Transcriptional Factor Binding Sites and Disease
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Mechanotransduction Signaling in Podocytes from Fluid Flow Shear Stress
Children's Mercy Kansas City SHARE @ Children's Mercy Manuscripts, Articles, Book Chapters and Other Papers 1-1-2018 Mechanotransduction signaling in podocytes from fluid flow shear stress. Tarak Srivastava Children's Mercy Hospital Hongying Dai Children's Mercy Hospital Daniel P. Heruth Children's Mercy Hospital Uri S. Alon Children's Mercy Hospital Robert E. Garola Children's Mercy Hospital See next page for additional authors Follow this and additional works at: https://scholarlyexchange.childrensmercy.org/papers Part of the Animal Structures Commons, Medical Biophysics Commons, Medical Genetics Commons, Nephrology Commons, and the Pathology Commons Recommended Citation Srivastava, T., Dai, H., Heruth, D. P., Alon, U. S., Garola, R. E., Zhou, J., Duncan, R. S., El-Meanawy, A., McCarthy, E. T., Sharma, R., Johnson, M. L., Savin, V. J., Sharma, M. Mechanotransduction signaling in podocytes from fluid flow shear stress. American journal of physiology. Renal physiology 314, 22-22 (2018). This Article is brought to you for free and open access by SHARE @ Children's Mercy. It has been accepted for inclusion in Manuscripts, Articles, Book Chapters and Other Papers by an authorized administrator of SHARE @ Children's Mercy. For more information, please contact [email protected]. Creator(s) Tarak Srivastava, Hongying Dai, Daniel P. Heruth, Uri S. Alon, Robert E. Garola, Jianping Zhou, R Scott Duncan, Ashraf El-Meanawy, Ellen T. McCarthy, Ram Sharma, Mark L. Johnson, Virginia J. Savin, and Mukut Sharma This article is available at SHARE @ Children's Mercy: https://scholarlyexchange.childrensmercy.org/papers/1227 Am J Physiol Renal Physiol 314: F22–F34, 2018. First published September 6, 2017; doi:10.1152/ajprenal.00325.2017. -

Differential Gene Expression of Minnesota (MN) Hygienic Honeybees (Apis Mellifera) Performing Hygienic Behavior
Minnesota State University, Mankato Cornerstone: A Collection of Scholarly and Creative Works for Minnesota State University, Mankato All Graduate Theses, Dissertations, and Other Graduate Theses, Dissertations, and Other Capstone Projects Capstone Projects 2016 Differential Gene Expression of Minnesota (MN) Hygienic Honeybees (Apis mellifera) Performing Hygienic Behavior Eric Northrup Minnesota State University Mankato Follow this and additional works at: https://cornerstone.lib.mnsu.edu/etds Part of the Bioinformatics Commons, Entomology Commons, and the Genetics Commons Recommended Citation Northrup, E. (2016). Differential Gene Expression of Minnesota (MN) Hygienic Honeybees (Apis mellifera) Performing Hygienic Behavior [Master’s thesis, Minnesota State University, Mankato]. Cornerstone: A Collection of Scholarly and Creative Works for Minnesota State University, Mankato. https://cornerstone.lib.mnsu.edu/etds/619/ This Thesis is brought to you for free and open access by the Graduate Theses, Dissertations, and Other Capstone Projects at Cornerstone: A Collection of Scholarly and Creative Works for Minnesota State University, Mankato. It has been accepted for inclusion in All Graduate Theses, Dissertations, and Other Capstone Projects by an authorized administrator of Cornerstone: A Collection of Scholarly and Creative Works for Minnesota State University, Mankato. Differential gene expression of Minnesota (MN) Hygienic honeybees (Apis mellifera) performing hygienic behavior By Eric Northrup A Thesis Submitted in Partial Fulfillment of the Requirements for the Degree of Masters of Science In Biological Sciences Minnesota State University, Mankato Mankato, Minnesota April 2016 Differential gene expression of MN Hygienic honeybees (Apis mellifera) performing hygienic behavior Eric Northrup This thesis has been examined and approved by the following members of the student’s committee. -

