Anti-UBQLN2 Antibody
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

UBQLN2 Polyclonal Antibody C-Terminal Ubiquitin-Associated Domain
UBQLN2 polyclonal antibody C-terminal ubiquitin-associated domain. They physically associate with both proteasomes and ubiquitin ligases, Catalog Number: PAB1699 and thus thought to functionally link the ubiquitination machinery to the proteasome to affect in vivo protein Regulatory Status: For research use only (RUO) degradation. This ubiquilin has also been shown to bind the ATPase domain of the Hsp70-like Stch protein. Product Description: Rabbit polyclonal antibody raised [provided by RefSeq] against synthetic peptide of UBQLN2. References: Immunogen: A synthetic peptide (conjugated with KLH) 1. Structural studies of the interaction between ubiquitin corresponding to C-terminus of human UBQLN2. family proteins and proteasome subunit S5a. Walters KJ, Kleijnen MF, Goh AM, Wagner G, Howley PM. Host: Rabbit Biochemistry. 2002 Feb 12;41(6):1767-77. 2. The hPLIC proteins may provide a link between the Reactivity: Human ubiquitination machinery and the proteasome. Kleijnen Applications: WB-Ce MF, Shih AH, Zhou P, Kumar S, Soccio RE, Kedersha (See our web site product page for detailed applications NL, Gill G, Howley PM. Mol Cell. 2000 Aug;6(2):409-19. information) 3. Selection system for genes encoding nuclear-targeted proteins. Ueki N, Oda T, Kondo M, Yano K, Noguchi T, Protocols: See our web site at Muramatsu M. Nat Biotechnol. 1998 http://www.abnova.com/support/protocols.asp or product Dec;16(13):1338-42. page for detailed protocols Form: Liquid Purification: Protein G purification Recommend Usage: Western Blot (1:1000) The optimal working dilution should be determined by the end user. Storage Buffer: In PBS (0.09% sodium azide) Storage Instruction: Store at 4°C. -

Machine Learning Models for Predicting Protein Condensate Formation from Sequence Determinants and Embeddings
bioRxiv preprint doi: https://doi.org/10.1101/2020.10.26.354753; this version posted October 26, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Machine learning models for predicting protein condensate formation from sequence determinants and embeddings Kadi L. Saar,1;2;† Alexey S. Morgunov,1;3;† Runzhang Qi,1 William E. Arter,1 Georg Krainer,1 Alpha A. Lee2 & Tuomas P. J. Knowles1;2;∗ 1 Department of Chemistry, University of Cambridge, Lensfield Road, Cambridge CB2 1EW, UK 2 Cavendish Laboratory, Department of Physics, University of Cambridge, J J Thomson Ave, Cambridge CB3 0HE, UK 3 Fluidic Analytics, Unit A, The Paddocks Business Centre, Cherry Hinton Rd, Cambridge CB1 8DH, UK † These authors contributed equally ∗ To whom correspondence should be addressed: [email protected] Abstract Intracellular phase separation of proteins into biomolecular condensates is increasingly recognised as an important phenomenon for cellular compartmentalisation and regulation of biological function. Different hypotheses about the parameters that determine the ten- dency of proteins to form condensates have been proposed with some of them probed ex- perimentally through the use of constructs generated by sequence alterations. To broaden the scope of these observations, here, we established an in silico strategy for understanding on a global level the associations between protein sequence and condensate formation, and used this information to construct machine learning classifiers for predicting liquid–liquid phase separation (LLPS) from protein sequence. -

