Discovery of a Polyomavirus in European Badgers (Meles Meles) and the Evolution of Host Range in the Family Polyomaviridae
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

And Giant Guitarfish (Rhynchobatus Djiddensis)
VIRAL DISCOVERY IN BLUEGILL SUNFISH (LEPOMIS MACROCHIRUS) AND GIANT GUITARFISH (RHYNCHOBATUS DJIDDENSIS) BY HISTOPATHOLOGY EVALUATION, METAGENOMIC ANALYSIS AND NEXT GENERATION SEQUENCING by JENNIFER ANNE DILL (Under the Direction of Alvin Camus) ABSTRACT The rapid growth of aquaculture production and international trade in live fish has led to the emergence of many new diseases. The introduction of novel disease agents can result in significant economic losses, as well as threats to vulnerable wild fish populations. Losses are often exacerbated by a lack of agent identification, delay in the development of diagnostic tools and poor knowledge of host range and susceptibility. Examples in bluegill sunfish (Lepomis macrochirus) and the giant guitarfish (Rhynchobatus djiddensis) will be discussed here. Bluegill are popular freshwater game fish, native to eastern North America, living in shallow lakes, ponds, and slow moving waterways. Bluegill experiencing epizootics of proliferative lip and skin lesions, characterized by epidermal hyperplasia, papillomas, and rarely squamous cell carcinoma, were investigated in two isolated poopulations. Next generation genomic sequencing revealed partial DNA sequences of an endogenous retrovirus and the entire circular genome of a novel hepadnavirus. Giant Guitarfish, a rajiform elasmobranch listed as ‘vulnerable’ on the IUCN Red List, are found in the tropical Western Indian Ocean. Proliferative skin lesions were observed on the ventrum and caudal fin of a juvenile male quarantined at a public aquarium following international shipment. Histologically, lesions consisted of papillomatous epidermal hyperplasia with myriad large, amphophilic, intranuclear inclusions. Deep sequencing and metagenomic analysis produced the complete genomes of two novel DNA viruses, a typical polyomavirus and a second unclassified virus with a 20 kb genome tentatively named Colossomavirus. -

Opportunistic Intruders: How Viruses Orchestrate ER Functions to Infect Cells
REVIEWS Opportunistic intruders: how viruses orchestrate ER functions to infect cells Madhu Sudhan Ravindran*, Parikshit Bagchi*, Corey Nathaniel Cunningham and Billy Tsai Abstract | Viruses subvert the functions of their host cells to replicate and form new viral progeny. The endoplasmic reticulum (ER) has been identified as a central organelle that governs the intracellular interplay between viruses and hosts. In this Review, we analyse how viruses from vastly different families converge on this unique intracellular organelle during infection, co‑opting some of the endogenous functions of the ER to promote distinct steps of the viral life cycle from entry and replication to assembly and egress. The ER can act as the common denominator during infection for diverse virus families, thereby providing a shared principle that underlies the apparent complexity of relationships between viruses and host cells. As a plethora of information illuminating the molecular and cellular basis of virus–ER interactions has become available, these insights may lead to the development of crucial therapeutic agents. Morphogenesis Viruses have evolved sophisticated strategies to establish The ER is a membranous system consisting of the The process by which a virus infection. Some viruses bind to cellular receptors and outer nuclear envelope that is contiguous with an intri‑ particle changes its shape and initiate entry, whereas others hijack cellular factors that cate network of tubules and sheets1, which are shaped by structure. disassemble the virus particle to facilitate entry. After resident factors in the ER2–4. The morphology of the ER SEC61 translocation delivering the viral genetic material into the host cell and is highly dynamic and experiences constant structural channel the translation of the viral genes, the resulting proteins rearrangements, enabling the ER to carry out a myriad An endoplasmic reticulum either become part of a new virus particle (or particles) of functions5. -

