAS, CRTAM, C5AR1, GBP5, NCF2, NCF1, NCF4, SERPING1, HLA-DQA2, HLA-DQA1, PDCD1LG2, LILRB1, CCR9, C1QA, C1QB, LILRB2, CCR7, CCR6, UNC13D, CCR5, CD40LG, CCR4, LILRB3, CCR2, LILRB4, HLA-DPA1, VSIG4, HLA-DRA, IL1R2, IL1R1, HLA-DRB1, OAS3, ACP5, OAS1, OAS2, CD74, IFI35, ZAP70, FCER1G, HLA-DRB5, HLA-DPB1, HLA-DOA, HLA-DOB, DHX58, BLNK, IL23R, KIR2DS4, CD300C, SLAMF7, OASL, RGS1, APOL1, CD300A, HMHB1, CD209, CLEC7A, LY86, LY9, CLNK, FCRL4, SH2D1A, NOD2, HAMP, CCL3L1, CCL3L3, TICAM2, ICAM1, GZMA, CMKLR1, LY96, WAS, IL18BP, LAX1, TNFSF12- TNFSF13, HLA-DQB1, CSF2, GPR183, CCR1, GPR65, CXCL9, NCF1C, IL7R, CLEC10A, CCL24, CCL22, CYP27B1, CCL23, FCGR1C, FTHL3, FCGR1A, FCGR1B, BCL3, C2, CD27, CD28, FYB, IL18R1, IL7, CD1C, CTLA4, CCL19, CD1B, CD1A, TRIM22, CD180, CD1E, CCL18, CCL17, CCL13, FCGR2B, FCGR2C, P2RY14, LIME1, CD14, IL16, IL18, TLR1, TNFSF15, -

Human Flow Cytometry Antibodies
Laser Fluorophore Ex (nm) Em (nm) Alterna�ve to mFluor UV375 351 387 mFluor UV460 364 461 ���� ��������� ���������� iFluor 350 344 488 AF350 iFluor 405 402 425 AF405 This list provides a comprehensive overview of the an�bodies mFluor Violet 450 406 445 Pacific Blue® for Flow Cytometry that ImTec can offer. We have arranged mFluor Violet 500 433 498 mFluor Violet 510 412 505 them by target, clone en fluorechrome. Simply click on the mFluor Violet 540 394 537 Pacific Orange® color of your laser to browse the list. iFluor 430 433 495 AF430 iFluor 450 451 502 iFluor 488 491 516 FITC, AF488 mFluor Blue 570 505 564 PE iFluor 514 527 554 AF514 iFluor 532 543 563 AF532 iFluor 546 541 557 AF546 iFluor 555 556 569 TRITC, Cy3®, AF555 iFluor 560 559 571 Cy3B iFluor 568 568 587 AF568 iFluor 594 587 603 AF594 mFluor Green 620 421 613 PE/Texas Red® iFluor 610 609 627 AF610 mFluor Red 700 525 623 APC/Cy5.5® iFluor 633 638 652 AF633 iFluor 647 654 669 Cy5®, AF647 iFluor 660 660 677 AF660 iFluor 670 669 682 Cy5B iFluor 680 683 700 AF680 iFluor 700 690 713 AF700 iFluor 710 712 736 Get the most out of your spectrum with our novel mFluor and iFluor 750 759 777 Cy7®, AF750 iFluor fluorophores. They give you more flexibility when iFluor A7 758 784 mFluor Red 780 610 627 APC/Cy7®, APC/AF750 designing your op�mized mul�color flow panel. The table on iFluor 790 786 811 AF790 the right shows their wavelenghts and for which conjugate iFluor 800 801 820 they can be an alterna�ve.