Failed Immune Responses Across Multiple Pathologies Share Pan-Tumor and Circulating Lymphocytic Targets
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A 1.37-Mb 12P11.22-P11.21 Deletion Coincident with a 367-Kb 22Q11.2
CORE Metadata, citation and similar papers at core.ac.uk Provided by Elsevier - Publisher Connector Taiwanese Journal of Obstetrics & Gynecology 53 (2014) 74e78 Contents lists available at ScienceDirect Taiwanese Journal of Obstetrics & Gynecology journal homepage: www.tjog-online.com Short Communication A 1.37-Mb 12p11.22ep11.21 deletion coincident with a 367-kb 22q11.2 duplication detected by array comparative genomic hybridization in an adolescent girl with autism and difficulty in self-care of menstruation Chih-Ping Chen a,b,c,d,e,f,*, Shuan-Pei Lin b,g,h,i, Schu-Rern Chern b, Peih-Shan Wu j, Jun-Wei Su a,k, Chen-Chi Lee a, Wayseen Wang b,l a Department of Obstetrics and Gynecology, Mackay Memorial Hospital, Taipei, Taiwan b Department of Medical Research, Mackay Memorial Hospital, Taipei, Taiwan c Department of Biotechnology, Asia University, Taichung, Taiwan d School of Chinese Medicine, College of Chinese Medicine, China Medical University, Taichung, Taiwan e Institute of Clinical and Community Health Nursing, National Yang-Ming University, Taipei, Taiwan f Department of Obstetrics and Gynecology, School of Medicine, National Yang-Ming University, Taipei, Taiwan g Department of Medicine, Mackay Medical College, New Taipei City, Taiwan h Department of Pediatrics, Mackay Memorial Hospital, Taipei, Taiwan i Mackay Junior College of Medicine, Nursing, and Management, Taipei, Taiwan j Gene Biodesign Co. Ltd, Taipei, Taiwan k Department of Obstetrics and Gynecology, China Medical University Hospital, Taichung, Taiwan l Department of Bioengineering, Tatung University, Taipei, Taiwan article info abstract Article history: Objective: To present an array comparative genomic hybridization (aCGH) characterization of a 12p11.22 Accepted 21 October 2013 ep11.21 microdeletion and 22q11.2 microduplication in an adolescent girl with autism, mental retar- dation, facial dysmorphism, microcephaly, behavior problems, and an apparently balanced reciprocal Keywords: translocation of t(8;12)(q24.3;p11.2). -

DNA Replication Stress Response Involving PLK1, CDC6, POLQ
DNA replication stress response involving PLK1, CDC6, POLQ, RAD51 and CLASPIN upregulation prognoses the outcome of early/mid-stage non-small cell lung cancer patients C. Allera-Moreau, I. Rouquette, B. Lepage, N. Oumouhou, M. Walschaerts, E. Leconte, V. Schilling, K. Gordien, L. Brouchet, Mb Delisle, et al. To cite this version: C. Allera-Moreau, I. Rouquette, B. Lepage, N. Oumouhou, M. Walschaerts, et al.. DNA replica- tion stress response involving PLK1, CDC6, POLQ, RAD51 and CLASPIN upregulation prognoses the outcome of early/mid-stage non-small cell lung cancer patients. Oncogenesis, Nature Publishing Group: Open Access Journals - Option C, 2012, 1, pp.e30. 10.1038/oncsis.2012.29. hal-00817701 HAL Id: hal-00817701 https://hal.archives-ouvertes.fr/hal-00817701 Submitted on 9 Jun 2021 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Distributed under a Creative Commons Attribution - NonCommercial - NoDerivatives| 4.0 International License Citation: Oncogenesis (2012) 1, e30; doi:10.1038/oncsis.2012.29 & 2012 Macmillan Publishers Limited All rights reserved 2157-9024/12 www.nature.com/oncsis ORIGINAL ARTICLE DNA replication stress response involving PLK1, CDC6, POLQ, RAD51 and CLASPIN upregulation prognoses the outcome of early/mid-stage non-small cell lung cancer patients C Allera-Moreau1,2,7, I Rouquette2,7, B Lepage3, N Oumouhou3, M Walschaerts4, E Leconte5, V Schilling1, K Gordien2, L Brouchet2, MB Delisle1,2, J Mazieres1,2, JS Hoffmann1, P Pasero6 and C Cazaux1 Lung cancer is the leading cause of cancer deaths worldwide. -

