Development of a Direct Activity Probe for Rho-Associated Protein Kinase" (2016)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Regulation of Calcium/Calmodulin-Dependent Protein Kinase II Activation by Intramolecular and Intermolecular Interactions
8394 • The Journal of Neuroscience, September 29, 2004 • 24(39):8394–8398 Mini-Review Regulation of Calcium/Calmodulin-Dependent Protein Kinase II Activation by Intramolecular and Intermolecular Interactions Leslie C. Griffith Department of Biology and Volen Center for Complex Systems, Brandeis University, Waltham, Massachusetts 02454-9110 Key words: calcium; calmodulin; learning; localization; NMDA; phosphatase; protein kinase As its name implies, calcium/calmodulin-dependent protein ki- The overlap of these subdomains is no accident. Binding of nase II (CaMKII) is calcium dependent. In its basal state, the Ca 2ϩ/CaM is the primary signal for release of autoinhibition. activity of CaMKII is extremely low. Regulation of intracellular Current models of activation posit that the binding of Ca 2ϩ/CaM calcium levels allows the neuron to link activity with phosphor- serves to disrupt the interactions of specific residues within the ylation by CaMKII. This review will briefly summarize our cur- autoinhibitory domain with the catalytic domain (Smith et al., rent understanding of the intramolecular mechanisms of activity 1992). Because there is no crystal structure for the catalytic and regulation and their modulation by Ca 2ϩ/CaM and will then regulatory parts of CaMKII, the interaction face of these two focus on the growing number of other modes of intermolecular domains has been inferred using the effects of charge-reversal regulation of CaMKII activity by substrate and scaffolding mutagenesis on activity and molecular modeling (Yang and molecules. Schulman, 1999). This study confirmed the role of Arg 297 at the P-3 position of the pseudosubstrate ligand (Mukherji and Soder- Regulation of CaMKII by its autoinhibitory domain ling, 1995) and identified residues in the catalytic domain that All members of the CaMKII family (␣, , ␥, and ␦ isozymes) share may have direct interactions with the regulatory region. -

Core Lab Brochure
CHOOSE THE MOST TRUSTED LONG-READ TECHNOLOGY FOR YOUR CORE Sequence with Confidence The Sequel® II and IIe Systems are powered by Single Molecule, Real-Time (SMRT®) Sequencing, a technology proven to produce highly accurate long reads, known as HiFi reads, for sequencing data you and your customers can trust. SMRT SEQUENCING IS SMART BUSINESS HiFi Reads: PacBio is the only sequencing technology to offer highly accurate long reads. Because HiFi reads are extremely accurate, downstream analysis is simplified and streamlined, requiring less compute time than the error-prone long reads of other technologies. High Throughput: The Sequel II and IIe Systems have high data yields on robust, highly automated platforms to increase productivity and reduce project costs. Efficient and Easy-To-Use Workflows: Our end-to-end solutions feature library preparation in <3 hours and many push- button analysis workflows, so you can run projects quickly and easily. Support: All of our products are backed by a global team of scientists, bioinformaticians, and engineers who stand ready to provide you with outstanding service. OUTSTANDING PERFORMANCE AND RELIABILITY 99% of runs on the Sequel II System completed successfully Sequel II Systems provide reliable performance with the total bases produced by the PacBio fleet steadily increasing, and 99% of runs completed successfully. “In our experience, the Sequel II System was essentially production-ready right out of the box. We have used it for a range of applications and sample types — from human genome sequencing to metagenome and microbiome profiling to non-model plant and animal genomes — and results have been very good.” — Luke Tallon, Director of the Genomics Resource Center at Maryland Genomics pacb.com/Sequel SMRT SEQUENCING APPLICATIONS – EFFICIENT AND COST EFFECTIVE The Sequel II and IIe Systems support a wide range of applications, each adding unique value to a sequencing study. -

