BCL9 Promotes Tumor Progression by Conferring Enhanced Proliferative, Metastatic, and Angiogenic Properties to Cancer Cells
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Constitutive Scaffolding of Multiple Wnt Enhanceosome Components By
RESEARCH ARTICLE Constitutive scaffolding of multiple Wnt enhanceosome components by Legless/ BCL9 Laurens M van Tienen, Juliusz Mieszczanek, Marc Fiedler, Trevor J Rutherford, Mariann Bienz* MRC Laboratory of Molecular Biology, Cambridge, United Kingdom Abstract Wnt/b-catenin signaling elicits context-dependent transcription switches that determine normal development and oncogenesis. These are mediated by the Wnt enhanceosome, a multiprotein complex binding to the Pygo chromatin reader and acting through TCF/LEF- responsive enhancers. Pygo renders this complex Wnt-responsive, by capturing b-catenin via the Legless/BCL9 adaptor. We used CRISPR/Cas9 genome engineering of Drosophila legless (lgs) and human BCL9 and B9L to show that the C-terminus downstream of their adaptor elements is crucial for Wnt responses. BioID proximity labeling revealed that BCL9 and B9L, like PYGO2, are constitutive components of the Wnt enhanceosome. Wnt-dependent docking of b-catenin to the enhanceosome apparently causes a rearrangement that apposes the BCL9/B9L C-terminus to TCF. This C-terminus binds to the Groucho/TLE co-repressor, and also to the Chip/LDB1-SSDP enhanceosome core complex via an evolutionary conserved element. An unexpected link between BCL9/B9L, PYGO2 and nuclear co-receptor complexes suggests that these b-catenin co-factors may coordinate Wnt and nuclear hormone responses. DOI: 10.7554/eLife.20882.001 *For correspondence: mb2@mrc- Introduction lmb.cam.ac.uk The Wnt/b-catenin signaling cascade is an ancient cell communication pathway that operates con- Competing interests: The text-dependent transcriptional switches to control animal development and tissue homeostasis authors declare that no (Cadigan and Nusse, 1997). -

Nucleo-Cytoplasmic Distribution of ß-Catenin Is Regulated by Retention
Research Article 1453 Nucleo-cytoplasmic distribution of -catenin is regulated by retention Eva Krieghoff, Jürgen Behrens* and Bernhard Mayr Nikolaus-Fiebiger-Center for Molecular Medicine, University of Erlangen-Nürnberg, Glückstr. 6, 91054 Erlangen, Germany *Author for correspondence (e-mail: [email protected]) Accepted 19 December 2005 Journal of Cell Science 119, 1453-1463 Published by The Company of Biologists 2006 doi:10.1242/jcs.02864 Summary -catenin is the central signalling molecule of the canonical of -catenin, i.e. increases the rate of -catenin nuclear Wnt pathway, where it activates target genes in a complex import or export. Moreover, the cytoplasmic enrichment of with LEF/TCF transcription factors in the nucleus. The -catenin by APC and axin is not abolished by inhibition regulation of -catenin activity is thought to occur mainly of CRM-1-dependent nuclear export. TCF4, APC, axin and on the level of protein degradation, but it has been axin2 move more slowly than -catenin in their respective suggested that -catenin nuclear localization and hence its compartment, and concomitantly decrease -catenin transcriptional activity may additionally be regulated via mobility. Together, these data indicate that -catenin nuclear import by TCF4 and BCL9 and via nuclear export interaction partners mainly regulate -catenin subcellular by APC and axin. Using live-cell microscopy and localization by retaining it in the compartment in which fluorescence recovery after photobleaching (FRAP), we they are localized, rather than by active transport into or have directly analysed the impact of these factors on the out of the nucleus. subcellular localization of -catenin, its nucleo-cytoplasmic shuttling and its mobility within the nucleus and the Supplementary material available online at cytoplasm. -