Transcriptomic and Proteomic Profiling Provides Insight Into
BASIC RESEARCH www.jasn.org Transcriptomic and Proteomic Profiling Provides Insight into Mesangial Cell Function in IgA Nephropathy † † ‡ Peidi Liu,* Emelie Lassén,* Viji Nair, Celine C. Berthier, Miyuki Suguro, Carina Sihlbom,§ † | † Matthias Kretzler, Christer Betsholtz, ¶ Börje Haraldsson,* Wenjun Ju, Kerstin Ebefors,* and Jenny Nyström* *Department of Physiology, Institute of Neuroscience and Physiology, §Proteomics Core Facility at University of Gothenburg, University of Gothenburg, Gothenburg, Sweden; †Division of Nephrology, Department of Internal Medicine and Department of Computational Medicine and Bioinformatics, University of Michigan, Ann Arbor, Michigan; ‡Division of Molecular Medicine, Aichi Cancer Center Research Institute, Nagoya, Japan; |Department of Immunology, Genetics and Pathology, Uppsala University, Uppsala, Sweden; and ¶Integrated Cardio Metabolic Centre, Karolinska Institutet Novum, Huddinge, Sweden ABSTRACT IgA nephropathy (IgAN), the most common GN worldwide, is characterized by circulating galactose-deficient IgA (gd-IgA) that forms immune complexes. The immune complexes are deposited in the glomerular mesangium, leading to inflammation and loss of renal function, but the complete pathophysiology of the disease is not understood. Using an integrated global transcriptomic and proteomic profiling approach, we investigated the role of the mesangium in the onset and progression of IgAN. Global gene expression was investigated by microarray analysis of the glomerular compartment of renal biopsy specimens from patients with IgAN (n=19) and controls (n=22). Using curated glomerular cell type–specific genes from the published literature, we found differential expression of a much higher percentage of mesangial cell–positive standard genes than podocyte-positive standard genes in IgAN. Principal coordinate analysis of expression data revealed clear separation of patient and control samples on the basis of mesangial but not podocyte cell–positive standard genes. -
Primepcr™Assay Validation Report
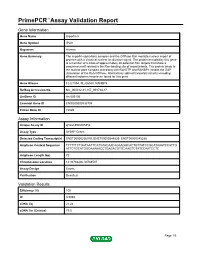
PrimePCR™Assay Validation Report Gene Information Gene Name importin 8 Gene Symbol IPO8 Organism Human Gene Summary The importin-alpha/beta complex and the GTPase Ran mediate nuclear import of proteins with a classical nuclear localization signal. The protein encoded by this gene is a member of a class of approximately 20 potential Ran targets that share a sequence motif related to the Ran-binding site of importin-beta. This protein binds to the nuclear pore complex and along with RanGTP and RANBP1 inhibits the GAP stimulation of the Ran GTPase. Alternatively spliced transcript variants encoding different isoforms have been found for this gene. Gene Aliases FLJ21954, FLJ26580, RANBP8 RefSeq Accession No. NC_000012.11, NT_009714.17 UniGene ID Hs.505136 Ensembl Gene ID ENSG00000133704 Entrez Gene ID 10526 Assay Information Unique Assay ID qHsaCED0005354 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000256079, ENST00000544829, ENST00000545286 Amplicon Context Sequence TTTTTTCTGATAATTCATGTACAACAGAAGGCACTGTTATCCGCATAAATCCATTG ATTCTGCATGGCAAAAACCTGAGACGTGCAAGTCTATCCAATCCTC Amplicon Length (bp) 72 Chromosome Location 12:30783406-30783507 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 100 R2 0.9998 cDNA Cq 21.24 cDNA Tm (Celsius) 79.5 Page 1/5 PrimePCR™Assay Validation Report gDNA Cq 24.52 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report IPO8, Human Amplification Plot Amplification of cDNA generated from -

Mouse Ipo8 Knockout Project (CRISPR/Cas9)