(Stch) Gene Reduces Prion Disease Incubation Time in Mice
Overexpression of the Hspa13 (Stch) gene reduces prion disease incubation time in mice Julia Grizenkovaa,b, Shaheen Akhtara,b, Holger Hummericha,b, Andrew Tomlinsona,b, Emmanuel A. Asantea,b, Adam Wenborna,b, Jérémie Fizeta,b, Mark Poultera,b, Frances K. Wisemanb, Elizabeth M. C. Fisherb, Victor L. J. Tybulewiczc, Sebastian Brandnerb, John Collingea,b, and Sarah E. Lloyda,b,1 aMedical Research Council (MRC) Prion Unit and bDepartment of Neurodegenerative Disease, University College London (UCL) Institute of Neurology, London WC1N 3BG, United Kingdom; and cDivision of Immune Cell Biology, MRC National Institute for Medical Research, London NW7 1AA, United Kingdom Edited by Reed B. Wickner, National Institutes of Health, Bethesda, MD, and approved July 10, 2012 (received for review May 28, 2012) Prion diseases are fatal neurodegenerative disorders that include way crosses and a heterogeneous cross (12–14). In these studies, bovine spongiform encephalopathy (BSE) and scrapie in animals the underlying functional polymorphism is unknown but could be and Creutzfeldt-Jakob disease (CJD) in humans. They are character- an amino acid change within the coding region of a protein or ized by long incubation periods, variation in which is determined by splicing variant or may occur within noncoding sequences such as many factors including genetic background. In some cases it is pos- untranslated regions, promoters, or other regulatory regions sible that incubation time may be directly correlated to the level of thereby influencing the pattern or level of expression. Although gene expression. To test this hypothesis, we combined incubation incubation time is a polygenic trait, it is possible that in some cases, time data from five different inbred lines of mice with quantitative there may be a direct correlation between incubation time and gene expression profiling in normal brains and identified five genes individual gene expression level. -

The Ubiquitin Proteasome System in Neuromuscular Disorders: Moving Beyond Movement
International Journal of Molecular Sciences Review The Ubiquitin Proteasome System in Neuromuscular Disorders: Moving Beyond Movement 1, , 2, 3,4 Sara Bachiller * y , Isabel M. Alonso-Bellido y , Luis Miguel Real , Eva María Pérez-Villegas 5 , José Luis Venero 2 , Tomas Deierborg 1 , José Ángel Armengol 5 and Rocío Ruiz 2 1 Experimental Neuroinflammation Laboratory, Department of Experimental Medical Science, Lund University, Sölvegatan 19, 221 84 Lund, Sweden; [email protected] 2 Departamento de Bioquímica y Biología Molecular, Facultad de Farmacia, Universidad de Sevilla/Instituto de Biomedicina de Sevilla-Hospital Universitario Virgen del Rocío/CSIC/Universidad de Sevilla, 41012 Sevilla, Spain; [email protected] (I.M.A.-B.); [email protected] (J.L.V.); [email protected] (R.R.) 3 Unidad Clínica de Enfermedades Infecciosas, Hospital Universitario de Valme, 41014 Sevilla, Spain; [email protected] 4 Departamento de Especialidades Quirúrgicas, Bioquímica e Inmunología, Facultad de Medicina, 29071 Universidad de Málaga, Spain 5 Departamento de Fisiología, Anatomía y Biología Celular, Universidad Pablo de Olavide, 41013 Sevilla, Spain; [email protected] (E.M.P.-V.); [email protected] (J.Á.A.) * Correspondence: [email protected] These authors contributed equally to the work. y Received: 14 July 2020; Accepted: 31 August 2020; Published: 3 September 2020 Abstract: Neuromuscular disorders (NMDs) affect 1 in 3000 people worldwide. There are more than 150 different types of NMDs, where the common feature is the loss of muscle strength. These disorders are classified according to their neuroanatomical location, as motor neuron diseases, peripheral nerve diseases, neuromuscular junction diseases, and muscle diseases. Over the years, numerous studies have pointed to protein homeostasis as a crucial factor in the development of these fatal diseases. -