BCL9 Provides Multi-Cellular Communication Properties in Colorectal Cancer by Interacting with Paraspeckle Proteins
ARTICLE https://doi.org/10.1038/s41467-019-13842-7 OPEN BCL9 provides multi-cellular communication properties in colorectal cancer by interacting with paraspeckle proteins Meng Jiang 1,2,8, Yue Kang1,3,8, Tomasz Sewastianik1,4, Jiao Wang1,5, Helen Tanton1, Keith Alder1, Peter Dennis1, Yu Xin1, Zhongqiu Wang1,6, Ruiyang Liu1, Mengyun Zhang1, Ying Huang1, Massimo Loda1, Amitabh Srivastava7, Runsheng Chen3, Ming Liu2 & Ruben D. Carrasco1,7* 1234567890():,; Colorectal cancer (CRC) is the third most commonly diagnosed cancer, which despite recent advances in treatment, remains incurable due to molecular heterogeneity of tumor cells. The B-cell lymphoma 9 (BCL9) oncogene functions as a transcriptional co-activator of the Wnt/ β-catenin pathway, which plays critical roles in CRC pathogenesis. Here we have identified a β-catenin-independent function of BCL9 in a poor-prognosis subtype of CRC tumors char- acterized by expression of stromal and neural associated genes. In response to spontaneous calcium transients or cellular stress, BCL9 is recruited adjacent to the interchromosomal regions, where it stabilizes the mRNA of calcium signaling and neural associated genes by interacting with paraspeckle proteins. BCL9 subsequently promotes tumor progression and remodeling of the tumor microenvironment (TME) by sustaining the calcium transients and neurotransmitter-dependent communication among CRC cells. These data provide additional insights into the role of BCL9 in tumor pathogenesis and point towards additional avenues for therapeutic intervention. 1 Department of Oncologic Pathology, Dana-Farber Cancer Institute, Harvard Medical School, Boston, MA 02115, USA. 2 Department of General Surgery, Fourth Affiliated Hospital of Harbin Medical University, Harbin Medical University, Harbin 150001, China. -

TBX3 Acts As Tissue-Specific Component of the Wnt/Β
bioRxiv preprint doi: https://doi.org/10.1101/2020.04.22.053561; this version posted April 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. TBX3 acts as tissue-specific component of the Wnt/b-catenin enhanceosome Dario Zimmerli1,6#, Costanza Borrelli2#, Amaia Jauregi-Miguel3,4#, Simon Söderholm3,4, Salome Brütsch1, Nikolaos Doumpas1, Jan Reichmuth1, Fabienne Murphy-Seiler5, Michel Aguet5, Konrad Basler1*, Andreas E. Moor2*, Claudio Cantù3,4* 1 Department of Molecular Life Sciences, University of Zurich, Zürich, Switzerland, CH-8057 2 Institute of Molecular Cancer Research, University of Zurich, Zürich, Switzerland, CH-8057 3 Wallenberg Centre for Molecular Medicine, Linköping University 4 Department of Biomedical and Clinical Sciences, Faculty of Health Science, SE-581 83 Linköping, Sweden 5Swiss Institute for Experimental Cancer Research (ISREC), Ecole Polytechnique Fédérale de Lausanne (EPFL), School of Life Sciences, CH-1015 Lausanne, Switzerland 6 Current address: Division of Molecular Pathology, The Netherlands Cancer Institute, Amsterdam, The Netherlands # These authors contributed equally to this work * For correspondence: [email protected] [email protected] [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.04.22.053561; this version posted April 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. -

Whole-Exome Sequencing of Metastatic Cancer and Biomarkers of Treatment Response