https://www.alphaknockout.com Mouse Ipo8 Knockout Project (CRISPR/Cas9) Objective: To create a Ipo8 knockout Mouse model (C57BL/6J) by CRISPR/Cas-mediated genome engineering. Strategy summary: The Ipo8 gene (NCBI Reference Sequence: NM_001081113 ; Ensembl: ENSMUSG00000040029 ) is located on Mouse chromosome 6. 25 exons are identified, with the ATG start codon in exon 1 and the TGA stop codon in exon 25 (Transcript: ENSMUST00000048418). Exon 2~4 will be selected as target site. Cas9 and gRNA will be co-injected into fertilized eggs for KO Mouse production. The pups will be genotyped by PCR followed by sequencing analysis. Note: Exon 2 starts from about 2.81% of the coding region. Exon 2~4 covers 13.14% of the coding region. The size of effective KO region: ~6522 bp. The KO region does not have any other known gene. Page 1 of 9 https://www.alphaknockout.com Overview of the Targeting Strategy Wildtype allele 5' gRNA region gRNA region 3' 1 2 3 4 25 Legends Exon of mouse Ipo8 Knockout region Page 2 of 9 https://www.alphaknockout.com Overview of the Dot Plot (up) Window size: 15 bp Forward Reverse Complement Sequence 12 Note: The 2000 bp section upstream of Exon 2 is aligned with itself to determine if there are tandem repeats. Tandem repeats are found in the dot plot matrix. The gRNA site is selected outside of these tandem repeats. Overview of the Dot Plot (down) Window size: 15 bp Forward Reverse Complement Sequence 12 Note: The 1322 bp section downstream of Exon 4 is aligned with itself to determine if there are tandem repeats. -

Open Data for Differential Network Analysis in Glioma
International Journal of Molecular Sciences Article Open Data for Differential Network Analysis in Glioma , Claire Jean-Quartier * y , Fleur Jeanquartier y and Andreas Holzinger Holzinger Group HCI-KDD, Institute for Medical Informatics, Statistics and Documentation, Medical University Graz, Auenbruggerplatz 2/V, 8036 Graz, Austria; [email protected] (F.J.); [email protected] (A.H.) * Correspondence: [email protected] These authors contributed equally to this work. y Received: 27 October 2019; Accepted: 3 January 2020; Published: 15 January 2020 Abstract: The complexity of cancer diseases demands bioinformatic techniques and translational research based on big data and personalized medicine. Open data enables researchers to accelerate cancer studies, save resources and foster collaboration. Several tools and programming approaches are available for analyzing data, including annotation, clustering, comparison and extrapolation, merging, enrichment, functional association and statistics. We exploit openly available data via cancer gene expression analysis, we apply refinement as well as enrichment analysis via gene ontology and conclude with graph-based visualization of involved protein interaction networks as a basis for signaling. The different databases allowed for the construction of huge networks or specified ones consisting of high-confidence interactions only. Several genes associated to glioma were isolated via a network analysis from top hub nodes as well as from an outlier analysis. The latter approach highlights a mitogen-activated protein kinase next to a member of histondeacetylases and a protein phosphatase as genes uncommonly associated with glioma. Cluster analysis from top hub nodes lists several identified glioma-associated gene products to function within protein complexes, including epidermal growth factors as well as cell cycle proteins or RAS proto-oncogenes. -

Genome-Wide Association Studies of Smooth Pursuit and Antisaccade Eye Movements in Psychotic Disorders: findings from the B-SNIP Study