Watsonjn2018.Pdf (1.780Mb)
UNIVERSITY OF CENTRAL OKLAHOMA Edmond, Oklahoma Department of Biology Investigating Differential Gene Expression in vivo of Cardiac Birth Defects in an Avian Model of Maternal Phenylketonuria A THESIS SUBMITTED TO THE GRADUATE FACULTY In partial fulfillment of the requirements For the degree of MASTER OF SCIENCE IN BIOLOGY By Jamie N. Watson Edmond, OK June 5, 2018 J. Watson/Dr. Nikki Seagraves ii J. Watson/Dr. Nikki Seagraves Acknowledgements It is difficult to articulate the amount of gratitude I have for the support and encouragement I have received throughout my master’s thesis. Many people have added value and support to my life during this time. I am thankful for the education, experience, and friendships I have gained at the University of Central Oklahoma. First, I would like to thank Dr. Nikki Seagraves for her mentorship and friendship. I lucked out when I met her. I have enjoyed working on this project and I am very thankful for her support. I would like thank Thomas Crane for his support and patience throughout my master’s degree. I would like to thank Dr. Shannon Conley for her continued mentorship and support. I would like to thank Liz Bullen and Dr. Eric Howard for their training and help on this project. I would like to thank Kristy Meyer for her friendship and help throughout graduate school. I would like to thank my committee members Dr. Robert Brennan and Dr. Lilian Chooback for their advisement on this project. Also, I would like to thank the biology faculty and staff. I would like to thank the Seagraves lab members: Jailene Canales, Kayley Pate, Mckayla Muse, Grace Thetford, Kody Harvey, Jordan Guffey, and Kayle Patatanian for their hard work and support. -

Microrna-205 Directly Targets Krüppel-Like Factor 12 and Is Involved in Invasion and Apoptosis in Basal-Like Breast Carcinoma
720 INTERNATIONAL JOURNAL OF ONCOLOGY 49: 720-734, 2016 MicroRNA-205 directly targets Krüppel-like factor 12 and is involved in invasion and apoptosis in basal-like breast carcinoma BING GUAN1, QING LI2*, LI SHEN3*, QIU RAO4, YAN WANG5, YUN ZHU6, XIAO-JUN ZHOU4 and XIAO-HONG LI1 1Department of Pathology, Shanghai 6th People's Hospital Jinshan Branch, Shanghai 201599; 2Department of Pathology, Shanghai Pudong New Area People's Hospital, Shanghai 201299; 3Department of Cardiothoracic Surgery, Shanghai Children's Hospital, Shanghai 201040; 4Department of Pathology, Nanjing Jinling Hospital, Nanjing, Jiangsu 210002; 5Department of Pathology, The Second Affiliated Hospital with Nanjing Medical University, Nanjing, Jiangsu 210000; 6Department of Pathology, Jiangsu Province People's Hospital, Nanjing, Jiangsu 210029, P.R. China Received March 31, 2016; Accepted May 23, 2016 DOI: 10.3892/ijo.2016.3573 Abstract. We investigated microRNAs (miRs) specific to control (NC). Gene ontology (GO) analysis suggested miR-205 its target gene and exerting distinct biological functions for may directly targeted Krüppel-like factor 12 (KLF12; basal-like breast carcinoma (BLBC). Total RNA was extracted degree=4). Luciferase assay revealed miR-205 directly and subjected to miR microarray and bioinformatics analysis. targeted KLF12 through binding its 3'-untranslated region Based on the comprehensive analysis, expression of miRs (3'-UTR; p=0.0016). qRT-PCR and western blot analysis including its target was analyzed by quantitative reverse showed miR-205 expression was low in cells (p=0.007) and transcription-polymerase chain reaction (qRT-PCR), western tumor tissues (n=6; p=0.0074), and KLF12 RNA/protein was blot analysis and immunohistochemistry (IHC). -

Prox1regulates the Subtype-Specific Development of Caudal Ganglionic
The Journal of Neuroscience, September 16, 2015 • 35(37):12869–12889 • 12869 Development/Plasticity/Repair Prox1 Regulates the Subtype-Specific Development of Caudal Ganglionic Eminence-Derived GABAergic Cortical Interneurons X Goichi Miyoshi,1 Allison Young,1 Timothy Petros,1 Theofanis Karayannis,1 Melissa McKenzie Chang,1 Alfonso Lavado,2 Tomohiko Iwano,3 Miho Nakajima,4 Hiroki Taniguchi,5 Z. Josh Huang,5 XNathaniel Heintz,4 Guillermo Oliver,2 Fumio Matsuzaki,3 Robert P. Machold,1 and Gord Fishell1 1Department of Neuroscience and Physiology, NYU Neuroscience Institute, Smilow Research Center, New York University School of Medicine, New York, New York 10016, 2Department of Genetics & Tumor Cell Biology, St. Jude Children’s Research Hospital, Memphis, Tennessee 38105, 3Laboratory for Cell Asymmetry, RIKEN Center for Developmental Biology, Kobe 650-0047, Japan, 4Laboratory of Molecular Biology, Howard Hughes Medical Institute, GENSAT Project, The Rockefeller University, New York, New York 10065, and 5Cold Spring Harbor Laboratory, Cold Spring Harbor, New York 11724 Neurogliaform (RELNϩ) and bipolar (VIPϩ) GABAergic interneurons of the mammalian cerebral cortex provide critical inhibition locally within the superficial layers. While these subtypes are known to originate from the embryonic caudal ganglionic eminence (CGE), the specific genetic programs that direct their positioning, maturation, and integration into the cortical network have not been eluci- dated. Here, we report that in mice expression of the transcription factor Prox1 is selectively maintained in postmitotic CGE-derived cortical interneuron precursors and that loss of Prox1 impairs the integration of these cells into superficial layers. Moreover, Prox1 differentially regulates the postnatal maturation of each specific subtype originating from the CGE (RELN, Calb2/VIP, and VIP). -

