Frequent Coexistence of Closely Related Tree Species on a Small Scale in Temperate
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Number 3, Spring 1998 Director’S Letter
Planning and planting for a better world Friends of the JC Raulston Arboretum Newsletter Number 3, Spring 1998 Director’s Letter Spring greetings from the JC Raulston Arboretum! This garden- ing season is in full swing, and the Arboretum is the place to be. Emergence is the word! Flowers and foliage are emerging every- where. We had a magnificent late winter and early spring. The Cornus mas ‘Spring Glow’ located in the paradise garden was exquisite this year. The bright yellow flowers are bright and persistent, and the Students from a Wake Tech Community College Photography Class find exfoliating bark and attractive habit plenty to photograph on a February day in the Arboretum. make it a winner. It’s no wonder that JC was so excited about this done soon. Make sure you check of themselves than is expected to seedling selection from the field out many of the special gardens in keep things moving forward. I, for nursery. We are looking to propa- the Arboretum. Our volunteer one, am thankful for each and every gate numerous plants this spring in curators are busy planting and one of them. hopes of getting it into the trade. preparing those gardens for The magnolias were looking another season. Many thanks to all Lastly, when you visit the garden I fantastic until we had three days in our volunteers who work so very would challenge you to find the a row of temperatures in the low hard in the garden. It shows! Euscaphis japonicus. We had a twenties. There was plenty of Another reminder — from April to beautiful seven-foot specimen tree damage to open flowers, but the October, on Sunday’s at 2:00 p.m. -

Literaturverzeichnis
Literaturverzeichnis Abaimov, A.P., 2010: Geographical Distribution and Ackerly, D.D., 2009: Evolution, origin and age of Genetics of Siberian Larch Species. In Osawa, A., line ages in the Californian and Mediterranean flo- Zyryanova, O.A., Matsuura, Y., Kajimoto, T. & ras. Journal of Biogeography 36, 1221–1233. Wein, R.W. (eds.), Permafrost Ecosystems. Sibe- Acocks, J.P.H., 1988: Veld Types of South Africa. 3rd rian Larch Forests. Ecological Studies 209, 41–58. Edition. Botanical Research Institute, Pretoria, Abbadie, L., Gignoux, J., Le Roux, X. & Lepage, M. 146 pp. (eds.), 2006: Lamto. Structure, Functioning, and Adam, P., 1990: Saltmarsh Ecology. Cambridge Uni- Dynamics of a Savanna Ecosystem. Ecological Stu- versity Press. Cambridge, 461 pp. dies 179, 415 pp. Adam, P., 1994: Australian Rainforests. Oxford Bio- Abbott, R.J. & Brochmann, C., 2003: History and geography Series No. 6 (Oxford University Press), evolution of the arctic flora: in the footsteps of Eric 308 pp. Hultén. Molecular Ecology 12, 299–313. Adam, P., 1994: Saltmarsh and mangrove. In Groves, Abbott, R.J. & Comes, H.P., 2004: Evolution in the R.H. (ed.), Australian Vegetation. 2nd Edition. Arctic: a phylogeographic analysis of the circu- Cambridge University Press, Melbourne, pp. marctic plant Saxifraga oppositifolia (Purple Saxi- 395–435. frage). New Phytologist 161, 211–224. Adame, M.F., Neil, D., Wright, S.F. & Lovelock, C.E., Abbott, R.J., Chapman, H.M., Crawford, R.M.M. & 2010: Sedimentation within and among mangrove Forbes, D.G., 1995: Molecular diversity and deri- forests along a gradient of geomorphological set- vations of populations of Silene acaulis and Saxi- tings. -

Post-Triassic Spermatophyta Timetree Adding the Quaternary Radiated Asarum Wild Gingers
