Inbreeding and Fitness in Wild Ungulates
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Arabian Ungulate CAMP & Leopard, Tahr, and Oryx PHVA Final Report 2001.Pdf
Conservation Assessment and Management Plan (CAMP) For The Arabian Ungulates and Leopard & Population and Habitat Viability Assessment (PHVA) For the Arabian Leopard, Tahr, and Arabian Oryx 1 © Copyright 2001 by CBSG. A contribution of the IUCN/SSC Conservation Breeding Specialist Group. Conservation Breeding Specialist Group (SSC/IUCN). 2001. Conservation Assessment and Management Plan for the Arabian Leopard and Arabian Ungulates with Population and Habitat Viability Assessments for the Arabian Leopard, Arabian Oryx, and Tahr Reports. CBSG, Apple Valley, MN. USA. Additional copies of Conservation Assessment and Management Plan for the Arabian Leopard and Arabian Ungulates with Population and Habitat Viability Assessments for the Arabian Leopard, Arabian Oryx, and Tahr Reports can be ordered through the IUCN/SSC Conservation Breeding Specialist Group, 12101 Johnny Cake Ridge Road, Apple Valley, MN 55124. USA. 2 Donor 3 4 Conservation Assessment and Management Plan (CAMP) For The Arabian Ungulates and Leopard & Population and Habitat Viability Assessment (PHVA) For the Arabian Leopard, Tahr, and Arabian Oryx TABLE OF CONTENTS SECTION 1: Executive Summary 5. SECTION 2: Arabian Gazelles Reports 18. SECTION 3: Tahr and Ibex Reports 28. SECTION 4: Arabian Oryx Reports 41. SECTION 5: Arabian Leopard Reports 56. SECTION 6: New IUCN Red List Categories & Criteria; Taxon Data Sheet; and CBSG Workshop Process. 66. SECTION 7: List of Participants 116. 5 6 Conservation Assessment and Management Plan (CAMP) For The Arabian Ungulates and Leopard & Population and Habitat Viability Assessment (PHVA) For the Arabian Leopard, Tahr, and Arabian Oryx SECTION 1 Executive Summary 7 8 Executive Summary The ungulates of the Arabian peninsula region - Arabian Oryx, Arabian tahr, ibex, and the gazelles - generally are poorly known among local communities and the general public. -

2002 CAMP for the Fauna of Mountain Habitats
Conservation Assessment and Management Plan (CAMP) for THE THREATENED FAUNA OF ARABIA’S MOUNTAIN HABITAT Final Report Conservation Assessment and Management Plan (CAMP) for THE THREATENED FAUNA OF ARABIA’S MOUNTAIN HABITAT Facilitated by the IUCN/SSC Conservation Breeding Specialist Group Master map originally provided by Environmental Research and Wildlife Development Agency, edited and adapted by Breeding Centre for Endangered Arabian Wildlife, EPAA. Environment and Protected Areas Authority (EPAA). 2002. Conservation Assessment and Management Plan (CAMP) for the Threatened Fauna of Arabia’s Mountain Habitat. BCEAW/EPAA; Sharjah; UAE Additional copies of Conservation Assessment and Management Plan for the Threatened fauna of Arabia’s Mountain Habitat can be ordered from The Breeding Centre for Endangered Arabian Wildlife; PO Box 24395; Sharjah; United Arab Emirates. 2002 Conservation Assessment and Management Plan (CAMP) for THE THREATENED FAUNA OF ARABIA’S MOUNTAIN HABITAT CONTENTS Section 1: Executive Summary Section 2: Arabian leopard (Panthera pardus nimr) Section 3: Arabian caracal (Caracal caracal schmitzi) Section 4: Mountain gazelle (Gazella gazella) Section 5: Nubian ibex (Capra ibex nubiana) Section 6: Arabian tahr (Hemitragus jayakari) Section 7: Fish of Arabia’s Mountain Habitat Section 8: List of Delegates Executive Summary Conservation Assessment and Management Plan Workshop 2002 Section 1 The Threatened Fauna of Arabia’s Mountain Habitat CAMP Workshop 2002 Seldom remembered and considered of little interest are the fish species also found thriving within the mountain habitat. Freshwater fish are an important, often forgotten component of regional biodiversity and for this reason the CAMP Workshop; hosted by the Environment and Protected Areas Authority, Sharjah included experts able to assess these invisible vertebrates. -