And Represses TNF Gene Expression Activating Transcription Factor-2
Histone Deacetylase 3, a Class I Histone Deacetylase, Suppresses MAPK11-Mediated Activating Transcription Factor-2 Activation and Represses TNF Gene Expression This information is current as of September 25, 2021. Ulrich Mahlknecht, Jutta Will, Audrey Varin, Dieter Hoelzer and Georges Herbein J Immunol 2004; 173:3979-3990; ; doi: 10.4049/jimmunol.173.6.3979 http://www.jimmunol.org/content/173/6/3979 Downloaded from References This article cites 45 articles, 31 of which you can access for free at: http://www.jimmunol.org/content/173/6/3979.full#ref-list-1 http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication by guest on September 25, 2021 *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2004 by The American Association of Immunologists All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Histone Deacetylase 3, a Class I Histone Deacetylase, Suppresses MAPK11-Mediated Activating Transcription Factor-2 Activation and Represses TNF Gene Expression1 Ulrich Mahlknecht,2* Jutta Will,* Audrey Varin,† Dieter Hoelzer,* and Georges Herbein2† During inflammatory events, the induction of immediate-early genes, such as TNF-␣, is regulated by signaling cascades including the JAK/STAT, NF-B, and the p38 MAPK pathways, which result in phosphorylation-dependent activation of transcription factors. -

Targeting Endothelial Kruppel-Like Factor 2 (KLF2) in Arteriovenous
Targeting Endothelial Krüppel-like Factor 2 (KLF2) in Arteriovenous Fistula Maturation Failure A dissertation submitted to the Graduate School of the University of Cincinnati in partial fulfillment of the requirements for the degree of DOCTOR OF PHILOSOPHY (Ph.D.) in the Biomedical Engineering Program Department of Biomedical Engineering College of Engineering and Applied Science 2018 by Keith Louis Saum B.S., Wright State University, 2012 Dissertation Committee: Albert Phillip Owens III, Ph.D. (Committee Chair) Begona Campos-Naciff, Ph.D Christy Holland, Ph.D. Daria Narmoneva, Ph.D. Prabir Roy-Chaudhury, M.D., Ph.D Charuhas Thakar, M.D. Abstract The arteriovenous fistula (AVF) is the preferred form of vascular access for hemodialysis. However, 25-60% of AVFs fail to mature to a state suitable for clinical use, resulting in significant morbidity, mortality, and cost for end-stage renal disease (ESRD) patients. AVF maturation failure is recognized to result from changes in local hemodynamics following fistula creation which lead to venous stenosis and thrombosis. In particular, abnormal wall shear stress (WSS) is thought to be a key stimulus which alters endothelial function and promotes AVF failure. In recent years, the transcription factor Krüppel-like factor-2 (KLF2) has emerged as a key regulator of endothelial function, and reduced KLF2 expression has been shown to correlate with disturbed WSS and AVF failure. Given KLF2’s importance in regulating endothelial function, the objective of this dissertation was to investigate how KLF2 expression is regulated by the hemodynamic and uremic stimuli within AVFs and determine if loss of endothelial KLF2 is responsible for impaired endothelial function. -

Accompanies CD8 T Cell Effector Function Global DNA Methylation
Global DNA Methylation Remodeling Accompanies CD8 T Cell Effector Function Christopher D. Scharer, Benjamin G. Barwick, Benjamin A. Youngblood, Rafi Ahmed and Jeremy M. Boss This information is current as of October 1, 2021. J Immunol 2013; 191:3419-3429; Prepublished online 16 August 2013; doi: 10.4049/jimmunol.1301395 http://www.jimmunol.org/content/191/6/3419 Downloaded from Supplementary http://www.jimmunol.org/content/suppl/2013/08/20/jimmunol.130139 Material 5.DC1 References This article cites 81 articles, 25 of which you can access for free at: http://www.jimmunol.org/content/191/6/3419.full#ref-list-1 http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists by guest on October 1, 2021 • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Global DNA Methylation Remodeling Accompanies CD8 T Cell Effector Function Christopher D. Scharer,* Benjamin G. Barwick,* Benjamin A. Youngblood,*,† Rafi Ahmed,*,† and Jeremy M. -