Post-Triassic Spermatophyta Timetree Adding the Quaternary Radiated Asarum Wild Gingers Soichi Osozawa ( [email protected] ) KawaOso Molecular Bio-Geology Institute https://orcid.org/0000-0001-5554-1320 Cunio Nackejima Japanese Society of Plant Systematics John Wakabayashi California State University, Fresno Research article Keywords: BEAST v.1.X, combined gene analysis, fossil and geological event calibrations, APG system, increased base substitution rate toward the Recent, Cretaceous peak, radiation, C4 plants, Quaternary glacier- inter glacier cycle Posted Date: November 3rd, 2020 DOI: https://doi.org/10.21203/rs.3.rs-99466/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/24 Abstract Background Angiospermae radiation was known as the mid-Cretaceous event, but adaptive radiation of Asarum is also expected in the Quaternary. In order to know such the Angiospermae evolutionary history through the time, we constructed a whole Spermatophyta timetree employing BEAST v1. X associated with robust fossil calibration function. Results We successfully and precisely dated the Spermatophyta phylogeny, and the Angiospermae topology was concordant to the APG system. Using another function of BEAST, we discovered the exponential increase in base substitution rate in recent geologic time, and another rise of rate at the mid-Cretaceous time. These increasing events correspond to the Quaternary and mid-Cretaceous Angiospermae radiations. Conclusions A probable cause of the recently increasing rate and the consequent radiation was ultimately generation of C4 grasses, reduction of atomospheric CO2, and the start of the Quaternary glacial period. Mid- Cretaceous event was explained by co-radiation with insect beetles as the food plant. -
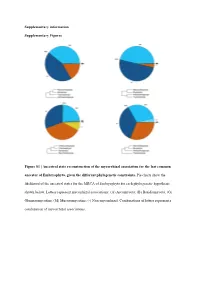
Ancestral State Reconstruction of the Mycorrhizal Association for the Last Common Ancestor of Embryophyta, Given the Different Phylogenetic Constraints
Supplementary information Supplementary Figures Figure S1 | Ancestral state reconstruction of the mycorrhizal association for the last common ancestor of Embryophyta, given the different phylogenetic constraints. Pie charts show the likelihood of the ancestral states for the MRCA of Embryophyta for each phylogenetic hypothesis shown below. Letters represent mycorrhizal associations: (A) Ascomycota; (B) Basidiomycota; (G) Glomeromycotina; (M) Mucoromycotina; (-) Non-mycorrhizal. Combinations of letters represent a combination of mycorrhizal associations. Austrocedrus chilensis Chamaecyparis obtusa Sequoiadendron giganteum Prumnopitys taxifolia Prumnopitys Prumnopitys montana Prumnopitys Prumnopitys ferruginea Prumnopitys Araucaria angustifolia Araucaria Dacrycarpus dacrydioides Dacrycarpus Taxus baccata Podocarpus oleifolius Podocarpus Afrocarpus falcatus Afrocarpus Ephedra fragilis Nymphaea alba Nymphaea Gnetum gnemon Abies alba Abies balsamea Austrobaileya scandens Austrobaileya Abies nordmanniana Thalictrum minus Thalictrum Abies homolepis Caltha palustris Caltha Abies magnifica ia repens Ranunculus Abies religiosa Ranunculus montanus Ranunculus Clematis vitalba Clematis Keteleeria davidiana Anemone patens Anemone Tsuga canadensis Vitis vinifera Vitis Tsuga mertensiana Saxifraga oppositifolia Saxifraga Larix decidua Hypericum maculatum Hypericum Larix gmelinii Phyllanthus calycinus Phyllanthus Larix kaempferi Hieronyma oblonga Hieronyma Pseudotsuga menziesii Salix reinii Salix Picea abies Salix polaris Salix Picea crassifolia Salix herbacea -
International Oaks No. 22.Pdf
INTERNATIONAL OAKS The Journal of the International Oak Society Issue No. 22 Spring 2011 ISSN 1941 2061 Spring 2011 International Oak Journal No. 22 1 The International Oak Society Officers and Board of Directors, 2009 Editorial Office: Membership Office: Béatrice Chassé (France), President Guy Sternberg (USA) Rudy Light (USA) Charles Snyers d'Attenhoven (Belgium), Starhill Forest 11535 East Road Vice-President 12000 Boy Scout Trail Redwood Valley, CA Jim Hitz (USA), Secretary Petersburg, IL 95470 US William Hess (USA), Treasurer 62675-9736 [email protected] Rudy Light (USA), Membership Director e-mail: Dirk Benoît (Belgium), [email protected] Tour Committee Director Allan Taylor (USA), Ron Allan Taylor (USA)USA) Editor, Oak News & Notes 787 17th Street