Tools and Technologies to Characterize Isoforms at Proteome-Scale
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Linked Mental Retardation Detected by Array CGH
JMG Online First, published on September 16, 2005 as 10.1136/jmg.2005.036178 J Med Genet: first published as 10.1136/jmg.2005.036178 on 16 September 2005. Downloaded from Chromosomal copy number changes in patients with non-syndromic X- linked mental retardation detected by array CGH D Lugtenberg1, A P M de Brouwer1, T Kleefstra1, A R Oudakker1, S G M Frints2, C T R M Schrander- Stumpel2, J P Fryns3, L R Jensen4, J Chelly5, C Moraine6, G Turner7, J A Veltman1, B C J Hamel1, B B A de Vries1, H van Bokhoven1, H G Yntema1 1Department of Human Genetics, Radboud University Nijmegen Medical Centre, Nijmegen, The Netherlands; 2Department of Clinical Genetics, University Hospital Maastricht, Maastricht, The Netherlands; 3Center for Human Genetics, University of Leuven, Leuven, Belgium; 4Max Planck Institute for Molecular Genetics, Berlin, Germany; 5INSERM 129-ICGM, Faculté de Médecine Cochin, Paris, France; 6Service de Génétique et INSERM U316, Hôpital Bretonneau, Tours, France; 7 GOLD Program, Hunter Genetics, University of Newcastle, Callaghan, New South Wales 2308, Australia http://jmg.bmj.com/ Corresponding author: on October 2, 2021 by guest. Protected copyright. Helger G. Yntema, PhD Department of Human Genetics Radboud University Nijmegen Medical Centre P.O. Box 9101 6500 HB Nijmegen The Netherlands E-mail: [email protected] tel: +31-24-3613799 fax: +31-24-3616658 1 Copyright Article author (or their employer) 2005. Produced by BMJ Publishing Group Ltd under licence. J Med Genet: first published as 10.1136/jmg.2005.036178 on 16 September 2005. Downloaded from ABSTRACT Introduction: Several studies have shown that array based comparative genomic hybridization (array CGH) is a powerful tool for the detection of copy number changes in the genome of individuals with a congenital disorder. -

Nuclear Organization and the Epigenetic Landscape of the Mus Musculus X-Chromosome Alicia Liu University of Connecticut - Storrs, [email protected]
University of Connecticut OpenCommons@UConn Doctoral Dissertations University of Connecticut Graduate School 8-9-2019 Nuclear Organization and the Epigenetic Landscape of the Mus musculus X-Chromosome Alicia Liu University of Connecticut - Storrs, [email protected] Follow this and additional works at: https://opencommons.uconn.edu/dissertations Recommended Citation Liu, Alicia, "Nuclear Organization and the Epigenetic Landscape of the Mus musculus X-Chromosome" (2019). Doctoral Dissertations. 2273. https://opencommons.uconn.edu/dissertations/2273 Nuclear Organization and the Epigenetic Landscape of the Mus musculus X-Chromosome Alicia J. Liu, Ph.D. University of Connecticut, 2019 ABSTRACT X-linked imprinted genes have been hypothesized to contribute parent-of-origin influences on social cognition. A cluster of imprinted genes Xlr3b, Xlr4b, and Xlr4c, implicated in cognitive defects, are maternally expressed and paternally silent in the murine brain. These genes defy classic mechanisms of autosomal imprinting, suggesting a novel method of imprinted gene regulation. Using Xlr3b and Xlr4c as bait, this study uses 4C-Seq on neonatal whole brain of a 39,XO mouse model, to provide the first in-depth analysis of chromatin dynamics surrounding an imprinted locus on the X-chromosome. Significant differences in long-range contacts exist be- tween XM and XP monosomic samples. In addition, XM interaction profiles contact a greater number of genes linked to cognitive impairment, abnormality of the nervous system, and abnormality of higher mental function. This is not a pattern that is unique to the imprinted Xlr3/4 locus. Additional Alicia J. Liu - University of Connecticut - 2019 4C-Seq experiments show that other genes on the X-chromosome, implicated in intellectual disability and/or ASD, also produce more maternal contacts to other X-linked genes linked to cognitive impairment. -