OPEN Citation: Transl Psychiatry (2017) 7, e1249; doi:10.1038/tp.2017.210 www.nature.com/tp ORIGINAL ARTICLE Genome-wide association studies of smooth pursuit and antisaccade eye movements in psychotic disorders: findings from the B-SNIP study R Lencer1, LJ Mills2, N Alliey-Rodriguez3, R Shafee4,5,AMLee6, JL Reilly7, A Sprenger8, JE McDowell9, SA McCarroll4, MS Keshavan10, GD Pearlson11,12, CA Tamminga13, BA Clementz9, ES Gershon3, JA Sweeney13,14 and JR Bishop6,15 Eye movement deviations, particularly deficits of initial sensorimotor processing and sustained pursuit maintenance, and antisaccade inhibition errors, are established intermediate phenotypes for psychotic disorders. We here studied eye movement measures of 849 participants from the Bipolar-Schizophrenia Network on Intermediate Phenotypes (B-SNIP) study (schizophrenia N = 230, schizoaffective disorder N = 155, psychotic bipolar disorder N = 206 and healthy controls N = 258) as quantitative phenotypes in relation to genetic data, while controlling for genetically derived ancestry measures, age and sex. A mixed-modeling genome-wide association studies approach was used including ~ 4.4 million genotypes (PsychChip and 1000 Genomes imputation). Across participants, sensorimotor processing at pursuit initiation was significantly associated with a single nucleotide polymorphism in IPO8 (12p11.21, P =8×10− 11), whereas suggestive associations with sustained pursuit maintenance were identified with SNPs in SH3GL2 (9p22.2, P =3×10− 8). In participants of predominantly African ancestry, sensorimotor processing was also significantly associated with SNPs in PCDH12 (5q31.3, P = 1.6 × 10 − 10), and suggestive associations were observed with NRSN1 (6p22.3, P = 5.4 × 10 −8) and LMO7 (13q22.2, P = 7.3x10−8), whereas antisaccade error rate was significantly associated with a non-coding region at chromosome 7 (P = 6.5 × 10− 9). -

Research2007herschkowitzetvolume Al
Open Access Research2007HerschkowitzetVolume al. 8, Issue 5, Article R76 Identification of conserved gene expression features between comment murine mammary carcinoma models and human breast tumors Jason I Herschkowitz¤*†, Karl Simin¤‡, Victor J Weigman§, Igor Mikaelian¶, Jerry Usary*¥, Zhiyuan Hu*¥, Karen E Rasmussen*¥, Laundette P Jones#, Shahin Assefnia#, Subhashini Chandrasekharan¥, Michael G Backlund†, Yuzhi Yin#, Andrey I Khramtsov**, Roy Bastein††, John Quackenbush††, Robert I Glazer#, Powel H Brown‡‡, Jeffrey E Green§§, Levy Kopelovich, reviews Priscilla A Furth#, Juan P Palazzo, Olufunmilayo I Olopade, Philip S Bernard††, Gary A Churchill¶, Terry Van Dyke*¥ and Charles M Perou*¥ Addresses: *Lineberger Comprehensive Cancer Center. †Curriculum in Genetics and Molecular Biology, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599, USA. ‡Department of Cancer Biology, University of Massachusetts Medical School, Worcester, MA 01605, USA. reports §Department of Biology and Program in Bioinformatics and Computational Biology, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599, USA. ¶The Jackson Laboratory, Bar Harbor, ME 04609, USA. ¥Department of Genetics, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599, USA. #Department of Oncology, Lombardi Comprehensive Cancer Center, Georgetown University, Washington, DC 20057, USA. **Department of Pathology, University of Chicago, Chicago, IL 60637, USA. ††Department of Pathology, University of Utah School of Medicine, Salt Lake City, UT 84132, USA. ‡‡Baylor College of Medicine, Houston, TX 77030, USA. §§Transgenic Oncogenesis Group, Laboratory of Cancer Biology and Genetics. Chemoprevention Agent Development Research Group, National Cancer Institute, Bethesda, MD 20892, USA. Department of Pathology, Thomas Jefferson University, Philadelphia, PA 19107, USA. Section of Hematology/Oncology, Department of Medicine, Committees on Genetics and Cancer Biology, University of Chicago, Chicago, IL 60637, USA. -