Allen Coombes (Mexico), Boulder, CO 80302 Development Director [email protected] Guy Sternberg (USA), 303-442-5662 Co-editor, IOS Journal Ron Lance (USA), Co-editor, IOS Journal Anyone interested in joining the International Oak Society or ordering information should contact the membership office or see the wesite for membership enrollment form. Benefits include International Oaks and Oak News and Notes publications, conference discounts, and exchanges of seeds and information among members from approximately 30 nations on six continents. International Oak Society website: http://www.internationaloaksociety.org ISSN 1941 2061 Cover photos: Front: Quercus chrysolepsis Liebm. or Uncle Oak, of Palomar Mountain photo©Guy Sternberg Back: Quercus alentejana (a new species) foliage and fruits photos©Michel Timacheff 2 International Oak Journal No. 22 Spring 2011 Table of Contents Message from the Editor Guy Sternberg ..................................................................................................5 Paternity and Pollination in Oaks: Answers Blowin’ in the Wind Mary V. -

Phytochemical Evaluation, Bioassay Screening and Aerial Plant
PHYTOCHEMICAL EVALUATION, BIOASSAY SCREENING AND AERIAL PLANT- MEDIATED SILVER NANOPARTICLES SYNTHESIS USING QUERCUS SEMECARPIFOLIA SMITH Ph. D Thesis By: AISHMA KHATTAK CENTRE OF BIOTECHNOLOGY AND MICROBIOLOGY UNVERSITY OF PESHAWAR Session 2013-2018 PHYTOCHEMICAL EVALUATION, BIOASSAY SCREENING AND AERIAL PLANT- MEDIATED SILVER NANOPARTICLES SYNTHESIS USING QUERCUS SEMECARPIFOLIA SMITH AISHMA KHATTAK A thesis submitted to the University of Peshawar in partial fulfillment of the requirements for the degree of Doctor of Philosophy in Biotechnology CENTRE OF BIOTECHNOLOGY AND MICROBIOLOGY UNVERSITY OF PESHAWAR Session 2013-2018 In the name of Allah, The Most Gracious, The Most Merciful Dedication I wish to dedicate this work to my parents who taught me to value myself and told me that I was the most precious thing in their life. CONTENTS Tables V Figures VII Schemes IX Acknowledgment X Summary XI C H A PTE R 1 IN TR O D U C TIO N & L ITE R A TU RE R E V IE W 1.1 General Introduction 1 1.2 Fagaceae (Family) 6 1.2.1 Description 6 1.2.2 Distribution 6 1.2.3 Importance 6 1.3 Quercus (Genus) 7 1.3.1 Description 7 1.3.2 Distribution 7 1.3.3 Importance 8 1.4 Quercus semecarpifolia Smith (Plant) 11 1.4.1 Description 11 1.4.2 Distribution 12 1.4.3 Importance 12 1.5 Preliminary Phytochemical Profile of the Genus Quercus 14 1.6 Nanotechnology 26 1.6.1 Background 26 1.6.2 Current Status 27 1.7 Nanobiotechnology 28 1.7.1 Background 28 1.7.2 Current Status 28 1.8 Silver(Ag) 29 1.8.1 History 29 1.8.2 Importance 30 1.9 Different Methods Used for the Synthesis -
Plant Breeding and Evaluation
SNA Research Conference Vol. 59 2014 Plant Breeding and Evaluation Tim Rinehart Section Editor Plant Breeding and Evaluation 180 SNA Research Conference Vol. 59 2014 Ploidy Levels and Interploid Hybridization in Panicle Hydrangea (Hydrangea paniculata) Winston T. Beck and Thomas G. Ranney Department of Horticultural Science, North Carolina State University Mountain Horticultural Crops Research and Extension Center 455 Research Dr., Mills River, NC 28759 [email protected] Index Words: cytotype, DNA content, genome size, plant breeding, polyploidy, reproductive biology Significance to Industry: Hydrangea paniculata is a medium to large shrub noted for its excellent floral displays throughout summer, followed by notable winter interest as the flowers dry and persist through the seasons. Although this species is native to Japan, Taiwan, and parts of China, it is grown in landscapes around the world (3, 9).Breeding efforts have been recently been directed towards smaller stature, quality architecture, panicle size, and coverage of sterile (showy) florets, among other attributes. In nature, H. paniculata occurs as diploid, tetraploid, and hexaploid cytotypes, where 1x = 