Viruses in Transplantation - Not Always Enemies
Viruses in transplantation - not always enemies Virome and transplantation ECCMID 2018 - Madrid Prof. Laurent Kaiser Head Division of Infectious Diseases Laboratory of Virology Geneva Center for Emerging Viral Diseases University Hospital of Geneva ESCMID eLibrary © by author Conflict of interest None ESCMID eLibrary © by author The human virome: definition? Repertoire of viruses found on the surface of/inside any body fluid/tissue • Eukaryotic DNA and RNA viruses • Prokaryotic DNA and RNA viruses (phages) 25 • The “main” viral community (up to 10 bacteriophages in humans) Haynes M. 2011, Metagenomic of the human body • Endogenous viral elements integrated into host chromosomes (8% of the human genome) • NGS is shaping the definition Rascovan N et al. Annu Rev Microbiol 2016;70:125-41 Popgeorgiev N et al. Intervirology 2013;56:395-412 Norman JM et al. Cell 2015;160:447-60 ESCMID eLibraryFoxman EF et al. Nat Rev Microbiol 2011;9:254-64 © by author Viruses routinely known to cause diseases (non exhaustive) Upper resp./oropharyngeal HSV 1 Influenza CNS Mumps virus Rhinovirus JC virus RSV Eye Herpes viruses Parainfluenza HSV Measles Coronavirus Adenovirus LCM virus Cytomegalovirus Flaviviruses Rabies HHV6 Poliovirus Heart Lower respiratory HTLV-1 Coxsackie B virus Rhinoviruses Parainfluenza virus HIV Coronaviruses Respiratory syncytial virus Parainfluenza virus Adenovirus Respiratory syncytial virus Coronaviruses Gastro-intestinal Influenza virus type A and B Human Bocavirus 1 Adenovirus Hepatitis virus type A, B, C, D, E Those that cause -

Diversity and Evolution of Viral Pathogen Community in Cave Nectar Bats (Eonycteris Spelaea)
viruses Article Diversity and Evolution of Viral Pathogen Community in Cave Nectar Bats (Eonycteris spelaea) Ian H Mendenhall 1,* , Dolyce Low Hong Wen 1,2, Jayanthi Jayakumar 1, Vithiagaran Gunalan 3, Linfa Wang 1 , Sebastian Mauer-Stroh 3,4 , Yvonne C.F. Su 1 and Gavin J.D. Smith 1,5,6 1 Programme in Emerging Infectious Diseases, Duke-NUS Medical School, Singapore 169857, Singapore; [email protected] (D.L.H.W.); [email protected] (J.J.); [email protected] (L.W.); [email protected] (Y.C.F.S.) [email protected] (G.J.D.S.) 2 NUS Graduate School for Integrative Sciences and Engineering, National University of Singapore, Singapore 119077, Singapore 3 Bioinformatics Institute, Agency for Science, Technology and Research, Singapore 138671, Singapore; [email protected] (V.G.); [email protected] (S.M.-S.) 4 Department of Biological Sciences, National University of Singapore, Singapore 117558, Singapore 5 SingHealth Duke-NUS Global Health Institute, SingHealth Duke-NUS Academic Medical Centre, Singapore 168753, Singapore 6 Duke Global Health Institute, Duke University, Durham, NC 27710, USA * Correspondence: [email protected] Received: 30 January 2019; Accepted: 7 March 2019; Published: 12 March 2019 Abstract: Bats are unique mammals, exhibit distinctive life history traits and have unique immunological approaches to suppression of viral diseases upon infection. High-throughput next-generation sequencing has been used in characterizing the virome of different bat species. The cave nectar bat, Eonycteris spelaea, has a broad geographical range across Southeast Asia, India and southern China, however, little is known about their involvement in virus transmission. -

Diversity and Evolution of Novel Invertebrate DNA Viruses Revealed by Meta-Transcriptomics
viruses Article Diversity and Evolution of Novel Invertebrate DNA Viruses Revealed by Meta-Transcriptomics Ashleigh F. Porter 1, Mang Shi 1, John-Sebastian Eden 1,2 , Yong-Zhen Zhang 3,4 and Edward C. Holmes 1,3,* 1 Marie Bashir Institute for Infectious Diseases and Biosecurity, Charles Perkins Centre, School of Life & Environmental Sciences and Sydney Medical School, The University of Sydney, Sydney, NSW 2006, Australia; [email protected] (A.F.P.); [email protected] (M.S.); [email protected] (J.-S.E.) 2 Centre for Virus Research, Westmead Institute for Medical Research, Westmead, NSW 2145, Australia 3 Shanghai Public Health Clinical Center and School of Public Health, Fudan University, Shanghai 201500, China; [email protected] 4 Department of Zoonosis, National Institute for Communicable Disease Control and Prevention, Chinese Center for Disease Control and Prevention, Changping, Beijing 102206, China * Correspondence: [email protected]; Tel.: +61-2-9351-5591 Received: 17 October 2019; Accepted: 23 November 2019; Published: 25 November 2019 Abstract: DNA viruses comprise a wide array of genome structures and infect diverse host species. To date, most studies of DNA viruses have focused on those with the strongest disease associations. Accordingly, there has been a marked lack of sampling of DNA viruses from invertebrates. Bulk RNA sequencing has resulted in the discovery of a myriad of novel RNA viruses, and herein we used this methodology to identify actively transcribing DNA viruses in meta-transcriptomic libraries of diverse invertebrate species. Our analysis revealed high levels of phylogenetic diversity in DNA viruses, including 13 species from the Parvoviridae, Circoviridae, and Genomoviridae families of single-stranded DNA virus families, and six double-stranded DNA virus species from the Nudiviridae, Polyomaviridae, and Herpesviridae, for which few invertebrate viruses have been identified to date. -