Development and Validation of a Serum Biomarker Panel for The
Journal of Cancer 2017, Vol. 8 2346 Ivyspring International Publisher Journal of Cancer 2017; 8(12): 2346-2355. doi: 10.7150/jca.19465 Research Paper Development and Validation of a Serum Biomarker Panel for the Detection of Esophageal Squamous Cell Carcinoma through RNA Transcriptome Sequencing Shan Xing1,2†, Xin Zheng1,2†, Li-Qiang Wei4†, Shi-Jian Song5, Dan Liu1,3, Ning Xue1,3, Xiao-Min Liu1,2, Mian-Tao Wu1,2, Qian Zhong1,3, Chu-Mei Huang6, Mu-Sheng Zeng1,3, Wan-Li Liu1,2 1. State Key Laboratory of Oncology in Southern China, Collaborative Innovation Center for Cancer Medicine, Guangzhou, China 2. Department of Clinical Laboratory, Sun Yat-sen University Cancer Center, Guangzhou, China 3. Department of Experimental Research, Sun Yat-sen University cancer center, Guangzhou, China 4. Department of Clinical Laboratory, Shaanxi Provincial People’s Hospital, Xian, China 5. Guangdong Experimental High School, Guangzhou, China 6. Department of Laboratory Medicine, Sun Yat-sen University First Affiliated Hospital, Guangzhou, China † Xing S, Zheng X and Wei LQ contributed equally to this work Corresponding authors: Wan-Li Liu. Email: [email protected]; Phone: 86-020-87343196. Department of Clinical Laboratory, Sun Yat-sen University Cancer Center, 651 Dongfeng Road East,Guangzhou 510060, P.R. China. Mu-Sheng Zeng; Email: [email protected] Phone: +86-020-87343192. Department of Experimental Research, Sun Yat-sen University cancer center, 651 Dongfeng Road East,Guangzhou 510060, P.R.China © Ivyspring International Publisher. This is an open access article distributed under the terms of the Creative Commons Attribution (CC BY-NC) license (https://creativecommons.org/licenses/by-nc/4.0/). -

Aggresomal Sequestration and STUB1-Mediated Ubiquitylation During Mammalian Proteaphagy of Inhibited Proteasomes
Aggresomal sequestration and STUB1-mediated ubiquitylation during mammalian proteaphagy of inhibited proteasomes Won Hoon Choia,b, Yejin Yuna,b, Seoyoung Parka,c, Jun Hyoung Jeona,b, Jeeyoung Leea,b, Jung Hoon Leea,c, Su-A Yangd, Nak-Kyoon Kime, Chan Hoon Jungb, Yong Tae Kwonb, Dohyun Hanf, Sang Min Lime, and Min Jae Leea,b,c,1 aDepartment of Biochemistry and Molecular Biology, Seoul National University College of Medicine, 03080 Seoul, Korea; bDepartment of Biomedical Sciences, Seoul National University Graduate School, 03080 Seoul, Korea; cNeuroscience Research Institute, Seoul National University College of Medicine, 03080 Seoul, Korea; dScience Division, Tomocube, 34109 Daejeon, Korea; eConvergence Research Center for Diagnosis, Korea Institute of Science and Technology, 02792 Seoul, Korea; and fProteomics Core Facility, Biomedical Research Institute, Seoul National University Hospital, 03080 Seoul, Korea Edited by Richard D. Vierstra, Washington University in St. Louis, St. Louis, MO, and approved July 1, 2020 (received for review November 18, 2019) The 26S proteasome, a self-compartmentalized protease complex, additional LC3-interacting region; the target cargoes can be plays a crucial role in protein quality control. Multiple levels of docked onto phosphatidylethanolamine-modified LC3 (LC3-II) regulatory systems modulate proteasomal activity for substrate on the expanding phagophore membrane, enveloped by an hydrolysis. However, the destruction mechanism of mammalian autophagosome, and eventually degraded in the autolysosomes. proteasomes is poorly understood. We found that inhibited pro- Notably, the enzymatic cascade attaching the lipid moiety at the teasomes are sequestered into the insoluble aggresome via C-terminal glycine of the cleaved LC3 protein in autophagy re- HDAC6- and dynein-mediated transport. -