Supplementary Online Content Beltran H, Eng K, Mosquera JM, et al. Whole-exome sequencing of metastatic cancer and biomarkers of treatment response. JAMA Oncol. Published online May 28, 2015. doi:10.1001/jamaoncol.2015.1313 eMethods eFigure 1. A schematic of the IPM Computational Pipeline eFigure 2. Tumor purity analysis eFigure 3. Tumor purity estimates from Pathology team versus computationally (CLONET) estimated tumor purities values for frozen tumor specimens (Spearman correlation 0.2765327, p- value = 0.03561) eFigure 4. Sequencing metrics Fresh/frozen vs. FFPE tissue eFigure 5. Somatic copy number alteration profiles by tumor type at cytogenetic map location resolution; for each cytogenetic map location the mean genes aberration frequency is reported eFigure 6. The 20 most frequently aberrant genes with respect to copy number gains/losses detected per tumor type eFigure 7. Top 50 genes with focal and large scale copy number gains (A) and losses (B) across the cohort eFigure 8. Summary of total number of copy number alterations across PM tumors eFigure 9. An example of tumor evolution looking at serial biopsies from PM222, a patient with metastatic bladder carcinoma eFigure 10. PM12 somatic mutations by coverage and allele frequency (A) and (B) mutation correlation between primary (y- axis) and brain metastasis (x-axis) eFigure 11. Point mutations across 5 metastatic sites of a 55 year old patient with metastatic prostate cancer at time of rapid autopsy eFigure 12. CT scans from patient PM137, a patient with recurrent platinum refractory metastatic urothelial carcinoma eFigure 13. Tracking tumor genomics between primary and metastatic samples from patient PM12 eFigure 14. -

Low BCL9 Expression Inhibited Ovarian Epithelial Malignant Tumor
Wang et al. Cancer Cell Int (2019) 19:330 https://doi.org/10.1186/s12935-019-1009-5 Cancer Cell International PRIMARY RESEARCH Open Access Low BCL9 expression inhibited ovarian epithelial malignant tumor progression by decreasing proliferation, migration, and increasing apoptosis to cancer cells Jing Wang1,2, Mingjun Zheng1,2, Liancheng Zhu1,2, Lu Deng1,2,3, Xiao Li1,2, Linging Gao1,2, Caixia Wang1,2, Huimin Wang1,2,4, Juanjuan Liu1,2 and Bei Lin1,2* Abstract Background: Abnormal activation of the classic Wnt signaling pathway is closely related to the occurrence of epithelial cancers. B-cell lymphoma 9 (BCL9), a transcription factor, is a novel oncogene discovered in the classic Wnt pathway and promotes the occurrence and development of various tumors. Ovarian cancer is the gynecological malignant tumor with the highest mortality because it is difcult to diagnose early, and easy to relapse and metas- tasis. The expression and role of BCL9 in epithelial ovarian cancer (EOC) have not been studied. Thus, in this research, we aimed to investigate the expression and clinical signifcance of BCL9 in EOC tissues and its efect on the malignant biological behavior of human ovarian cancer cells. Methods: We detect the expression of BCL9 in ovarian epithelial tumor tissues and normal ovarian tissues using immunohistochemistry and analyzed the relationship between it and clinicopathological parameters and patient prognosis. The expression of proteins was detected by Western blot. The MTT assay, fow cytometry, the scratch assay, and the transwell assay were used to detect cell proliferation, apoptosis, migration, and invasion, respectively. A total of 374 ovarian cancer tissue samples were collected using TCGA database. -

Association Between the UBE2Z Rs46522 and TCF7L2 Rs7903146 Polymorphisms with Type 2 Diabetes in South Western Iran