Virus World As an Evolutionary Network of Viruses and Capsidless Selfish Elements
Virus World as an Evolutionary Network of Viruses and Capsidless Selfish Elements Koonin, E. V., & Dolja, V. V. (2014). Virus World as an Evolutionary Network of Viruses and Capsidless Selfish Elements. Microbiology and Molecular Biology Reviews, 78(2), 278-303. doi:10.1128/MMBR.00049-13 10.1128/MMBR.00049-13 American Society for Microbiology Version of Record http://cdss.library.oregonstate.edu/sa-termsofuse Virus World as an Evolutionary Network of Viruses and Capsidless Selfish Elements Eugene V. Koonin,a Valerian V. Doljab National Center for Biotechnology Information, National Library of Medicine, Bethesda, Maryland, USAa; Department of Botany and Plant Pathology and Center for Genome Research and Biocomputing, Oregon State University, Corvallis, Oregon, USAb Downloaded from SUMMARY ..................................................................................................................................................278 INTRODUCTION ............................................................................................................................................278 PREVALENCE OF REPLICATION SYSTEM COMPONENTS COMPARED TO CAPSID PROTEINS AMONG VIRUS HALLMARK GENES.......................279 CLASSIFICATION OF VIRUSES BY REPLICATION-EXPRESSION STRATEGY: TYPICAL VIRUSES AND CAPSIDLESS FORMS ................................279 EVOLUTIONARY RELATIONSHIPS BETWEEN VIRUSES AND CAPSIDLESS VIRUS-LIKE GENETIC ELEMENTS ..............................................280 Capsidless Derivatives of Positive-Strand RNA Viruses....................................................................................................280 -

Chronic Viral Infections Vs. Our Immune System: Revisiting Our View of Viruses As Pathogens
Chronic Viral Infections vs. Our Immune System: Revisiting our view of viruses as pathogens Tiffany A. Reese Assistant Professor Departments of Immunology and Microbiology Challenge your idea of classic viral infection and disease • Define the microbiome and the virome • Brief background on persistent viruses • Illustrate how viruses change disease susceptibility – mutualistic symbiosis – gene + virus = disease phenotype – virome in immune responses Bacteria-centric view of the microbiome The microbiome defined Definition of microbiome – Merriam-Webster 1 :a community of microorganisms (such as bacteria, fungi, and viruses) that inhabit a particular environment and especially the collection of microorganisms living in or on the human body 2 :the collective genomes of microorganisms inhabiting a particular environment and especially the human body Virome Ø Viral component of the microbiome Ø Includes both commensal and pathogenic viruses Ø Viruses that infect host cells Ø Virus-derived elements in host chromosomes Ø Viruses that infect other organisms in the body e.g. phage/bacteria Viruses are everywhere! • “intracellular parasites with nucleic acids that are capable of directing their own replication and are not cells” – Roossinck, Nature Reviews Microbiology 2011. • Viruses infect all living things. • We are constantly eating and breathing viruses from our environment • Only a small subset of viruses cause disease. • We even carry viral genomes as part of our own genetic material! Diverse viruses all over the body Adenoviridae Picornaviridae -