The Drug Sensitivity and Resistance Testing (DSRT) Approach
A phenotypic screening and machine learning platform eciently identifies triple negative breast cancer-selective and readily druggable targets Prson Gautam 1 Alok Jaiswal 1 Tero Aittokallio 1, 2 Hassan Al Ali 3 Krister Wennerberg 1,4 Identifying eective oncogenic targets is challenged by the complexity of genetic alterations in 1Institute for Molecular Medicine Finland (FIMM), HiLIFE, University of Helsinki, Finland cancer and their poorly understood relation to cell function and survival. There is a need for meth- Current kinome coverage of kinase inhibitors in TNBC exhibit diverse kinase dependencies MFM-223 is selectively addicted to FGFR2 2Department of Mathematics and Statistics, University of Turku, Finland 3The Miami Project to Cure Paralysis, Peggy and Harold Katz Family Drug Discovery Center, A A Sylvester Comprehensive Cancer Center, and Department of Neurological Surgery and Medicine ods that rapidly and accurately identify “pharmacologically eective” targets without the require- clinical evaluation TN Kinases MFM-223 CAL-120 MDA-MB-231 TNBC TNBC TNBC TNBC TNBC TNBC HER2+ 100 University of Miami Miller School of Medicine, Miami, FL 33136, USA. non- HER2+ FGFR1 0.97 0.00 0.00 MFM-223 BL1 BL2 M MSL IM LAR ER+, PR+ 50 ment for priori knowledge of complex signaling networks. We developed an approach that uses ma- cancerous FGFR2 56.46 0.00 0.00 CAL-120 25 4 MDA-MB-231 Biotech Research & Innovation Centre (BRIC) and Novo Nordisk Foundation Center HCC1937 CAL-85-1 CAL-120 MDA-MB-231 DU4475 CAL-148 MCF-10A SK-BR-3 BT-474 FGFR3 25.10 0.00 0.00 0 chine learning to relate results from unbiased phenotypic screening of kinase inhibitors to their bio- for Stem Cell Biology (DanStem), University of Copenhagen, Denmark HCC1599 HDQ-P1 BT-549 MDA-MB-436 MFM-223 FGFR4 0.00 0.00 0.00 MAXIS*Bk Clinical status MDA-MB-468 CAL-51 Hs578T MDA-MB-453 score chemical activity data. -

Masre, Siti Fathiah (2015) Analysis of ROCK2 Activation in Transgenic Mouse Skin Carcinogenesis. Phd Thesis
Masre, Siti Fathiah (2015) Analysis of ROCK2 activation in transgenic mouse skin carcinogenesis. PhD thesis. http://theses.gla.ac.uk/6628/ Copyright and moral rights for this thesis are retained by the author A copy can be downloaded for personal non-commercial research or study This thesis cannot be reproduced or quoted extensively from without first obtaining permission in writing from the Author The content must not be changed in any way or sold commercially in any format or medium without the formal permission of the Author When referring to this work, full bibliographic details including the author, title, awarding institution and date of the thesis must be given Glasgow Theses Service http://theses.gla.ac.uk/ [email protected] Analysis of ROCK2 Activation in Transgenic Mouse Skin Carcinogenesis Siti Fathiah Masre A Thesis submitted to the University of Glasgow In fulfilment of the requirements for the Degree of Doctor of Philosophy School of Medicine College of Medical, Veterinary and Life Sciences University of Glasgow August 2015 2 Summary The purpose of this study was to investigate ROCK2 activation in squamous cell carcinogenesis and assess its co-operation with rasHa and fos oncogene activation together with loss of PTEN mediated AKT regulation. The analysis of ROCK deregulation with these genes in the MAP Kinase and PI3K pathways, two of the most commonly deregulated signalling systems, employed a well-characterised, transgenic mouse skin model of multi-stage carcinogenesis. A major goal was to study co-operation between these genes in the conversion of benign tumours to malignancy and investigate subsequent progression to aggressive carcinomas, given these are the most significant clinical stages of carcinogenesis from a patient’s viewpoint; and also investigated effects of ROCK2 deregulation on the processes of normal epidermal differentiation. -