Association between the UBE2Z rs46522 and TCF7L2 rs7903146 polymorphisms with type 2 diabetes in south western Iran Ali Mohammad Foroughmand1, Sana Shafidelpour1, Merhrnoosh Zakerkish2, Mehdi Pourmehdi Borujeni3 1. Department of Genetics, Faculty of Sciences, Shahid Chamran University of Ahvaz, Ahvaz, Iran. 2. Department of Endocrinology and Metabolism, Health Research Institute, Diabetes Research Center, Ahvaz Jundishapur University of Medical Sciences, Ahvaz, Iran. 3. Department of Food Hygiene and Quality control, Faculty of Veterinary Medicine, Shahid Chamran University of Ahvaz, Ahvaz, Iran. Abstract Background: Transcription factor 7-like 2 Protein (TCF7L2) has a strong role in the pathogenesis of type 2 diabetes melli- tus (T2DM). Association between rs7903146 and T2D risk reported in some of populations. Also many loci such as UBE2Z rs46522 are affecting by TCF7L2 transcription factor have been found associated with T2D. The present study aimed to evaluate association of the SNPs with risk of T2D among our population. Methods: This case-control study was conducted on 150 T2D patients and 150 healthy people (as a control group) in south western Iran. Genotyping was performed by PCR-RFLP. Results: The frequency of genotypes showed no remarkable difference between T2DM patients and control group. The odds ratios of rs7903146 (C/T) poly¬morphism for CC and TC genotypes were 1.9 (95% CI, 0.85 to 4.24; P=0.12) and 0.81 (95% CI, 0.47 to 1.38; P=0.43) compared with the TT genotype, respectively. The odds ratios of rs46522 (C/T) poly¬morphism for TT and TC genotypes were 1.75 (95% CI, 0.86 to 3.59; P=0.13) and 1.38 (95% CI, 0.81to 2.35; P=0.24) compared with the CC genotype, respectively. -

Proximal Microdeletions and Microduplications of 1Q21.1 Contribute to Variable Abnormal Phenotypes
European Journal of Human Genetics (2012) 20, 754–761 & 2012 Macmillan Publishers Limited All rights reserved 1018-4813/12 www.nature.com/ejhg ARTICLE Proximal microdeletions and microduplications of 1q21.1 contribute to variable abnormal phenotypes Jill A Rosenfeld1, Ryan N Traylor1, G Bradley Schaefer2, Elizabeth W McPherson3, Blake C Ballif1, Eva Klopocki4, Stefan Mundlos4, Lisa G Shaffer*,1 and Arthur S Aylsworth5, 1q21.1 Study Group6 Chromosomal band 1q21.1 can be divided into two distinct regions, proximal and distal, based on segmental duplications that mediate recurrent rearrangements. Microdeletions and microduplications of the distal region within 1q21.1, which are susceptibility factors for a variety of neurodevelopmental phenotypes, have been more extensively studied than proximal microdeletions and microduplications. Proximal microdeletions are known as a susceptibility factor for thrombocytopenia-absent radius (TAR) syndrome, but it is unclear if these proximal microdeletions have other phenotypic consequences. Therefore, to elucidate the clinical significance of rearrangements of the proximal 1q21.1 region, we evaluated the phenotypes in patients identified with 1q21.1 rearrangements after referral for clinical microarray testing. We report clinical information for 55 probands with copy number variations (CNVs) involving proximal 1q21.1: 22 microdeletions and 20 reciprocal microduplications limited to proximal 1q21.1 and 13 microdeletions that include both the proximal and distal regions. Six individuals with proximal microdeletions have TAR syndrome. Three individuals with proximal microdeletions and two individuals with larger microdeletions of proximal and distal 1q21.1 have a ‘partial’ TAR phenotype. Furthermore, one subject with TAR syndrome has a smaller, atypical deletion, narrowing the critical deletion region for the syndrome. -