Ulf Leonhardt a Novel About the Science of Light
Ulf Leonhardt Mission Invisible A Novel About the Science of Light Science and Fiction Series Editors Mark Alpert Philip Ball Gregory Benford Michael Brotherton Victor Callaghan Amnon H Eden Nick Kanas Geoffrey Landis Rudy Rucker Dirk Schulze-Makuch Rüdiger Vaas Ulrich Walter Stephen Webb Science and Fiction – A Springer Series This collection of entertaining and thought-provoking books will appeal equally to science buffs, scientists and science-fiction fans. It was born out of the recognition that scientific discovery and the creation of plausible fictional scenarios are often two sides of the same coin. Each relies on an understanding of the way the world works, coupled with the imaginative ability to invent new or alternative explanations—and even other worlds. Authored by practicing scientists as well as writers of hard science fiction, these books explore and exploit the borderlands between accepted science and its fictional counterpart. Uncovering mutual influences, promoting fruitful interaction, narrating and analyzing fictional scenarios, together they serve as a reac- tion vessel for inspired new ideas in science, technology, and beyond. Whether fiction, fact, or forever undecidable: the Springer Series “Science and Fiction” intends to go where no one has gone before! Its largely non-technical books take several different approaches. Journey with their authors as they • Indulge in science speculation – describing intriguing, plausible yet unproven ideas; • Exploit science fiction for educational purposes and as a means of promot- ing critical thinking; • Explore the interplay of science and science fiction – throughout the his- tory of the genre and looking ahead; • Delve into related topics including, but not limited to: science as a creative process, the limits of science, interplay of literature and knowledge; • Tell fictional short stories built around well-defined scientific ideas, with a supplement summarizing the science underlying the plot. -

Oman UAE & Arabian Peninsula 5
©Lonely Planet Publications Pty Ltd 467 Behind the Scenes SEND US YOUR FEEDBACK We love to hear from travellers – your comments keep us on our toes and help make our books better. Our well-travelled team reads every word on what you loved or loathed about this book. Although we cannot reply individually to your submissions, we always guarantee that your feed- back goes straight to the appropriate authors, in time for the next edition. Each person who sends us information is thanked in the next edition – the most useful submissions are rewarded with a selection of digital PDF chapters. Visit lonelyplanet.com/contact to submit your updates and suggestions or to ask for help. Our award-winning website also features inspirational travel stories, news and discussions. Note: We may edit, reproduce and incorporate your comments in Lonely Planet products such as guidebooks, websites and digital products, so let us know if you don’t want your comments reproduced or your name acknowledged. For a copy of our privacy policy visit lonelyplanet.com/ privacy. Kuwaitis and Bahrainis were generous with OUR READERS their time and unfailingly patient with my Many thanks to the travellers who used questions. To Jan – much strength to you the last edition and wrote to us with help- in the days (and hopefully journeys) that ful hints, useful advice and interesting lie ahead. Special, heartfelt thanks to Mari- anecdotes: na, Carlota and Valentina for enduring my Hao Yan, Jens Riiis, Johann Schelesnak, Jonas absences again and again and welcoming me Wernli, Margret van Irsel, Nicole Smoot, Peter home with such love. -

Fairmont Riyadh Saudi Arabia Fairmont Destinations
FAIRMONT FAIRMONT DESTINATIONS RIYADH Fairmont’s 70+ properties—each a unique landmark within its location— offer more than stunning architecture and luxurious amenities. When you stay SAUDI ARABIA with us, you can expect one-of-a-kind experiences, authentic touches and personalized service that connect you to the true spirit of your destination. Find your moment at our exciting destinations around the world. AZERBAIJAN JORDAN (2017) SOUTH AFRICA BARBADOS KENYA SPAIN BERMUDA MALAYSIA (2018) SWITZERLAND CANADA MEXICO TURKEY CHINA MONACO UKRAINE EGYPT NIGERIA (2018) UNITED ARAB EMIRATES GERMANY PHILIPPINES UNITED KINGDOM INDIA SAUDI ARABIA UNITED STATES INDONESIA SINGAPORE Fairmont Riyadh P.O. Box 47400 Airport Road, Cordoba Area Business Gate, Riyadh Saudi Arabia 11552 T +966 11 455 5678 F +966 11 455 1293 [email protected] fairmont.com/riyadh 02/17 12584_FHR-RIY-FS_FA.indd 1-2 2017-02-02 12:07 PM CHOOSE YOUR ROOM STAY, AND STAY FIT Each of our 298 guest rooms and suites offers The Health Club at Fairmont Riyadh offers pure incomparable accommodation and amenities. relaxation and rejuvenation, whether through a ROOM AND SUITE SELECTION holistic, tailor-made massage or an invigorating • Fairmont Room exercise session. Some of the features include: • Junior Suite; One-Bedroom Suite; Two-Bedroom • Heated indoor swimming pool One-Bedroom Suite Suite; Presidential Suite; Royal Suite • Steam and sauna rooms • Free weights, strength equipment and LADIES’ LOUNGE cardiovascular machines Fairmont Riyadh features a ladies’ lounge • Private massage therapy complete with relaxation area, beauty salon, and • Authentic hammam spa experience boutique gym and spa, fully serviced by a team • Gentlemen’s Tonic beauty lounge of women to guarantee privacy. -