Identification and Characterization of RHOA-Interacting Proteins in Bovine Spermatozoa1
BIOLOGY OF REPRODUCTION 78, 184–192 (2008) Published online before print 10 October 2007. DOI 10.1095/biolreprod.107.062943 Identification and Characterization of RHOA-Interacting Proteins in Bovine Spermatozoa1 Sarah E. Fiedler, Malini Bajpai, and Daniel W. Carr2 Department of Medicine, Oregon Health & Sciences University and Veterans Affairs Medical Center, Portland, Oregon 97239 ABSTRACT Guanine nucleotide exchange factors (GEFs) catalyze the GDP for GTP exchange [2]. Activation is negatively regulated by In somatic cells, RHOA mediates actin dynamics through a both guanine nucleotide dissociation inhibitors (RHO GDIs) GNA13-mediated signaling cascade involving RHO kinase and GTPase-activating proteins (GAPs) [1, 2]. Endogenous (ROCK), LIM kinase (LIMK), and cofilin. RHOA can be RHO can be inactivated via C3 exoenzyme ADP-ribosylation, negatively regulated by protein kinase A (PRKA), and it and studies have demonstrated RHO involvement in actin-based interacts with members of the A-kinase anchoring (AKAP) cytoskeletal response to extracellular signals, including lyso- family via intermediary proteins. In spermatozoa, actin poly- merization precedes the acrosome reaction, which is necessary phosphatidic acid (LPA) [2–4]. LPA is known to signal through for normal fertility. The present study was undertaken to G-protein-coupled receptors (GPCRs) [4, 5]; specifically, LPA- determine whether the GNA13-mediated RHOA signaling activated GNA13 (formerly Ga13) promotes RHO activation pathway may be involved in acrosome reaction in bovine through GEFs [4, 6]. Activated RHO-GTP then signals RHO caudal sperm, and whether AKAPs may be involved in its kinase (ROCK), resulting in the phosphorylation and activation targeting and regulation. GNA13, RHOA, ROCK2, LIMK2, and of LIM-kinase (LIMK), which in turn phosphorylates and cofilin were all detected by Western blot in bovine caudal inactivates cofilin, an actin depolymerizer, the end result being sperm. -

Application of a MYC Degradation
SCIENCE SIGNALING | RESEARCH ARTICLE CANCER Copyright © 2019 The Authors, some rights reserved; Application of a MYC degradation screen identifies exclusive licensee American Association sensitivity to CDK9 inhibitors in KRAS-mutant for the Advancement of Science. No claim pancreatic cancer to original U.S. Devon R. Blake1, Angelina V. Vaseva2, Richard G. Hodge2, McKenzie P. Kline3, Thomas S. K. Gilbert1,4, Government Works Vikas Tyagi5, Daowei Huang5, Gabrielle C. Whiten5, Jacob E. Larson5, Xiaodong Wang2,5, Kenneth H. Pearce5, Laura E. Herring1,4, Lee M. Graves1,2,4, Stephen V. Frye2,5, Michael J. Emanuele1,2, Adrienne D. Cox1,2,6, Channing J. Der1,2* Stabilization of the MYC oncoprotein by KRAS signaling critically promotes the growth of pancreatic ductal adeno- carcinoma (PDAC). Thus, understanding how MYC protein stability is regulated may lead to effective therapies. Here, we used a previously developed, flow cytometry–based assay that screened a library of >800 protein kinase inhibitors and identified compounds that promoted either the stability or degradation of MYC in a KRAS-mutant PDAC cell line. We validated compounds that stabilized or destabilized MYC and then focused on one compound, Downloaded from UNC10112785, that induced the substantial loss of MYC protein in both two-dimensional (2D) and 3D cell cultures. We determined that this compound is a potent CDK9 inhibitor with a previously uncharacterized scaffold, caused MYC loss through both transcriptional and posttranslational mechanisms, and suppresses PDAC anchorage- dependent and anchorage-independent growth. We discovered that CDK9 enhanced MYC protein stability 62 through a previously unknown, KRAS-independent mechanism involving direct phosphorylation of MYC at Ser . -