HSP90-Incorporating Chaperome Networks As Biosensor for Disease-Related Pathways in Patient- Specific Midbrain Dopamine Neurons
ARTICLE DOI: 10.1038/s41467-018-06486-6 OPEN HSP90-incorporating chaperome networks as biosensor for disease-related pathways in patient- specific midbrain dopamine neurons Sarah Kishinevsky1,2,3,4, Tai Wang3, Anna Rodina3, Sun Young Chung1,2, Chao Xu3, John Philip5, Tony Taldone3, Suhasini Joshi3, Mary L. Alpaugh3,14, Alexander Bolaender3, Simon Gutbier6, Davinder Sandhu7, Faranak Fattahi1,2, Bastian Zimmer1,2, Smit K. Shah3, Elizabeth Chang5, Carmen Inda3,15, John Koren 3rd3,16, Nathalie G. Saurat1,2, Marcel Leist 6, Steven S. Gross7, Venkatraman E. Seshan8, Christine Klein9, Mark J. Tomishima1,2,10, Hediye Erdjument-Bromage11,12, Thomas A. Neubert 11,12, Ronald C. Henrickson5, 1234567890():,; Gabriela Chiosis3,13 & Lorenz Studer1,2 Environmental and genetic risk factors contribute to Parkinson’s Disease (PD) pathogenesis and the associated midbrain dopamine (mDA) neuron loss. Here, we identify early PD pathogenic events by developing methodology that utilizes recent innovations in human pluripotent stem cells (hPSC) and chemical sensors of HSP90-incorporating chaperome networks. We show that events triggered by PD-related genetic or toxic stimuli alter the neuronal proteome, thereby altering the stress-specific chaperome networks, which produce changes detected by chemical sensors. Through this method we identify STAT3 and NF-κB signaling activation as examples of genetic stress, and phospho-tyrosine hydroxylase (TH) activation as an example of toxic stress-induced pathways in PD neurons. Importantly, pharmacological inhibition of the stress chaperome network reversed abnormal phospho- STAT3 signaling and phospho-TH-related dopamine levels and rescued PD neuron viability. The use of chemical sensors of chaperome networks on hPSC-derived lineages may present a general strategy to identify molecular events associated with neurodegenerative diseases. -

Evaluation of Reference Genes and Normalization Strategy for Quantitative Real-Time PCR in Human Pancreatic Carcinoma
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Crossref Disease Markers 32 (2012) 203–210 203 DOI 10.3233/DMA-2011-0875 IOS Press Evaluation of reference genes and normalization strategy for quantitative real-time PCR in human pancreatic carcinoma Beatrice Mohelnikova-Duchonovaa,b, Martin Oliveriusc, Eva Honsovad and Pavel Souceka,∗ aDepartment of Toxicogenomics, National Institute of Public Health, Prague, Czech Republic bFirst Faculty of Medicine, Charles University in Prague, Czech Republic cDepartment of Transplantation Surgery, Institute of Clinical and Experimental Medicine, Prague, Czech Republic dDepartment of Clinical and Transplantation Pathology, Institute of Clinical and Experimental Medicine, Prague, Czech Republic Abstract. Histologically verified pairs (n = 10) of pancreatic tumors and non-neoplastic tissues were used for quantitative real-time PCR and the stability of 24 reference genes was analyzed with geNorm and NormFinder software. Raw Cq values correlated with the degree of RNA degradation. This correlation was abolished by normalization to Cq of 18S endogenous control gene. Both geNorm and NormFinder programs suggested EIF2B1, ELF1, MRPL19,andPOP4 as the same most stable genes. We have thus identified suitable reference genes for future expression studies in pancreatic carcinoma. Normalization method reducing the effects of RNA degradation on the quality of results was also developed. Keywords: Pancreas, carcinoma, transcript, quantification, reference gene, normalization 1. Introduction The relative quantification by real-time PCR method (qPCR) is adequate strategy for most purposes where Pancreatic cancer is the fourth leading cause of can- investigation of physiological changes in gene expres- cer death in the United States [1] as well as in the Czech sion levels is conducted [6–8]. -