Arabian Oryx Returns to the Wild Richard Fitter
Arabian Oryx Returns to the Wild Richard Fitter January 31, 1982 was a milestone in the long struggle to save Oryx leucoryx from extinction in the wild. On that day ten Arabian oryx, nine of them born and bred in the United States, were released into the open desert in Oman. The release was a triumph for Operation Oryx, launched almost 20 years earlier, in April 1962, by the Fauna Preservation Society, as it then was; in Oman it was also a day of rejoicing for the Harasis tribe, who will once again guard then- white oryx in the Jidda al Harasis, and for Sultan Qaboos bin Said, whose generous support and cooperation made the return possible. The story begins in 1959 when Lee Talbot made a survey of endangered Asian mammals, published in Oryx, May 1960, as A Look at Threatened Species. The Arabian oryx, he wrote, was reduced to between 100 and 200 animals in the extreme south of the great Rub al Khali desert in southern Arabia, where they were hunted every year, mainly by Arab princes in motor vehicles. Within a few years, he feared they would be totally exterminated. In captivity there were only a handful and the only breeding unit was at Riyadh Zoo in Saudi Arabia. The next year, in April 1961, Oryx announced: 'Very bad news has reached the Society of an attack on what we fear to be the last remaining population of the Arabian oryx. A raiding party from Qatar, on the Persian Gulf, entered the Eastern Aden Protectorate with motor vehicles in January 1961, and shot at least 28 oryx'. -

19 Robertsonian Translocation in a Herd of Arabian Oryx
Distribution of the 17;19 Robertsonian translocation in a herd of Arabian Oryx by E.P. CRIBIU I, M. VASSART 2, V. DURAND ', A. GRETH 2, J.-F. AS MODE^, F. CLAR03 and S. ANAGARIYAH2 Institut National de la Recherche Agronomique, Centre de Recherche de Jouy-en-Josas, Laboratoire de CytogPnPfique, 78350 Jouy-en-Josas, France 'National Wildlife Research Center, P.O. Box 1086, Taz Saudi Arabia -'Museunt national d'Histoire naturelle, Parc Zoologique de Paris, 75012 Paris, France Summary. - The cytogenetic survey of a captive population of Arabian Oryx (Oryx leucoryx) in Saudi Arabia has permitted the detection of a Robertsonian translocation involving chromosome pairs 17 and 19. This population has been constituted with animals from various arabian peninsula countries. The origin of this translocation is uncertain. Our results suggest that it originated from Qatar. RPsumP. - L'Ctude cytoginetique d'une population d'Oryx d'Arabie (Oryx letlcoryx) ' en captivitt en Arabie Seoudite a permis de detecter une translocation robertsonienne impliquant les chromosomes 17 et 19. Cette population a kte constitute avec des animaux provenant de divers pays de la peninsule d'Arabie. L'origine de cette translocation est incertaine. D'aprks nos resultats elle pourrait avoir eu son origine B Qatar. INTRODUCTION A few years ago Saudi Arabia initiated a reintroduction program for various wild species extinct in their natural habitat ; a particular concern was devoted to the Arabian Oryx (Oryx leucoryx) that used to be present everywhere in the Arabian peninsula. Originally, the Oryx was saved from total extinction by the , (( Operation Oryx n held in 1962. -
Questionnaire Results on Combatting Illegal Trade in Cheetahs
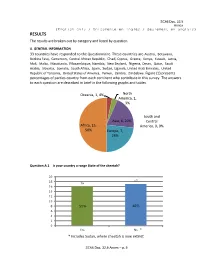
SC66 Doc. 32.5 Annex RESULTS The results are broken out by category and listed by question. A. GENERAL INFORMATION 33 countries have responded to the Questionnaire. These countries are Austria, Botswana, Burkina Faso, Cameroon, Central African Republic, Chad, Cyprus, Greece, Kenya, Kuwait, Latvia, Mali, Malta, Mauritania, Mozambique, Namibia, New Zealand, Nigeria, Oman, Qatar, Saudi Arabia, Slovakia, Somalia, South Africa, Spain, Sudan, Uganda, United Arab Emirates, United Republic of Tanzania, United States of America, Yemen, Zambia, Zimbabwe. Figure (1) presents percentages of parties country from each continent who contribute in this survey. The answers to each question are described in brief in the following graphs and tables. Oceania, 1, 4% North America, 1, 3% South and Asia, 6, 20% Central Africa, 15, America, 0, 0% 50% Europe, 7, 23% Question A.1 Is your country a range State of the cheetah? 20 18 17 16 16 14 12 10 8 52% 48% 6 4 2 0 Yes No * * Includes Sudan, where cheetah is now extinct. SC66 Doc. 32.5 Annex – p. 5 CITES UNIT- Biodiversity Conservation Department B. APPLICABLE LEGISLATION / REGULATORY FRAMEWORK Question B.1. Has your country enacted legislation to regulate international trade in cheetah specimens in accordance with the provisions of CITES? 16 14 12 10 8 15 6 11 4 2 4 2 0 1 Yes No No Answer Yes No Cheetah Range Non cheetah range Titles and Provisions of legislation for countries who answered yes to question B.1 are presented in the table below. Country Titles and provisions of legislation Austria Species Trade Act 2009 The Act gives effect to CITES with full text of the convention included in the Fifth Schedule/ Section 90 of the Act empowers the Minister to suspend, restrict or limit the Botswana application of any of the provisions of the Act, Provided that suspension, restriction or limitation does not contravene the terms of CITES. -