Regulation of Calmodulin-Stimulated Cyclic Nucleotide Phosphodiesterase (PDE1): Review
95-105 5/6/06 13:44 Page 95 INTERNATIONAL JOURNAL OF MOLECULAR MEDICINE 18: 95-105, 2006 95 Regulation of calmodulin-stimulated cyclic nucleotide phosphodiesterase (PDE1): Review RAJENDRA K. SHARMA, SHANKAR B. DAS, ASHAKUMARY LAKSHMIKUTTYAMMA, PONNIAH SELVAKUMAR and ANURAAG SHRIVASTAV Department of Pathology and Laboratory Medicine, College of Medicine, University of Saskatchewan, Cancer Research Division, Saskatchewan Cancer Agency, 20 Campus Drive, Saskatoon SK S7N 4H4, Canada Received January 16, 2006; Accepted March 13, 2006 Abstract. The response of living cells to change in cell 6. Differential inhibition of PDE1 isozymes and its environment depends on the action of second messenger therapeutic applications molecules. The two second messenger molecules cAMP and 7. Role of proteolysis in regulating PDE1A2 Ca2+ regulate a large number of eukaryotic cellular events. 8. Role of PDE1A1 in ischemic-reperfused heart Calmodulin-stimulated cyclic nucleotide phosphodiesterase 9. Conclusion (PDE1) is one of the key enzymes involved in the complex interaction between cAMP and Ca2+ second messenger systems. Some PDE1 isozymes have similar kinetic and 1. Introduction immunological properties but are differentially regulated by Ca2+ and calmodulin. Accumulating evidence suggests that the A variety of cellular activities are regulated through mech- activity of PDE1 is selectively regulated by cross-talk between anisms controlling the level of cyclic nucleotides. These Ca2+ and cAMP signalling pathways. These isozymes are mechanisms include synthesis, degradation, efflux and seque- also further distinguished by various pharmacological agents. stration of cyclic adenosine 3':5'-monophosphate (cAMP) and We have demonstrated a potentially novel regulation of PDE1 cyclic guanosine 3':5'- monophosphate (cGMP) within the by calpain. -

Cellular and Molecular Signatures in the Disease Tissue of Early
Cellular and Molecular Signatures in the Disease Tissue of Early Rheumatoid Arthritis Stratify Clinical Response to csDMARD-Therapy and Predict Radiographic Progression Frances Humby1,* Myles Lewis1,* Nandhini Ramamoorthi2, Jason Hackney3, Michael Barnes1, Michele Bombardieri1, Francesca Setiadi2, Stephen Kelly1, Fabiola Bene1, Maria di Cicco1, Sudeh Riahi1, Vidalba Rocher-Ros1, Nora Ng1, Ilias Lazorou1, Rebecca E. Hands1, Desiree van der Heijde4, Robert Landewé5, Annette van der Helm-van Mil4, Alberto Cauli6, Iain B. McInnes7, Christopher D. Buckley8, Ernest Choy9, Peter Taylor10, Michael J. Townsend2 & Costantino Pitzalis1 1Centre for Experimental Medicine and Rheumatology, William Harvey Research Institute, Barts and The London School of Medicine and Dentistry, Queen Mary University of London, Charterhouse Square, London EC1M 6BQ, UK. Departments of 2Biomarker Discovery OMNI, 3Bioinformatics and Computational Biology, Genentech Research and Early Development, South San Francisco, California 94080 USA 4Department of Rheumatology, Leiden University Medical Center, The Netherlands 5Department of Clinical Immunology & Rheumatology, Amsterdam Rheumatology & Immunology Center, Amsterdam, The Netherlands 6Rheumatology Unit, Department of Medical Sciences, Policlinico of the University of Cagliari, Cagliari, Italy 7Institute of Infection, Immunity and Inflammation, University of Glasgow, Glasgow G12 8TA, UK 8Rheumatology Research Group, Institute of Inflammation and Ageing (IIA), University of Birmingham, Birmingham B15 2WB, UK 9Institute of -