Xo PANEL DNA GENE LIST
xO PANEL DNA GENE LIST ~1700 gene comprehensive cancer panel enriched for clinically actionable genes with additional biologically relevant genes (at 400 -500x average coverage on tumor) Genes A-C Genes D-F Genes G-I Genes J-L AATK ATAD2B BTG1 CDH7 CREM DACH1 EPHA1 FES G6PC3 HGF IL18RAP JADE1 LMO1 ABCA1 ATF1 BTG2 CDK1 CRHR1 DACH2 EPHA2 FEV G6PD HIF1A IL1R1 JAK1 LMO2 ABCB1 ATM BTG3 CDK10 CRK DAXX EPHA3 FGF1 GAB1 HIF1AN IL1R2 JAK2 LMO7 ABCB11 ATR BTK CDK11A CRKL DBH EPHA4 FGF10 GAB2 HIST1H1E IL1RAP JAK3 LMTK2 ABCB4 ATRX BTRC CDK11B CRLF2 DCC EPHA5 FGF11 GABPA HIST1H3B IL20RA JARID2 LMTK3 ABCC1 AURKA BUB1 CDK12 CRTC1 DCUN1D1 EPHA6 FGF12 GALNT12 HIST1H4E IL20RB JAZF1 LPHN2 ABCC2 AURKB BUB1B CDK13 CRTC2 DCUN1D2 EPHA7 FGF13 GATA1 HLA-A IL21R JMJD1C LPHN3 ABCG1 AURKC BUB3 CDK14 CRTC3 DDB2 EPHA8 FGF14 GATA2 HLA-B IL22RA1 JMJD4 LPP ABCG2 AXIN1 C11orf30 CDK15 CSF1 DDIT3 EPHB1 FGF16 GATA3 HLF IL22RA2 JMJD6 LRP1B ABI1 AXIN2 CACNA1C CDK16 CSF1R DDR1 EPHB2 FGF17 GATA5 HLTF IL23R JMJD7 LRP5 ABL1 AXL CACNA1S CDK17 CSF2RA DDR2 EPHB3 FGF18 GATA6 HMGA1 IL2RA JMJD8 LRP6 ABL2 B2M CACNB2 CDK18 CSF2RB DDX3X EPHB4 FGF19 GDNF HMGA2 IL2RB JUN LRRK2 ACE BABAM1 CADM2 CDK19 CSF3R DDX5 EPHB6 FGF2 GFI1 HMGCR IL2RG JUNB LSM1 ACSL6 BACH1 CALR CDK2 CSK DDX6 EPOR FGF20 GFI1B HNF1A IL3 JUND LTK ACTA2 BACH2 CAMTA1 CDK20 CSNK1D DEK ERBB2 FGF21 GFRA4 HNF1B IL3RA JUP LYL1 ACTC1 BAG4 CAPRIN2 CDK3 CSNK1E DHFR ERBB3 FGF22 GGCX HNRNPA3 IL4R KAT2A LYN ACVR1 BAI3 CARD10 CDK4 CTCF DHH ERBB4 FGF23 GHR HOXA10 IL5RA KAT2B LZTR1 ACVR1B BAP1 CARD11 CDK5 CTCFL DIAPH1 ERCC1 FGF3 GID4 HOXA11 -

BCL9 and C9orf5 Are Associated with Negative Symptoms in Schizophrenia: Meta-Analysis of Two Genome-Wide Association Studies
University of Texas Rio Grande Valley ScholarWorks @ UTRGV Health and Biomedical Sciences Faculty Publications and Presentations College of Health Professions 2013 BCL9 and C9orf5 are associated with negative symptoms in schizophrenia: meta-analysis of two genome-wide association studies Chun Xu The University of Texas Rio Grande Valley Nagesh Aragam Xia Li Erika Cynthia Villa Liang Wang See next page for additional authors Follow this and additional works at: https://scholarworks.utrgv.edu/hbs_fac Part of the Psychiatry and Psychology Commons Recommended Citation Xu, C., Aragam, N., Li, X., Villla, E. C., Wang, L., Briones, D., Petty, L., Posada, Y., Arana, T. B., Cruz, G., Mao, C., Camarillo, C., Su, B. B., Escamilla, M. A., & Wang, K. (2013). BCL9 and C9orf5 are associated with negative symptoms in schizophrenia: meta-analysis of two genome-wide association studies. PloS one, 8(1), e51674. https://doi.org/10.1371/journal.pone.0051674 This Article is brought to you for free and open access by the College of Health Professions at ScholarWorks @ UTRGV. It has been accepted for inclusion in Health and Biomedical Sciences Faculty Publications and Presentations by an authorized administrator of ScholarWorks @ UTRGV. For more information, please contact [email protected], [email protected]. Authors Chun Xu, Nagesh Aragam, Xia Li, Erika Cynthia Villa, Liang Wang, David Briones, Brenda Bin Su, Michael A. Escamilla, and KeSheng Wang This article is available at ScholarWorks @ UTRGV: https://scholarworks.utrgv.edu/hbs_fac/85 BCL9 and C9orf5 Are Associated with Negative Symptoms in Schizophrenia: Meta-Analysis of Two Genome-Wide Association Studies Chun Xu1*, Nagesh Aragam2,3, Xia Li4, Erika Cynthia Villla5, Liang Wang2, David Briones1, Leonora Petty1, Yolanda Posada6, Tania Bedard Arana1, Grace Cruz1, ChunXiang Mao7, Cynthia Camarillo1, Brenda Bin Su7, Michael A. -