Location Analysis of Estrogen Receptor Target Promoters Reveals That
Location analysis of estrogen receptor ␣ target promoters reveals that FOXA1 defines a domain of the estrogen response Jose´ e Laganie` re*†, Genevie` ve Deblois*, Ce´ line Lefebvre*, Alain R. Bataille‡, Franc¸ois Robert‡, and Vincent Gigue` re*†§ *Molecular Oncology Group, Departments of Medicine and Oncology, McGill University Health Centre, Montreal, QC, Canada H3A 1A1; †Department of Biochemistry, McGill University, Montreal, QC, Canada H3G 1Y6; and ‡Laboratory of Chromatin and Genomic Expression, Institut de Recherches Cliniques de Montre´al, Montreal, QC, Canada H2W 1R7 Communicated by Ronald M. Evans, The Salk Institute for Biological Studies, La Jolla, CA, July 1, 2005 (received for review June 3, 2005) Nuclear receptors can activate diverse biological pathways within general absence of large scale functional data linking these putative a target cell in response to their cognate ligands, but how this binding sites with gene expression in specific cell types. compartmentalization is achieved at the level of gene regulation is Recently, chromatin immunoprecipitation (ChIP) has been used poorly understood. We used a genome-wide analysis of promoter in combination with promoter or genomic DNA microarrays to occupancy by the estrogen receptor ␣ (ER␣) in MCF-7 cells to identify loci recognized by transcription factors in a genome-wide investigate the molecular mechanisms underlying the action of manner in mammalian cells (20–24). This technology, termed 17-estradiol (E2) in controlling the growth of breast cancer cells. ChIP-on-chip or location analysis, can therefore be used to deter- We identified 153 promoters bound by ER␣ in the presence of E2. mine the global gene expression program that characterize the Motif-finding algorithms demonstrated that the estrogen re- action of a nuclear receptor in response to its natural ligand. -
![Downloaded from [266]](https://docslib.b-cdn.net/cover/7352/downloaded-from-266-347352.webp)
Downloaded from [266]
Patterns of DNA methylation on the human X chromosome and use in analyzing X-chromosome inactivation by Allison Marie Cotton B.Sc., The University of Guelph, 2005 A THESIS SUBMITTED IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF DOCTOR OF PHILOSOPHY in The Faculty of Graduate Studies (Medical Genetics) THE UNIVERSITY OF BRITISH COLUMBIA (Vancouver) January 2012 © Allison Marie Cotton, 2012 Abstract The process of X-chromosome inactivation achieves dosage compensation between mammalian males and females. In females one X chromosome is transcriptionally silenced through a variety of epigenetic modifications including DNA methylation. Most X-linked genes are subject to X-chromosome inactivation and only expressed from the active X chromosome. On the inactive X chromosome, the CpG island promoters of genes subject to X-chromosome inactivation are methylated in their promoter regions, while genes which escape from X- chromosome inactivation have unmethylated CpG island promoters on both the active and inactive X chromosomes. The first objective of this thesis was to determine if the DNA methylation of CpG island promoters could be used to accurately predict X chromosome inactivation status. The second objective was to use DNA methylation to predict X-chromosome inactivation status in a variety of tissues. A comparison of blood, muscle, kidney and neural tissues revealed tissue-specific X-chromosome inactivation, in which 12% of genes escaped from X-chromosome inactivation in some, but not all, tissues. X-linked DNA methylation analysis of placental tissues predicted four times higher escape from X-chromosome inactivation than in any other tissue. Despite the hypomethylation of repetitive elements on both the X chromosome and the autosomes, no changes were detected in the frequency or intensity of placental Cot-1 holes. -