RTL8 Promotes Nuclear Localization of UBQLN2 to Subnuclear Compartments Associated with Protein Quality Control
bioRxiv preprint doi: https://doi.org/10.1101/2021.04.21.440788; this version posted April 22, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Title: RTL8 promotes nuclear localization of UBQLN2 to subnuclear compartments associated with protein quality control Harihar Milaganur Mohan1,2, Amit Pithadia1, Hanna Trzeciakiewicz1, Emily V. Crowley1, Regina Pacitto1 Nathaniel Safren1,3, Chengxin Zhang4, Xiaogen Zhou4, Yang Zhang4, Venkatesha Basrur5, Henry L. Paulson1,*, Lisa M. Sharkey1,* 1. Department of Neurology, University of Michigan, Ann Arbor, MI 48109-2200 2. Graduate Program in Cellular and Molecular Biology, University of Michigan, Ann Arbor, MI 48109- 2200 3. Present address: Department of Neurology, Northwestern University Feinberg School of Medicine, Chicago, IL 60611 4. Department of Computational Medicine and Bioinformatics, University of Michigan, Ann Arbor, MI 48109-2200 5. Department of Pathology, University of Michigan Medical School, Ann Arbor, MI 48109. *Corresponding authors: Henry Paulson: Department of Neurology, University of Michigan, Ann Arbor, MI 48109-2200; [email protected]; Tel. (734) 615-5632; Fax. (734) 615-5655 Lisa M Sharkey: Department of Neurology, University of Michigan, Ann Arbor, MI 48109-2200; [email protected]; Tel. (734) 763-3496; Fax (734) 615-5655 Keywords: Ubiquilin, UBQLN2, RTL8, Nuclear Protein Quality Control, Ubiquitin Proteasome System Declarations Funding: This work was supported by NIH 9R01NS096785-06, 1P30AG053760-01, The Amyotrophic Lateral Sclerosis Foundation and the UM Protein Folding Disease Initiative. Conflicts of interest/Competing interests: None declared Availability of data and material: • The authors confirm that the data supporting the findings in this are available within the article, at repository links provided within the article, and within its supplementary files. -

Deciphering the Molecular Profile of Plaques, Memory Decline And
ORIGINAL RESEARCH ARTICLE published: 16 April 2014 AGING NEUROSCIENCE doi: 10.3389/fnagi.2014.00075 Deciphering the molecular profile of plaques, memory decline and neuron loss in two mouse models for Alzheimer’s disease by deep sequencing Yvonne Bouter 1†,Tim Kacprowski 2,3†, Robert Weissmann4, Katharina Dietrich1, Henning Borgers 1, Andreas Brauß1, Christian Sperling 4, Oliver Wirths 1, Mario Albrecht 2,5, Lars R. Jensen4, Andreas W. Kuss 4* andThomas A. Bayer 1* 1 Division of Molecular Psychiatry, Georg-August-University Goettingen, University Medicine Goettingen, Goettingen, Germany 2 Department of Bioinformatics, Institute of Biometrics and Medical Informatics, University Medicine Greifswald, Greifswald, Germany 3 Department of Functional Genomics, Interfaculty Institute for Genetics and Functional Genomics, University Medicine Greifswald, Greifswald, Germany 4 Human Molecular Genetics, Department for Human Genetics of the Institute for Genetics and Functional Genomics, Institute for Human Genetics, University Medicine Greifswald, Ernst-Moritz-Arndt University Greifswald, Greifswald, Germany 5 Institute for Knowledge Discovery, Graz University of Technology, Graz, Austria Edited by: One of the central research questions on the etiology of Alzheimer’s disease (AD) is the Isidro Ferrer, University of Barcelona, elucidation of the molecular signatures triggered by the amyloid cascade of pathological Spain events. Next-generation sequencing allows the identification of genes involved in disease Reviewed by: Isidro Ferrer, University of Barcelona, processes in an unbiased manner. We have combined this technique with the analysis of Spain two AD mouse models: (1) The 5XFAD model develops early plaque formation, intraneu- Dietmar R. Thal, University of Ulm, ronal Ab aggregation, neuron loss, and behavioral deficits. (2)TheTg4–42 model expresses Germany N-truncated Ab4–42 and develops neuron loss and behavioral deficits albeit without plaque *Correspondence: formation. -