Global Mef2 Target Gene Analysis in Skeletal and Cardiac Muscle
GLOBAL MEF2 TARGET GENE ANALYSIS IN SKELETAL AND CARDIAC MUSCLE STEPHANIE ELIZABETH WALES A DISSERTATION SUBMITTED TO THE FACULTY OF GRADUATE STUDIES IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF DOCTOR OF PHILOSOPHY GRADUATE PROGRAM IN BIOLOGY YORK UNIVERSITY TORONTO, ONTARIO FEBRUARY 2016 © Stephanie Wales 2016 ABSTRACT A loss of muscle mass or function occurs in many genetic and acquired pathologies such as heart disease, sarcopenia and cachexia which are predominantly found among the rapidly increasing elderly population. Developing effective treatments relies on understanding the genetic networks that control these disease pathways. Transcription factors occupy an essential position as regulators of gene expression. Myocyte enhancer factor 2 (MEF2) is an important transcription factor in striated muscle development in the embryo, skeletal muscle maintenance in the adult and cardiomyocyte survival and hypertrophy in the progression to heart failure. We sought to identify common MEF2 target genes in these two types of striated muscles using chromatin immunoprecipitation and next generation sequencing (ChIP-seq) and transcriptome profiling (RNA-seq). Using a cell culture model of skeletal muscle (C2C12) and primary cardiomyocytes we found 294 common MEF2A binding sites within both cell types. Individually MEF2A was recruited to approximately 2700 and 1600 DNA sequences in skeletal and cardiac muscle, respectively. Two genes were chosen for further study: DUSP6 and Hspb7. DUSP6, an ERK1/2 specific phosphatase, was negatively regulated by MEF2 in a p38MAPK dependent manner in striated muscle. Furthermore siRNA mediated gene silencing showed that MEF2D in particular was responsible for repressing DUSP6 during C2C12 myoblast differentiation. Using a p38 pharmacological inhibitor (SB 203580) we observed that MEF2D must be phosphorylated by p38 to repress DUSP6. -