A Private 16Q24.2Q24.3 Microduplication in a Boy with Intellectual Disability, Speech Delay and Mild Dysmorphic Features
G C A T T A C G G C A T genes Article A Private 16q24.2q24.3 Microduplication in a Boy with Intellectual Disability, Speech Delay and Mild Dysmorphic Features Orazio Palumbo * , Pietro Palumbo , Ester Di Muro, Luigia Cinque, Antonio Petracca, Massimo Carella and Marco Castori Division of Medical Genetics, Fondazione IRCCS-Casa Sollievo della Sofferenza, San Giovanni Rotondo, 71013 Foggia, Italy; [email protected] (P.P.); [email protected] (E.D.M.); [email protected] (L.C.); [email protected] (A.P.); [email protected] (M.C.); [email protected] (M.C.) * Correspondence: [email protected]; Tel.: +39-088-241-6350 Received: 5 June 2020; Accepted: 24 June 2020; Published: 26 June 2020 Abstract: No data on interstitial microduplications of the 16q24.2q24.3 chromosome region are available in the medical literature and remain extraordinarily rare in public databases. Here, we describe a boy with a de novo 16q24.2q24.3 microduplication at the Single Nucleotide Polymorphism (SNP)-array analysis spanning ~2.2 Mb and encompassing 38 genes. The patient showed mild-to-moderate intellectual disability, speech delay and mild dysmorphic features. In DECIPHER, we found six individuals carrying a “pure” overlapping microduplication. Although available data are very limited, genomic and phenotype comparison of our and previously annotated patients suggested a potential clinical relevance for 16q24.2q24.3 microduplication with a variable and not (yet) recognizable phenotype predominantly affecting cognition. Comparing the cytogenomic data of available individuals allowed us to delineate the smallest region of overlap involving 14 genes. Accordingly, we propose ANKRD11, CDH15, and CTU2 as candidate genes for explaining the related neurodevelopmental manifestations shared by these patients. -

At Elevated Temperatures, Heat Shock Protein Genes Show Altered Ratios Of
EXPERIMENTAL AND THERAPEUTIC MEDICINE 22: 900, 2021 At elevated temperatures, heat shock protein genes show altered ratios of different RNAs and expression of new RNAs, including several novel HSPB1 mRNAs encoding HSP27 protein isoforms XIA GAO1,2, KEYIN ZHANG1,2, HAIYAN ZHOU3, LUCAS ZELLMER4, CHENGFU YUAN5, HAI HUANG6 and DEZHONG JOSHUA LIAO2,6 1Department of Pathology, Guizhou Medical University Hospital; 2Key Lab of Endemic and Ethnic Diseases of The Ministry of Education of China in Guizhou Medical University; 3Clinical Research Center, Guizhou Medical University Hospital, Guiyang, Guizhou 550004, P.R. China; 4Masonic Cancer Center, University of Minnesota, Minneapolis, MN 55455, USA; 5Department of Biochemistry, China Three Gorges University, Yichang, Hubei 443002; 6Center for Clinical Laboratories, Guizhou Medical University Hospital, Guiyang, Guizhou 550004, P.R. China Received December 16, 2020; Accepted May 10, 2021 DOI: 10.3892/etm.2021.10332 Abstract. Heat shock proteins (HSP) serve as chaperones genes may engender multiple protein isoforms. These results to maintain the physiological conformation and function of collectively suggested that, besides increasing their expres‑ numerous cellular proteins when the ambient temperature is sion, certain HSP and associated genes also use alternative increased. To determine how accurate the general assumption transcription start sites to produce multiple RNA transcripts that HSP gene expression is increased in febrile situations is, and use alternative splicing of a transcript to produce multiple the RNA levels of the HSF1 (heat shock transcription factor 1) mature RNAs, as important mechanisms for responding to an gene and certain HSP genes were determined in three cell increased ambient temperature in vitro. lines cultured at 37˚C or 39˚C for three days. -