18. There is evidence that most plants in cultivation are tetraploids (10). However, little is known about the ploidy levels of specific cultivars or potential for interploid hybridization and fertility of anisoploid progeny. To better understand the reproductive biology and to further breeding efforts within H. paniculata the objectives of this study were to: 1) determine the ploidy levels of diverse clones and cultivars, and 2) determine fertility and cytotypes of interploid and anisoploid hybrids. Although most cultivated plants were tetraploids, pentaploids and hexaploids were also found. Information on ploidy levels of specific cultivars will facilitate the development of more strategic and efficient breeding programs. -

The Journal of the International Oak Society Proceedings 7Th International Oak Society Conference 9-29-12 / 10-2-12
XXX International Oaks The Journal of the International Oak Society Proceedings 7th International Oak Society Conference 9-29-12 / 10-2-12 Issue No. 24 / 2013 / ISSN 1941-2061 1 International Oaks The Journal of the International Oak Society Proceedings 7th International Oak Society Conference 9-29-12 / 10-2-12 Issue No. 24 / 2013 / ISSN 1941-2061 ,QWHUQDWLRQDO2DN6RFLHW\2IʏFHUVDQG%RDUGRI'LUHFWRUV 2IʏFHUV President Béatrice Chassé (France) Vice-President Charles Snyers d’Attenhoven (Belgium) Secretary Gert Fortgens (The Netherlands) Treasurer James E. Hitz (USA) %RDUGRI'LUHFWRUV (GLWRULDO&RPPLWWHH Membership Director Chairman Emily Griswold (USA) Béatrice Chassé Tour Director Members Shaun Haddock (France) Roderick Cameron International Oaks Allen Coombes Editor Béatrice Chassé Shaun Haddock Co-Editor Allen Coombes (Mexico) Eike Jablonski (Luxemburg) Oak News & Notes Ryan Russell Editor Ryan Russell (USA) Charles Snyers d’Attenhoven International Editor Roderick Cameron (Uruguay) 5HYLHZHUV Website Administrator Andrew Hipp (USA) Charles Snyers d’Attenhoven Antoine Kremer (France) )RUFRQWULEXWLRQVWRInternational Oaks contact Béatrice Chassé [email protected] 0033553621353 Les Pouyouleix 24800 St.-Jory-de-Chalais France Author’s guidelines for submissions can be found at http://www.internationaloaksociety.org/content/author-guidelines-journal-ios © 2013 International Oak Society. Text ʏgures and photographs © of individual authors and photographers. Cover: Dominique Mansion (Quercus robur pollards in Brittany). Photos: p. 7: -

Genomic Landscape of the Global Oak Phylogeny
Research Genomic landscape of the global oak phylogeny Andrew L. Hipp1,2 , Paul S. Manos3, Marlene Hahn1, Michael Avishai4†, Catherine Bodenes5, Jeannine Cavender- Bares6 , Andrew A. Crowl3 , Min Deng7 , Thomas Denk8 , Sorel Fitz-Gibbon9 , Oliver Gailing10 , M. Socorro Gonzalez-Elizondo11 , Antonio Gonzalez-Rodrıguez12, Guido W. Grimm13 , Xiao-Long Jiang7 , Antoine Kremer5 , Isabelle Lesur5, John D. McVay3 , Christophe Plomion5 , Hernando Rodrıguez- Correa12 , Ernst-Detlef Schulze14, Marco C. Simeone15 , Victoria L. Sork9 and Susana Valencia-Avalos16 1The Morton Arboretum, Lisle, IL 60532-1293, USA; 2The Field Museum, Chicago, IL 60605, USA; 3Duke University, Durham, NC 27708, USA; 4 Previously of, The Hebrew University of Jerusalem, Botanical Garden, Zalman Shne’ur St. 1, Jerusalem, Israel; 5INRA, UMR1202 BIOGECO, Cestas F-33610, France; 6University of Minnesota, Minneapolis, MN 55455, USA; 7Shanghai Chenshan Plant Science Research Center, Chinese Academy of Sciences, Shanghai 201602, China; 8Swedish Museum of Natural History, Stockholm 10405, Sweden; 9University of California, Los Angeles, Los Angeles, CA 90095, USA; 10Busgen-Institute,€ Georg-August-University G€ottingen, G€ottingen D-37077, Germany; 11CIIDIR Unidad Durango, Instituto Politecnico Nacional, Durango 34220, Mexico; 12Escuela Nacional de Estudios Superiores Unidad Morelia, Universidad Nacional Auto´noma de Me´xico, Morelia 58190, Me´xico; 13Orle´ans 45100, France; 14Max Planck Institute for Biogeochemistry, Hans-Knoell-Str. 10, Jena 07745, Germany; 15Universita` degli Studi della Tuscia, Viterbo 01100, Italy; 16Herbario de la Facultad de Ciencias, Departamento de Biologı´a Comparada, Universidad Nacional Auto´noma de Me´xico, Circuito Exterior, s.n., Ciudad Universitaria, Coyoaca´n, Me´xico City CP 04510, Me´xico Summary Author for correspondence: The tree of life is highly reticulate, with the history of population divergence emerging from Andrew L. -