Methylome and Transcriptome Maps of Human Visceral and Subcutaneous
www.nature.com/scientificreports OPEN Methylome and transcriptome maps of human visceral and subcutaneous adipocytes reveal Received: 9 April 2019 Accepted: 11 June 2019 key epigenetic diferences at Published: xx xx xxxx developmental genes Stephen T. Bradford1,2,3, Shalima S. Nair1,3, Aaron L. Statham1, Susan J. van Dijk2, Timothy J. Peters 1,3,4, Firoz Anwar 2, Hugh J. French 1, Julius Z. H. von Martels1, Brodie Sutclife2, Madhavi P. Maddugoda1, Michelle Peranec1, Hilal Varinli1,2,5, Rosanna Arnoldy1, Michael Buckley1,4, Jason P. Ross2, Elena Zotenko1,3, Jenny Z. Song1, Clare Stirzaker1,3, Denis C. Bauer2, Wenjia Qu1, Michael M. Swarbrick6, Helen L. Lutgers1,7, Reginald V. Lord8, Katherine Samaras9,10, Peter L. Molloy 2 & Susan J. Clark 1,3 Adipocytes support key metabolic and endocrine functions of adipose tissue. Lipid is stored in two major classes of depots, namely visceral adipose (VA) and subcutaneous adipose (SA) depots. Increased visceral adiposity is associated with adverse health outcomes, whereas the impact of SA tissue is relatively metabolically benign. The precise molecular features associated with the functional diferences between the adipose depots are still not well understood. Here, we characterised transcriptomes and methylomes of isolated adipocytes from matched SA and VA tissues of individuals with normal BMI to identify epigenetic diferences and their contribution to cell type and depot-specifc function. We found that DNA methylomes were notably distinct between diferent adipocyte depots and were associated with diferential gene expression within pathways fundamental to adipocyte function. Most striking diferential methylation was found at transcription factor and developmental genes. Our fndings highlight the importance of developmental origins in the function of diferent fat depots. -
Integrative Bulk and Single-Cell Profiling of Premanufacture T-Cell Populations Reveals Factors Mediating Long-Term Persistence of CAR T-Cell Therapy
Published OnlineFirst April 5, 2021; DOI: 10.1158/2159-8290.CD-20-1677 RESEARCH ARTICLE Integrative Bulk and Single-Cell Profiling of Premanufacture T-cell Populations Reveals Factors Mediating Long-Term Persistence of CAR T-cell Therapy Gregory M. Chen1, Changya Chen2,3, Rajat K. Das2, Peng Gao2, Chia-Hui Chen2, Shovik Bandyopadhyay4, Yang-Yang Ding2,5, Yasin Uzun2,3, Wenbao Yu2, Qin Zhu1, Regina M. Myers2, Stephan A. Grupp2,5, David M. Barrett2,5, and Kai Tan2,3,5 Downloaded from cancerdiscovery.aacrjournals.org on October 1, 2021. © 2021 American Association for Cancer Research. Published OnlineFirst April 5, 2021; DOI: 10.1158/2159-8290.CD-20-1677 ABSTRACT The adoptive transfer of chimeric antigen receptor (CAR) T cells represents a breakthrough in clinical oncology, yet both between- and within-patient differences in autologously derived T cells are a major contributor to therapy failure. To interrogate the molecular determinants of clinical CAR T-cell persistence, we extensively characterized the premanufacture T cells of 71 patients with B-cell malignancies on trial to receive anti-CD19 CAR T-cell therapy. We performed RNA-sequencing analysis on sorted T-cell subsets from all 71 patients, followed by paired Cellular Indexing of Transcriptomes and Epitopes (CITE) sequencing and single-cell assay for transposase-accessible chromatin sequencing (scATAC-seq) on T cells from six of these patients. We found that chronic IFN signaling regulated by IRF7 was associated with poor CAR T-cell persistence across T-cell subsets, and that the TCF7 regulon not only associates with the favorable naïve T-cell state, but is maintained in effector T cells among patients with long-term CAR T-cell persistence. -
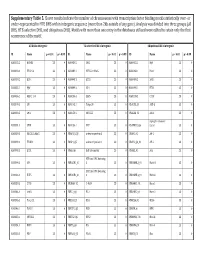
Supplementary Table 5. Clover Results Indicate the Number Of
Supplementary Table 5. Clover results indicate the number of chromosomes with transcription factor binding motifs statistically over‐ or under‐represented in HTE DHS within intergenic sequence (more than 2kb outside of any gene). Analysis was divided into three groups (all DHS, HTE‐selective DHS, and ubiquitous DHS). Motifs with more than one entry in the databases utilized were edited to retain only the first occurrence of the motif. All DHS x Intergenic TEselective DHS x Intergenic Ubiquitous DHS x Intergenic ID Name p < 0.01 p > 0.99 ID Name p < 0.01 p > 0.99 ID Name p < 0.01 p > 0.99 MA0002.2 RUNX1 23 0 MA0080.2 SPI1 23 0 MA0055.1 Myf 23 0 MA0003.1 TFAP2A 23 0 MA0089.1 NFE2L1::MafG 23 0 MA0068.1 Pax4 23 0 MA0039.2 Klf4 23 0 MA0098.1 ETS1 23 0 MA0080.2 SPI1 23 0 MA0055.1 Myf 23 0 MA0099.2 AP1 23 0 MA0098.1 ETS1 23 0 MA0056.1 MZF1_1‐4 23 0 MA0136.1 ELF5 23 0 MA0139.1 CTCF 23 0 MA0079.2 SP1 23 0 MA0145.1 Tcfcp2l1 23 0 V$ALX3_01 ALX‐3 23 0 MA0080.2 SPI1 23 0 MA0150.1 NFE2L2 23 0 V$ALX4_02 Alx‐4 23 0 myocyte enhancer MA0081.1 SPIB 23 0 MA0156.1 FEV 23 0 V$AMEF2_Q6 factor 23 0 MA0089.1 NFE2L1::MafG 23 0 V$AP1FJ_Q2 activator protein 1 23 0 V$AP1_01 AP‐1 23 0 MA0090.1 TEAD1 23 0 V$AP4_Q5 activator protein 4 23 0 V$AP2_Q6_01 AP‐2 23 0 MA0098.1 ETS1 23 0 V$AR_Q6 half‐site matrix 23 0 V$ARX_01 Arx 23 0 BTB and CNC homolog MA0099.2 AP1 23 0 V$BACH1_01 1 23 0 V$BARHL1_01 Barhl‐1 23 0 BTB and CNC homolog MA0136.1 ELF5 23 0 V$BACH2_01 2 23 0 V$BARHL2_01 Barhl2 23 0 MA0139.1 CTCF 23 0 V$CMAF_02 C‐MAF 23 0 V$BARX1_01 Barx1 23 0 MA0144.1 Stat3 23 0