Proteomic Signatures of Serum Albumin
Proteomic signatures of serum albumin- bound proteins from stroke patients with and without endovascular closure of PFO are significantly different and suggest a novel mechanism for cholesterol efflux The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Lopez, M. F., B. Krastins, D. A. Sarracino, G. Byram, M. S. Vogelsang, A. Prakash, S. Peterman, et al. 2015. “Proteomic signatures of serum albumin-bound proteins from stroke patients with and without endovascular closure of PFO are significantly different and suggest a novel mechanism for cholesterol efflux.” Clinical Proteomics 12 (1): 2. doi:10.1186/1559-0275-12-2. http:// dx.doi.org/10.1186/1559-0275-12-2. Published Version doi:10.1186/1559-0275-12-2 Citable link http://nrs.harvard.edu/urn-3:HUL.InstRepos:14065419 Terms of Use This article was downloaded from Harvard University’s DASH repository, and is made available under the terms and conditions applicable to Other Posted Material, as set forth at http:// nrs.harvard.edu/urn-3:HUL.InstRepos:dash.current.terms-of- use#LAA Lopez et al. Clinical Proteomics 2015, 12:2 http://www.clinicalproteomicsjournal.com/content/12/1/2 CLINICAL PROTEOMICS RESEARCH Open Access Proteomic signatures of serum albumin-bound proteins from stroke patients with and without endovascular closure of PFO are significantly different and suggest a novel mechanism for cholesterol efflux Mary F Lopez1*, Bryan Krastins1, David A Sarracino1, Gregory Byram1, Maryann S Vogelsang1, Amol Prakash1, Scott Peterman1, Shadab Ahmad1, Gouri Vadali1, Wenjun Deng2, Ignacio Inglessis2, Tom Wickham2, Kathleen Feeney2, G William Dec2, Igor Palacios2, Ferdinando S Buonanno2, Eng H Lo2 and MingMing Ning2 Abstract Background: The anatomy of PFO suggests that it can allow thrombi and potentially harmful circulatory factors to travel directly from the venous to the arterial circulation – altering circulatory phenotype. -

Proteograph Platform Delivers Unbiased
Application of the Proteograph™ Product Suite to the Identification of Differential Protein Isoform Plasma Abundance in Early Lung Cancer vs. Healthy Controls Asim Siddiqui*, John E. Blume, Margaret K. R. Donovan, Marwin Ko, Ryan W. Benz, Theodore L. Platt, Juan C. Cuevas, Serafim Batzoglou and Omid C. Farokhzad Proteograph Platform Delivers Unbiased, Utilizing Proteograph Platform to Interrogate Protein Isoforms in a Proteograph Data Sheds Light on Biological Deep and Rapid Proteomics at Scale Non-small Cell Lung Cancer (NSCLC) Plasma Proteome Study Consequences of Protein Isoforms The ~20,000 genes in the human genome encode over one million Proteograph Platform Enables Deep and Unbiased Plasma Proteomics BMP1 Shows Differential Isoform Abundance Pattern protein variants, because of alternative splice forms, allelic variation and protein modifications. Though large-scale genomics studies Percentage of Samples Number of Peptides Detected The lowest abundant putative protein isoform, BMP1 from the list of have expanded our understanding of cancer biology through in Which Protein Group Detected per Protein candidate proteins identified in this study, comprises four protein analysis of both tissue and biofluids, similarly-scaled unbiased, deep 1A 1B 1C coding isoforms. Two of these isoforms are substantially longer n=80 proteomics studies of biofluids have remained impractical due to 2499 (~400-800 residues) than the other two isoforms covering additional Median 8 complexity of workflows. We have previously described the exons. Peptides mapping to exons that cover all four protein isoforms Proteograph Product Suite, a novel platform that leverages the n=61 (5A) have higher abundance in cancer relative to controls, whereas nano-bio interactions of nanoparticles for deep and unbiased 1992 peptides mapping to exons that cover only the two longer isoforms proteomic sampling at scale. -