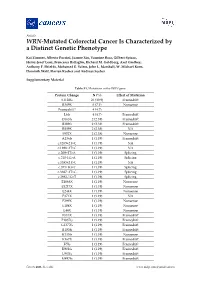
WRN-Mutated Colorectal Cancer Is Characterized by a Distinct Genetic Phenotype
Article WRN-Mutated Colorectal Cancer Is Characterized by a Distinct Genetic Phenotype Kai Zimmer, Alberto Puccini, Joanne Xiu, Yasmine Baca, Gilbert Spizzo, Heinz-Josef Lenz, Francesca Battaglin, Richard M. Goldberg, Axel Grothey, Anthony F. Shields, Mohamed E. Salem, John L. Marshall, W. Michael Korn, Dominik Wolf, Florian Kocher and Andreas Seeber Supplementary Material Table S1. Mutations in the WRN gene. Protein Change N (%) Effect of Mutation S1128fs 26 (30.9) Frameshift R369X 6 (7.1) Nonsense Frameshift* 4 (4.7) L6fs 4 (4.7) Frameshift D163fs 2 (2.38) Frameshift R389fs 2 (2.38) Frameshift R889X 2 (2.38) NA S952X 2 (2.38) Nonsense A136fs 1 (1.19) Frameshift c.1269+2T>C 1 (1.19) NA c.1898+2T>G 1 (1.19) NA c.209+2T>A 1 (1.19) Splicing c.210-1G>A 1 (1.19) Splicing c.3383+2T>C 1 (1.19) NA c.355+1G>T 1 (1.19) Splicing c.3687+2T>G 1 (1.19) Splicing c.3983-1G>T 1 (1.19) Splicing E1068X 1 (1.19) Nonsense E1217X 1 (1.19) Nonsense E244X 1 (1.19) Nonsense E371X 1 (1.19) NA E399X 1 (1.19) Nonsense E488X 1 (1.19) Nonsense E48X 1 (1.19) Nonsense E513X 1 (1.19) Frameshift F1037fs 1 (1.19) Frameshift G1377fs 1 (1.19) Frameshift I1183fs 1 (1.19) Frameshift K135fs 1 (1.19) Nonsense K167X 1 (1.19) Frameshift K5fs 1 (1.19) Frameshift K901fs 1 (1.19) Frameshift L967fs 1 (1.19) Frameshift M497fs 1 (1.19) Frameshift Cancers 2020, 12, x; doi: www.mdpi.com/journal/cancers Cancers 2020, 12 S2 of 13 N166fs 1 (1.19) Frameshift Q11fs 1 (1.19) Frameshift R1305X 1 (1.19) Nonsense R279fs 1 (1.19) Frameshift R565X 1 (1.19) Nonsene R741X 1 (1.19) Nonsense R987X 1 (1.19) NA T1011fs 1 (1.19) Frameshift W1014X 1 (1.19) Nonsense Y57X 1 (1.19) Nonsense Grand Total 84 *4 Frameshift mutations could not be described more precisely. -

N-Terminal Region (P100600 T100) Data Sheet
CTNNB1 antibody - N-terminal region (P100600_T100) Data Sheet Product Number P100600_T100 Product Name CTNNB1 antibody - N-terminal region (P100600_T100) Size 100ug Gene Symbol CTNNB1 Alias Symbols CTNNB Nucleotide Accession# NM_001904 Protein Size (# AA) 781 amino acids Molecular Weight 85kDa Product Format Lyophilized powder NCBI Gene Id 1499 Host Rabbit Clonality Polyclonal Official Gene Full Name Catenin (cadherin-associated protein), beta 1, 88kDa Gene Family ARMC This is a rabbit polyclonal antibody against CTNNB1. It was validated on Western Blot using a cell lysate as a Description positive control. Aviva Systems Biology strives to provide antibodies covering each member of a whole protein family of your interest. We also use our best efforts to provide you antibodies recognize various epitopes of a target protein. For availability of antibody needed for your experiment, please inquire (). Peptide Sequence Synthetic peptide located within the following region: SLSGKGNPEEEDVDTSQVLYEWEQGFSQSFTQEQVADIDGQYAMTRAQRV Target Reference Strausberg, R.L., et al., (2002) Proc. Natl. Acad. Sci. U.S.A. 99 (26), 16899-16903. Description of Target CTNNB1 is involved in the regulation of cell adhesion and in signal transduction through the Wnt pathway. APC, APC, AXIN2, BCL3, BCL9, BTRC, CCND1, CD82, CDH1, CDH1, CDH1, CTNNA1, DKK1, ESR1, FAS, Partner Proteins FGF20, FOXO1, FOXO3, FOXO4, GSK3B, IGFBP2, KLK3, LEF1, MUC1, NFKB1, NR5A2, PKD1, PSEN1, RUVBL2, SGK1, TCF7L2, TCF7L2, TFF1, ACP1, AJAP1, APC, APC2, AR, ARHGAP32, AXIN1, AXIN2, BCL3, BCL9, BCL9L, BOC, BTRC, CA9, Reconstitution and Add 100 ul of distilled water. Final anti-CTNNB1 antibody concentration is 1 mg/ml in PBS buffer with 2% Storage sucrose. For longer periods of storage, store at -20C.