Mutant Uromodulin Expression Leads to Altered Homeostasis of the Endoplasmic Reticulum and Activates the Unfolded Protein Response
RESEARCH ARTICLE Mutant uromodulin expression leads to altered homeostasis of the endoplasmic reticulum and activates the unfolded protein response CeÂline Schaeffer1, Stefania Merella2, Elena Pasqualetto1, Dejan Lazarevic2, Luca Rampoldi1* a1111111111 1 Molecular Genetics of Renal Disorders, Division of Genetics and Cell Biology, IRCCS San Raffaele Scientific Institute, Milan, Italy, 2 Center of Translational Genomics and Bioinformatics, IRCCS San Raffaele a1111111111 Scientific Institute, Milan, Italy a1111111111 a1111111111 * [email protected] a1111111111 Abstract Uromodulin is the most abundant urinary protein in physiological conditions. It is exclusively OPEN ACCESS produced by renal epithelial cells lining the thick ascending limb of Henle's loop (TAL) and it Citation: Schaeffer C, Merella S, Pasqualetto E, plays key roles in kidney function and disease. Mutations in UMOD, the gene encoding uro- Lazarevic D, Rampoldi L (2017) Mutant uromodulin expression leads to altered modulin, cause autosomal dominant tubulointerstitial kidney disease uromodulin-related homeostasis of the endoplasmic reticulum and (ADTKD-UMOD), characterised by hyperuricemia, gout and progressive loss of renal func- activates the unfolded protein response. PLoS ONE tion. While the primary effect of UMOD mutations, retention in the endoplasmic reticulum 12(4): e0175970. https://doi.org/10.1371/journal. (ER), is well established, its downstream effects are still largely unknown. To gain insight pone.0175970 into ADTKD-UMOD pathogenesis, we performed transcriptional profiling and biochemical Editor: Salvatore V. Pizzo, Duke University School characterisation of cellular models (immortalised mouse TAL cells) of robust expression of of Medicine, UNITED STATES wild type or mutant GFP-tagged uromodulin. In this model mutant uromodulin accumulation Received: January 5, 2017 in the ER does not impact on cell viability and proliferation. -