Post-Translational Modifications - Essential for Protein Regulation Related Publications AUGUST Research Tools 2015
CYTOSKELETON NEWS NEWS FROM CYTOSKELETON INC. this issue Post-translational Modifications - Essential for Protein Regulation Related Publications AUGUST Research Tools 2015 Meetings Post-translational Modifications - Essential for Protein Regulation To maintain homeostasis, cells need to respond to changes in the SUMOylation, where the former plays a prominent role in protein 2015 European Cytoskeletal intracellular and extracellular milieu. Some of the changes have to degradation through proteasomal pathway7 and the latter is News Forum Meeting be acted upon quickly to avoid detrimental effects that can lead important for regulating cellular processes, including cell cycle, Aug 30-Sept 4 2015 to cell damage or even death. One way that cells act is through apoptosis, DNA repair, and signal transduction pathways8. Ljubljana, Slovenia protein post-translational modifications (PTMs) which enable cells to respond quickly to various changes by influencing numerous PTM crosstalk – Adding layers of complexity to the proteomic network The Triangle Cytoskeleton Meeting properties of proteins including enzymatic activity, protein Sept 21st, 2015 interactions, and subcellular location. Advanced developments PTM crosstalk has gained increasing interest in recent proteomics Saxapahawm NC in mass-spectrometry (MS) methods allow the identification of studies. Two independent groups employed bioinformatics and PTMs on a large scale. Today, more than 200 PTMs are known1,2, database analyses of protein sequences and have identified Neuroscience 2015 ranging from small chemical modifications (e.g., phosphorylation “hotspots” where PTMs of different types are located within a and acetylation) close proximity, about 15 amino acids apart9,10. Some of these Oct 17-21, 2015 NIH3T3 +/- pervanadate 1,2 Ljubljana, Slovenia to the addition of IP: APY03-bead “hotspots” have been reported in the literature to crosstalk . -

Datasheet: MCA1370Z Product Details
Datasheet: MCA1370Z Description: HAMSTER ANTI MOUSE CD31:Preservative Free Specificity: CD31 Other names: PECAM-1 Format: Preservative Free Product Type: Monoclonal Antibody Clone: 2H8 Isotype: IgG Quantity: 0.5 mg Product Details Applications This product has been reported to work in the following applications. This information is derived from testing within our laboratories, peer-reviewed publications or personal communications from the originators. Please refer to references indicated for further information. For general protocol recommendations, please visit www.bio-rad-antibodies.com/protocols. Yes No Not Determined Suggested Dilution Flow Cytometry 0.1ug/ml Immunohistology - Frozen Immunohistology - Paraffin ELISA Immunoprecipitation Western Blotting Immunofluorescence Functional Assays Where this antibody has not been tested for use in a particular technique this does not necessarily exclude its use in such procedures. Suggested working dilutions are given as a guide only. It is recommended that the user titrates the antibody for use in their own system using appropriate negative/positive controls. Target Species Mouse Product Form Purified IgG - liquid Preparation Purified IgG prepared by Caprylic Acid Precipitation Buffer Solution Phosphate buffered saline Preservative None present. Stabilisers Sterile filtered. Approx. Protein IgG concentration 0.5 mg/ml Concentrations Immunogen D10.G4.1 cells (Kaye et al. 1984). Page 1 of 3 External Database Links UniProt: Q08481 Related reagents Entrez Gene: 18613 Pecam1 Related reagents Synonyms Pecam, Pecam-1 Fusion Partners Splenic lymphocytes from an immunized Armenian hamster were fused with cells from the SP2/0 murine myeloma. Specificity Hamster anti Mouse CD31 monoclonal antibody, clone 2H8 recognizes murine CD31, also known as Platelet endothelial cell adhesion molecule or PECAM-1. -