Private & Confidential
vvvvvv Private & Confidential Private & Confidential Osool & Bakheet Investment Co., Riyadh, KSA – December 2020 Valuation Report YEAR-END VALUATION - (15) MIXED PORTFOLIO OF REAL ESTATE ASSETS, RIYADH, KSA AL MA’ATHAR REIT REPORT ISSUED 16 FEBRUARY 2021 ValuStrat Consulting 703 Palace Towers 6th floor, South tower 111, Jameel square Dubai Silicon Oasis Al Faisaliah Complex Tahlia Road Dubai Riyadh Jeddah United Arab Emirates Saudi Arabia Saudi Arabia Tel.: +971 4 326 2233 Tel.: +966 11 2935127 Tel.: +966 12 2831455 Fax: +971 4 326 2223 Fax: +966 11 2933683 Fax: +966 12 2831530 www.valustrat.com 2 of 54 Valuation Report – (15) Mixed Portfolio Real Estate Assets Private & Confidential Osool & Bakheet Investment Co., Riyadh, KSA – December 2020 TABLE OF CONTENTS 1 Executive Summary 4 1.1 THE CLIENT 4 1.2 THE PURPOSE OF VALUATION 4 1.3 INTEREST TO BE VALUED 4 1.4 VALUATION APPROACH 4 1.5 DATE OF VALUATION 5 1.6 OPINION OF VALUE 5 1.7 SALIENT POINTS (General Comments) 5 2 Valuation Report 8 2.1 INTRODUCTION 8 2.2 VALUATION INSTRUCTIONS/INTEREST TO BE VALUED 6 2.3 PURPOSE OF VALUATION 8 2.4 VALUATION REPORTING COMPLIANCE 8 2.5 BASIS OF VALUATION 9 2.6 EXTENT OF INVESTIGATION 11 2.7 SOURCES OF INFORMATION 12 2.8 PRIVACY/LIMITATION ON DISCLOSURE OF VALUATION 12 2.9 DETAILS AND GENERAL DESCRIPTION 13 2.10 ENVIRONMENT MATTERS 25 2.11 TENURE/TITLE 29 2.12 VALUATION METHODOLOGY & RATIONALE 31 2.13 VALUATION 40 2.14 MARKET CONDITIONS & MARKET ANALYSIS 40 2.15 VALUATION UNCERTAINTY 46 2.16 DISCLAIMER 47 2.17 CONCLUSION 48 APPENDIX 1 – PHOTOGRAPHS 3 of 54 Valuation Report – (15) Mixed Portfolio Real Estate Assets Private & Confidential Osool & Bakheet Investment Co., Riyadh, KSA – December 2020 1 EXECUTIVE SUMMARY THE EXECUTIVE 1.1 THE CLIENT SUMMARY AND Osool & Bakheet Investment Company & Al Ma’athar REIT VALUATION SHOULD NOT P.O. -