Protein Kinases Phosphorylation/Dephosphorylation Protein Phosphorylation Is One of the Most Important Mechanisms of Cellular Re
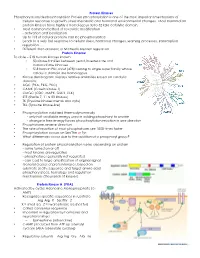
Protein Kinases Phosphorylation/dephosphorylation Protein phosphorylation is one of the most important mechanisms of cellular responses to growth, stress metabolic and hormonal environmental changes. Most mammalian protein kinases have highly a homologous 30 to 32 kDa catalytic domain. • Most common method of reversible modification - activation and localization • Up to 1/3 of cellular proteins can be phosphorylated • Leads to a very fast response to cellular stress, hormonal changes, learning processes, transcription regulation .... • Different than allosteric or Michealis Menten regulation Protein Kinome To date – 518 human kinases known • 50 kinase families between yeast, invertebrate and mammaliane kinomes • 518 human PKs, most (478) belong to single super family whose catalytic domain are homologous. • Kinase dendrogram displays relative similarities based on catalytic domains. • AGC (PKA, PKG, PKC) • CAMK (Casein kinase 1) • CMGC (CDC, MAPK, GSK3, CLK) • STE (Sterile 7, 11 & 20 kinases) • TK (Tryosine kinases memb and cyto) • TKL (Tyrosine kinase-like) • Phosphorylation stabilized thermodynamically - only half available energy used in adding phosphoryl to protein - change in free energy forces phosphorylation reaction in one direction • Phosphatases reverse direction • The rate of reaction of most phosphatases are 1000 times faster • Phosphorylation occurs on Ser/The or Tyr • What differences occur due to the addition of a phosphoryl group? • Regulation of protein phosphorylation varies depending on protein - some turned on or off -

Genomic Characterization of Additional Cancer-Driver Genes Using a Weighted Iterative Regression Accurately Modelling Background Mutation Rate
bioRxiv preprint doi: https://doi.org/10.1101/437061; this version posted October 8, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Genomic characterization of additional cancer-driver genes using a weighted iterative regression accurately modelling background mutation rate Lin Jiang1,2,#, Jingjing Zheng1, #, Johnny Sheung Him Kwan9,10,11,#, Sheng Dai1, Cong Li1, Ka Fai TO9,10,11, Pak Chung Sham5,6,7,8,*, Yonghong Zhu1,2,* and Miaoxin Li1,2,3,4,5,6,7* 1Zhongshan School of Medicine,2First Affiliated Hospital, 3Center for Genome Research, 4Center for Precision Medicine, Sun Yat-sen University, Guangzhou 510080, China; 5Key Laboratory of Tropical Disease Control (SYSU), Ministry of Education, Guangzhou 510080, China; 6The Centre for Genomic Sciences, 7Department of Psychiatry, 8State Key Laboratory for Cognitive and Brain Sciences, the University of Hong Kong, Pokfulam, Hong Kong; 9Department of Anatomical and Cellular Pathology, 10State Key Laboratory in Oncology in South China, 11Li Ka-Shing Institute of Health Sciences, The Chinese University of Hong Kong, New Territories, Hong Kong Short title: a powerful approach to detect cancer-driver genes * To whom correspondence should be addressed to Miaoxin Li. Tel: +86-2087335080; Email: [email protected] Medical Science and Technology Building, Zhongshan School of Medicine, Sun Yat-sen University, China. Yonghong Zhu. Tel: +86-20 87331451; Email: [email protected] Medical Science and Technology Building, Zhongshan School of Medicine, Sun Yat-sen University, China. Pak-Chung Sham. Tel: +852-28315425; Fax: +852-28185653; Email: [email protected]; The University of Hong Kong, 5 Sassoon Road, Pokfulam Hong Kong SAR”. -