E-Mutpath: Computational Modelling Reveals the Functional Landscape of Genetic Mutations Rewiring Interactome Networks
bioRxiv preprint doi: https://doi.org/10.1101/2020.08.22.262386; this version posted August 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. e-MutPath: Computational modelling reveals the functional landscape of genetic mutations rewiring interactome networks Yongsheng Li1, Daniel J. McGrail1, Brandon Burgman2,3, S. Stephen Yi2,3,4,5 and Nidhi Sahni1,6,7,8,* 1Department oF Systems Biology, The University oF Texas MD Anderson Cancer Center, Houston, TX 77030, USA 2Department oF Oncology, Livestrong Cancer Institutes, Dell Medical School, The University oF Texas at Austin, Austin, TX 78712, USA 3Institute For Cellular and Molecular Biology (ICMB), The University oF Texas at Austin, Austin, TX 78712, USA 4Institute For Computational Engineering and Sciences (ICES), The University oF Texas at Austin, Austin, TX 78712, USA 5Department oF Biomedical Engineering, Cockrell School of Engineering, The University oF Texas at Austin, Austin, TX 78712, USA 6Department oF Epigenetics and Molecular Carcinogenesis, The University oF Texas MD Anderson Science Park, Smithville, TX 78957, USA 7Department oF BioinFormatics and Computational Biology, The University oF Texas MD Anderson Cancer Center, Houston, TX 77030, USA 8Program in Quantitative and Computational Biosciences (QCB), Baylor College oF Medicine, Houston, TX 77030, USA *To whom correspondence should be addressed. Nidhi Sahni. Tel: +1 512 2379506; Email: [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.08.22.262386; this version posted August 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. -
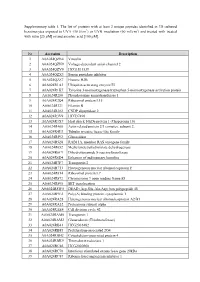
Supplementary Table 1. the List of Proteins with at Least 2 Unique
Supplementary table 1. The list of proteins with at least 2 unique peptides identified in 3D cultured keratinocytes exposed to UVA (30 J/cm2) or UVB irradiation (60 mJ/cm2) and treated with treated with rutin [25 µM] or/and ascorbic acid [100 µM]. Nr Accession Description 1 A0A024QZN4 Vinculin 2 A0A024QZN9 Voltage-dependent anion channel 2 3 A0A024QZV0 HCG1811539 4 A0A024QZX3 Serpin peptidase inhibitor 5 A0A024QZZ7 Histone H2B 6 A0A024R1A3 Ubiquitin-activating enzyme E1 7 A0A024R1K7 Tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein 8 A0A024R280 Phosphoserine aminotransferase 1 9 A0A024R2Q4 Ribosomal protein L15 10 A0A024R321 Filamin B 11 A0A024R382 CNDP dipeptidase 2 12 A0A024R3V9 HCG37498 13 A0A024R3X7 Heat shock 10kDa protein 1 (Chaperonin 10) 14 A0A024R408 Actin related protein 2/3 complex, subunit 2, 15 A0A024R4U3 Tubulin tyrosine ligase-like family 16 A0A024R592 Glucosidase 17 A0A024R5Z8 RAB11A, member RAS oncogene family 18 A0A024R652 Methylenetetrahydrofolate dehydrogenase 19 A0A024R6C9 Dihydrolipoamide S-succinyltransferase 20 A0A024R6D4 Enhancer of rudimentary homolog 21 A0A024R7F7 Transportin 2 22 A0A024R7T3 Heterogeneous nuclear ribonucleoprotein F 23 A0A024R814 Ribosomal protein L7 24 A0A024R872 Chromosome 9 open reading frame 88 25 A0A024R895 SET translocation 26 A0A024R8W0 DEAD (Asp-Glu-Ala-Asp) box polypeptide 48 27 A0A024R9E2 Poly(A) binding protein, cytoplasmic 1 28 A0A024RA28 Heterogeneous nuclear ribonucleoprotein A2/B1 29 A0A024RA52 Proteasome subunit alpha 30 A0A024RAE4 Cell division cycle 42 31