Reconstructions Phylogénétiques Du Genre Quercus À Partir De Séquences Du Génome Nucléaire Et Chloroplastique François Hubert
Reconstructions phylogénétiques du genre Quercus à partir de séquences du génome nucléaire et chloroplastique François Hubert To cite this version: François Hubert. Reconstructions phylogénétiques du genre Quercus à partir de séquences du génome nucléaire et chloroplastique. Biologie végétale. Université Sciences et Technologies - Bordeaux I, 2013. Français. NNT : 2013BOR14804. tel-01124107 HAL Id: tel-01124107 https://tel.archives-ouvertes.fr/tel-01124107 Submitted on 6 Mar 2015 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. THESE Présentée à L’UNIVERSITE DE BORDEAUX 1 Ecole Doctorale Sciences et Environnements Par François HUBERT Pour l’obtention du grade de DOCTEUR SPECIALITE: Ecologie évolutive, fonctionnelle et des communautés RECONSTRUCTIONS PHYLOGENETIQUES DU GENRE QUERCUS A PARTIR DE SEQUENCES DU GENOME NUCLEAIRE ET CHLOROPLASTIQUE Soutenue le 21 juin 2013 Devant la commission d’examen formée de : Mme Nathalie FRASCARIA-LACOSTE Professeur AgroParisTech, Orsay Rapporteur Mr Daniel PRAT Professeur de l’Université de Lyon 1 Rapporteur Mr Richard MICHALET Professeur de l’Université de Bordeaux 1 Rapporteur Mr Jean-Pierre RENAUDIN Professeur de l’Université de Bordeaux 1 Président Mr Sylvain JEANDROZ Professeur AgroSup, Dijon Examinateur Mr Antoine KREMER Directeur de Recherche, INRA, Bordeaux Directeur de thèse UMR BIOGECO 1202, 69 route d’Arcachon, 33612 CESTAS Cedex, France N° ordre : 4804 1 2 REMERCIEMENTS Ces six années et demie passées au sein de l’INRA font suite { plusieurs passages au sein de divers laboratoires, nationaux ou internationaux. -

Effects of Changes in Forest Use on Biodiversity 3
Chapter 3 EFFECTS OF CHANGES IN FOREST USE ON BIODIVERSITY 3. Introduction Chapter 3 Chapter 3 Introduction Satoshi Yamashita Research Institute for Humanity and Nature In Chapter 2, the changes in forest utilization during the past 50 to 100 years were revealed for each study site. Based on this understanding, we will discuss the effects of forest utilization on the biological community at each study site in section 3.1 and the effects of human activities on ecosystem functions in section 3.2. In section 3.1, we discuss the effects of forest utilization on communities or populations of living organisms at each study site. At Lambir, species diversity and the species composition of plants (Momose et al.), insects (i.e. butterflies (Itioka et al.), beetles (Kishimoto-Yamada et al.), ants (Matsumoto et al.) ), mammals (i.e. bats (Fukuda et al.), small mammals (Nakagawa et al.)), and macrofungi (Yamashita et al.) were compared among several forest types (i.e. rubber plantation, secondary forest after swidden agriculture, isolated natural forest, and primary forest). In every biological community except for the beetle community, species diversity was highest in the primary forest and generally decreased with increasing disturbance of the forest. At Sabah, species diversity and the species composition of plants (Aiba et al.), insects (i.e. flies (Akutsu et al.)), soil animals (Ito et al.) and mammals (Onoguchi & Matsubayashi) were compared between forest managed under reduced-impact logging, forest managed under conventional logging, and primary forest. Species diversity of plants and mammals was higher in forest managed under reduced-impact logging than in forest managed under conventional logging methods.