Differential Effects of STCH and Stress-Inducible Hsp70 on the Stability and Maturation of NKCC2
International Journal of Molecular Sciences Article Differential Effects of STCH and Stress-Inducible Hsp70 on the Stability and Maturation of NKCC2 Dalal Bakhos-Douaihy 1,2, Elie Seaayfan 1,2,† , Sylvie Demaretz 1,2,†, Martin Komhoff 3,‡ and Kamel Laghmani 1,2,*,‡ 1 Centre de Recherche des Cordeliers, INSERM, Sorbonne Université, USPC, Université Paris Descartes, Université Paris Diderot, 75006 Paris, France; [email protected] (D.B.-D.); [email protected] (E.S.); [email protected] (S.D.) 2 CNRS, ERL8228, 75006 Paris, France 3 University Children’s Hospital, Philipps-University, 35043 Marburg, Germany; [email protected] * Correspondence: [email protected]; Tel.: +33-1-44-27-50-68; Fax: +33-1-44-27-51-19 † These authors contributed equally to this work. ‡ These authors contributed equally to this work. Abstract: Mutations in the Na-K-2Cl co-transporter NKCC2 lead to type I Bartter syndrome, a life- threatening kidney disease. We previously showed that export from the ER constitutes the limiting step in NKCC2 maturation and cell surface expression. Yet, the molecular mechanisms involved in this process remain obscure. Here, we report the identification of chaperone stress 70 protein (STCH) and the stress-inducible heat shock protein 70 (Hsp70), as two novel binding partners of the ER-resident form of NKCC2. STCH knock-down increased total NKCC2 expression whereas Hsp70 knock-down or its inhibition by YM-01 had the opposite effect. Accordingly, overexpressing of STCH and Hsp70 exerted opposite actions on total protein abundance of NKCC2 and its folding mutants. -
Download Validation Data
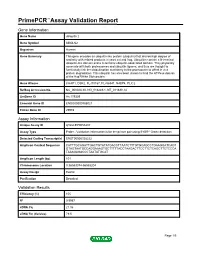
PrimePCR™Assay Validation Report Gene Information Gene Name ubiquilin 2 Gene Symbol UBQLN2 Organism Human Gene Summary This gene encodes an ubiquitin-like protein (ubiquilin) that shares high degree of similarity with related products in yeast rat and frog. Ubiquilins contain a N-terminal ubiquitin-like domain and a C-terminal ubiquitin-associated domain. They physically associate with both proteasomes and ubiquitin ligases; and thus are thought to functionally link the ubiquitination machinery to the proteasome to affect in vivo protein degradation. This ubiquilin has also been shown to bind the ATPase domain of the Hsp70-like Stch protein. Gene Aliases CHAP1, DSK2, FLJ10167, FLJ56541, N4BP4, PLIC2 RefSeq Accession No. NC_000023.10, NG_016249.1, NT_011630.14 UniGene ID Hs.179309 Ensembl Gene ID ENSG00000188021 Entrez Gene ID 29978 Assay Information Unique Assay ID qHsaCEP0055207 Assay Type Probe - Validation information is for the primer pair using SYBR® Green detection Detected Coding Transcript(s) ENST00000338222 Amplicon Context Sequence CATTTGCAGATTGACTGTATATGACCTTAATCTTTGTGCAGCCTGAAGGATCAGT GTAGTAATGCCAGGAAAGTGCTTTTTACCTAAGACTTCCTTCTCAGCTTCTCCCA TAAAGAGACCCTAATATGCAT Amplicon Length (bp) 101 Chromosome Location X:56593074-56593204 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 105 R2 0.9967 cDNA Cq 21.06 cDNA Tm (Celsius) 79.5 Page 1/5 PrimePCR™Assay Validation Report gDNA Cq 25.2 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay -

UBQLN2/Ubiquilin 2 Mutation and Pathology in Familial Amyotrophic Lateral Sclerosis Kelly L
Neurobiology of Aging 33 (2012) 2527.e3–2527.e10 www.elsevier.com/locate/neuaging UBQLN2/ubiquilin 2 mutation and pathology in familial amyotrophic lateral sclerosis Kelly L. Williamsa,b, Sadaf T. Warraicha,b, Shu Yanga, Jennifer A. Solskia, Ruvini Fernandoa, Guy A. Rouleauc, Garth A. Nicholsona,b,d, Ian P. Blaira,b,* a Northcott Neuroscience Laboratory, ANZAC Research Institute, Sydney, New South Wales, Australia b Sydney Medical School, University of Sydney, New South Wales, Australia c Department of Medicine, University of Montreal, Montreal, Canada d Molecular Medicine Laboratory, Concord Hospital, New South Wales, Australia Received 9 March 2012; received in revised form 18 May 2012; accepted 20 May 2012 Abstract Amyotrophic lateral sclerosis (ALS) shows clinical and pathological overlap with frontotemporal dementia that includes the presence of hallmark ubiquitinated inclusions in affected neurons. Mutations in UBQLN2, which encodes ubiquilin 2, were recently identified in X-linked juvenile and adult-onset ALS and ALS/dementia. As part of an established exome sequencing program to identify disease genes in familial ALS, we identified a novel missense UBQLN2 mutation (c.1460CϾT, p.T487I) in 2 apparently unrelated multigenerational ALS families with no evidence of frontotemporal dementia. This mutation segregated with the disease and was absent in 820 healthy controls and all public single nucleotide polymorphism databases. The UBQLN2 p.T487I mutation substitutes a highly conserved residue and is located immediately upstream of a PXX region where all previous mutations have been identified. Immunostaining of spinal cord from a patient with UBQLN2 p.T487I mutation showed colocalization of ubiquilin 2 with ubiquitin in all neuronal inclusions examined and frequent colocalization with TAR DNA-binding protein 43 (TDP-43) and fused in sarcoma protein (FUS).