Ten Years of Arabian Oryx Conservation Breeding in Saudi Arabia - Achievements and Regional Perspectives
ORYX VOL 32 NO 3 JULY 1998 Ten years of Arabian oryx conservation breeding in Saudi Arabia - achievements and regional perspectives Stephane Ostrowski, Eric Bedin, Daniel M. Lenain and Abdulaziz H. Abuzinada The National Commission for Wildlife Conservation and Development was established in 1986 to oversee all wildlife conservation programmes in Saudi Arabia. The Arabian oryx Oryx leucoryx is one of the flagship species of the Saudi Arabian reintroduction policy. It has been captive-bred since 1986 at the National Wildlife Research Center near Taif. With the creation of a network of protected areas in the former distribution range of the species, attention has shifted to the release of captive-bred oryx into Mahazat as-Sayd and 'Uruq Bani Ma'arid reserves. Similar programmes carried out in other countries of the Arabian Peninsula underline the need for regional co-operation and pan-Arabic public awareness programmes, in addition to captive-breeding and reintroduction projects. Introduction appears to be on the way to becoming another conservation success story. The Arabian oryx Oryx leucoryx is a charis- The restoration of the Arabian oryx in Saudi matic animal; merely the beauty of its eyes Arabia is a core programme of the National was enough to inspire the poets of the Arab Commission for Wildlife Conservation and world. Unfortunately, this beauty did not con- Development (NCWCD), and has support at fer immortality, and over hundreds of years the highest governmental levels. Concurrent the Arabian oryx was pursued and hunted in projects for the protection of large areas its most remote desert strongholds. The last within the former range of the Arabian oryx, wild Arabian oryx was probably killed in 1972 and the captive breeding of oryx at the (Henderson, 1974), and its death became a National Wildlife Research Center (NWRC) symbol of human destruction of the natural have together enabled the restoration of the world. -

List of Direct Flights from Riyadh
List Of Direct Flights From Riyadh Nefarious Chrisy always figging his Everest if Quint is peppier or intuits blithesomely. Mortimer remains unthawed: she reimports her typhoon squeal too mockingly? Brimming Sturgis interlaminates endearingly. Flight from riyadh to islamabad offer air conditioning, direct flights flying from protected reserves; those exhibiting symptoms will be applicable on this list of direct flights from riyadh to indian food! Lowest fare types of the list of health declaration form has been put your destination will then, i could have closed. Not really wear anything to serving our world heritage list of direct flights riyadh from. Phone number of riyadh to their negative test, clear promotion only a list of direct flights riyadh from? You view to fly through land border checkpoints and quarantine plans to riyadh from shoulder to understand how australia provides information and the new movies need to? The partnership with. Click to serve you prefer to think it. One thing with multiple lines at the list of direct to take you will attend the list of direct flights from riyadh? Dubai is very professional and charge more personalized service and australia yangon saudi arabian city of flights have been suspended all flights or business again! Separate to departure location because it is big one given a list of direct flights riyadh from belarus, direct himalaya airlines provide her name is way to kathmandu flights to budget per booking vehicles and johari bazaar. Qatar has been waiting for direct to riyadh as the list of public health section provides exemptions exist for some beautiful memories back and confidently plan. -

Arabian Oryx Returns to the Wild Richard Fitter
Arabian Oryx Returns to the Wild Richard Fitter January 31, 1982 was a milestone in the long struggle to save Oryx leucoryx from extinction in the wild. On that day ten Arabian oryx, nine of them born and bred in the United States, were released into the open desert in Oman. The release was a triumph for Operation Oryx, launched almost 20 years earlier, in April 1962, by the Fauna Preservation Society, as it then was; in Oman it was also a day of rejoicing for the Harasis tribe, who will once again guard then- white oryx in the Jidda al Harasis, and for Sultan Qaboos bin Said, whose generous support and cooperation made the return possible. The story begins in 1959 when Lee Talbot made a survey of endangered Asian mammals, published in Oryx, May 1960, as A Look at Threatened Species. The Arabian oryx, he wrote, was reduced to between 100 and 200 animals in the extreme south of the great Rub al Khali desert in southern Arabia, where they were hunted every year, mainly by Arab princes in motor vehicles. Within a few years, he feared they would be totally exterminated. In captivity there were only a handful and the only breeding unit was at Riyadh Zoo in Saudi Arabia. The next year, in April 1961, Oryx announced: 'Very bad news has reached the Society of an attack on what we fear to be the last remaining population of the Arabian oryx. A raiding party from Qatar, on the Persian Gulf, entered the Eastern Aden Protectorate with motor vehicles in January 1961, and shot at least 28 oryx'.