Transcriptional Control of Autophagy–Lysosome Function Drives Pancreatic Cancer Metabolism
Transcriptional control of autophagy–lysosome function drives pancreatic cancer metabolism The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation Perera, Rushika M.; Stoykova, Svetlana; Nicolay, Brandon N.; Ross, Kenneth N.; Fitamant, Julien; Boukhali, Myriam; Lengrand, Justine et al. “Transcriptional Control of Autophagy–lysosome Function Drives Pancreatic Cancer Metabolism.” Nature 524, no. 7565 (July 2015): 361–365 © 2015 Macmillan Publishers Limited, part of Springer Nature As Published http://dx.doi.org/10.1038/nature14587 Publisher Nature Publishing Group Version Author's final manuscript Citable link http://hdl.handle.net/1721.1/109748 Terms of Use Article is made available in accordance with the publisher's policy and may be subject to US copyright law. Please refer to the publisher's site for terms of use. HHS Public Access Author manuscript Author ManuscriptAuthor Manuscript Author Nature. Manuscript Author Author manuscript; Manuscript Author available in PMC 2016 October 31. Published in final edited form as: Nature. 2015 August 20; 524(7565): 361–365. doi:10.1038/nature14587. Transcriptional control of the autophagy-lysosome system in pancreatic cancer Rushika M. Perera1,2,3, Svetlana Stoykova1,2,#, Brandon N. Nicolay1,3,#, Kenneth N. Ross1,2,3, Julien Fitamant1,2,3, Myriam Boukhali1, Justine Lengrand1,2, Vikram Deshpande3,4, Martin K. Selig4, Cristina R. Ferrone1,3,5, Jeff Settleman1, Gregory Stephanopoulos6, Nicholas J. Dyson1,3, Roberto Zoncu7, Sridhar -

Proteomics of Nucleocytoplasmic Partitioning
Available online at www.sciencedirect.com ScienceDirect Proteomics of nucleocytoplasmic partitioning Thao Nguyen, Nishant Pappireddi and Martin Wu¨ hr The partitioning of the proteome between nucleus and activation. Kinase activity can also be regulated via sub- cytoplasm affects nearly every aspect of eukaryotic biology. cellular localization: cyclin B needs to relocalize into the Despite this central role, we still have a poor understanding of nucleus to induce NE breakdown, which is required for which proteins localize in the nucleus and how this varies in the transition from G2 to mitosis [3]. Considering the different cell types and conditions. Recent advances in importance of subcellular localization in encoding impor- quantitative proteomics and high-throughput imaging are tant cellular information, it is not surprising that mis- starting to close this knowledge gap. Studies on protein regulation of nuclear transport has been associated with interaction are beginning to reveal the spectrum of cargos of multiple diseases, including developmental defects and nuclear import and export receptors.We anticipate that it will cancer [4–6]. Targeting mis-regulation of NC partitioning soon be possible to predict each protein’s nucleocytoplasmic has emerged as a promising therapeutic approach, partic- localization based on its importin/exportin interactions and its ularly for cancer treatment [7–11]. estimated diffusion rate through the nuclear pore. This insight is likely to provide us with a fundamental understanding of Many previous studies have reported this subcellular how cells use nucleocytoplasmic partitioning to encode and localization of individual proteins. Recent technological relay information. advances in methods such as mass spectrometry (MS) now allow us to look at the entire proteome at once.