Supplementary Table 1. in Vitro Side Effect Profiling Study for LDN/OSU-0212320. Neurotransmitter Related Steroids
Supplementary Table 1. In vitro side effect profiling study for LDN/OSU-0212320. Percent Inhibition Receptor 10 µM Neurotransmitter Related Adenosine, Non-selective 7.29% Adrenergic, Alpha 1, Non-selective 24.98% Adrenergic, Alpha 2, Non-selective 27.18% Adrenergic, Beta, Non-selective -20.94% Dopamine Transporter 8.69% Dopamine, D1 (h) 8.48% Dopamine, D2s (h) 4.06% GABA A, Agonist Site -16.15% GABA A, BDZ, alpha 1 site 12.73% GABA-B 13.60% Glutamate, AMPA Site (Ionotropic) 12.06% Glutamate, Kainate Site (Ionotropic) -1.03% Glutamate, NMDA Agonist Site (Ionotropic) 0.12% Glutamate, NMDA, Glycine (Stry-insens Site) 9.84% (Ionotropic) Glycine, Strychnine-sensitive 0.99% Histamine, H1 -5.54% Histamine, H2 16.54% Histamine, H3 4.80% Melatonin, Non-selective -5.54% Muscarinic, M1 (hr) -1.88% Muscarinic, M2 (h) 0.82% Muscarinic, Non-selective, Central 29.04% Muscarinic, Non-selective, Peripheral 0.29% Nicotinic, Neuronal (-BnTx insensitive) 7.85% Norepinephrine Transporter 2.87% Opioid, Non-selective -0.09% Opioid, Orphanin, ORL1 (h) 11.55% Serotonin Transporter -3.02% Serotonin, Non-selective 26.33% Sigma, Non-Selective 10.19% Steroids Estrogen 11.16% 1 Percent Inhibition Receptor 10 µM Testosterone (cytosolic) (h) 12.50% Ion Channels Calcium Channel, Type L (Dihydropyridine Site) 43.18% Calcium Channel, Type N 4.15% Potassium Channel, ATP-Sensitive -4.05% Potassium Channel, Ca2+ Act., VI 17.80% Potassium Channel, I(Kr) (hERG) (h) -6.44% Sodium, Site 2 -0.39% Second Messengers Nitric Oxide, NOS (Neuronal-Binding) -17.09% Prostaglandins Leukotriene, -

Rho-Associated Kinase 2 Polymorphism of Vasospastic
Editorial http://dx.doi.org/10.4070/kcj.2012.42.6.379 Print ISSN 1738-5520 • On-line ISSN 1738-5555 Korean Circulation Journal Rho-Associated Kinase 2 Polymorphism of Vasospastic Angina in Korean Population Chul Soo Park, MD Division of Cardiology, Department of Internal Medicine, College of Medicine, The Catholic University of Korea, Yeouido St. Mary’s Hospital, Seoul, Korea Refer to the page 406-413 periments, we can speculate that an increased activity of ROCKs can be one of the important pathophysiologic mechanisms of vaso- spastic angina, and fasudil, a ROCK inhibitor has exerted beneficial Rho-associated kinases (ROCKs), the immediate downstream effect on VA.4) targets of RhoA, are ubiquitously expressed serine-threonine pro- Rho-associated kinases could regulate other cellular functions tein kinases, which are involved in diverse cellular functions, includ- that are independent of their effects on the actin cytoskeleton. For ing smooth muscle contraction, actin cytoskeleton organization, cell examples, Rock inhibits insulin signaling, reduces cardiac hypertro- adhesion and motility, and gene expression. Recent studies have phy and involves an in tissue differentiation from adipocyte to myo- shown that ROCKs may play a pivotal role in cardiovascular diseases, cyte. such as vasospastic angina, ischemic stroke, and heart failure. In addition to ROCK’s effect on actin cytoskeleton system, inhibi- Rho-associated kinases are important regulators of cellular apop- tory effect on endothelial nitric oxide synthase (eNOS) activity can tosis, growth, metabolism, and migration via control of the actin cy- be another mechanism involved in the pathogenesis of vasospasm. toskeletal assembly and cell contraction.1) ROCKs phosphorylate var- Enothelium derived NO plays an important role in the regulation ious targets and mediate broad range of cellular responses that in- of vascular tone, inhibition of platelet aggregation, and the suppres- volve in actin cytoskeleton in response to GTPase-RhoA.