Alma Mater Studiorum – Università Di Bologna
Alma Mater Studiorum – Università di Bologna DOTTORATO DI RICERCA IN Scienze Biotecnologiche e Farmaceutiche Ciclo XXXI Settore Concorsuale: 06/A3 - MICROBIOLOGIA E MICROBIOLOGIA CLINICA Settore Scientifico Disciplinare: MED/07 - MICROBIOLOGIA E MICROBIOLOGIA CLINICA TITOLO TESI “Human Parvovirus B19: from the development of a reverse genetics system to antiviral strategies” Presentata da: Ilaria Conti Matricola n. 0000772499 Coordinatore Dottorato Supervisore Chiar.mo Prof. Chiar.mo Prof. Santi Mario Spampinato Giorgio Gallinella Esame finale anno 2019 Abstract Dott. Ilaria Conti Supervisor: Prof. Giorgio Gallinella PhD course: Scienze Biotecnologiche e Farmaceutiche, XXXI ciclo Thesis: “Human Parvovirus B19: from the development of a reverse genetics system to antiviral strategies” Human Parvovirus B19 (B19V) is a human pathogenic virus which belongs to the Parvoviridae family. It is worldwide distributed and is responsible of various clinical manifestations in human, although neither an antiviral therapy nor a vaccine are available. The virus is not well adapted to grow in cellular cultures and this causes difficulties for its propagation, maintaining and characterization. B19V has a narrow tropism for the erythroid progenitor cells of human bone marrow and very few cellular systems can support the viral replication (such as, UT7/EpoS1 cells which is a megakaryoblastoid cell line). In this research, a reverse genetic approach was developed to allow the generation of mature and infectious viral particles from an established consensus sequence. Synthetic clones that differ for the length and isomerism of their terminal regions (ITRs) were constructed. After their transfection in UT7/EpoS1 cells, the obtained viral particles were used to infect EPCs (erythroid progenitor cells) in serial passages, in order to evaluate the capability for each clone to generate a new viral stock and to propagate it. -
Supporting Information
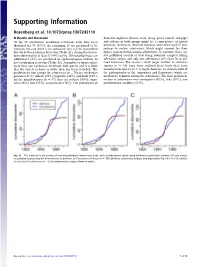
Supporting Information Rosenberg et al. 10.1073/pnas.1307243110 SI Results and Discussion domestic ungulates (horses, cows, sheep, goats, camels, and pigs) Of the 83 arboviruses, nonhuman vertebrate hosts have been and rodents in both groups might be a consequence of spatial identified for 70 (84%); the remaining 13 are presumed to be proximity to humans. Sentinel monkeys were often used in pro- zoonoses because there is no indication they can be transmitted cedures to isolate arboviruses, which might account for their directly between humans by vectors (Table S1). Animal hosts have higher representation among arboviruses. In contrast, there are been identified for at least 57 (44%) of the 130 nonarboviruses; an few published records of bats being routinely sampled during additional 5 (8%) are presumed on epidemiological evidence to arbovirus studies, and only two arboviruses (3%) have been iso- have nonhuman reservoirs (Table S1). A number of viruses infect lated from bats. The reason a much larger number of arbovirus more than one nonhuman vertebrate host species and it is likely species (n = 16) have been isolated from birds than have that the variety of hosts is wider than has been recorded. The nonarbovirus species (n = 1) might, however, be characteristic of predominant host groups for arboviruses (n = 70) are nonhuman the pathogenicity of the togaviruses and flaviviruses, which are primates (31%), rodents (29%), ungulates (26%), and birds (23%); much more common among the arboviruses. The most prominent for the nonarboviruses (n = 57), they are rodents (30%), ungu- vectors of arboviruses were mosquitoes (67%), ticks (19%), and lates (26%), bats (23%), and primates (16%). -

Foodborne Viruses
Available online at www.sciencedirect.com ScienceDirect Foodborne viruses 1,2 1,2 1,2 Albert Bosch , Rosa M Pinto´ and Susana Guix Among the wide variety of viral agents liable to be found as food mortality, although the actual global burden of unsafe contaminants, noroviruses and hepatitis A virus are responsible food consumption remains hard to estimate [1]. Several for most well characterized foodborne virus outbreaks. factors, among them the increasing population and the Additionally, hepatitis E virus has emerged as a potential demand for continuous availability of seasonal products zoonotic threat.Molecular methods, including an ISO standard, all year-around, lead to global food trade among regions are available for norovirus and hepatitis A virus detection in with different hygienic standards and the vulnerability of foodstuffs, although the significance of genome copy the food supply. detection with regard to the associated health risk is yet to be determined through viability assays.More precise and rapid The World Health Organization (WHO) Foodborne Dis- methods for early foodborne outbreak investigation are ease Burden Epidemiology Reference Group provided in being developed and they will need to be validated versus 2015 the first estimates of global foodborne disease inci- the ISO standard. In addition, protocols for next-generation dence, mortality, and disease burden in terms of Disability sequencing characterization of outbreak-related samples Adjusted Life Years (DALYs) [1]. The global burden of must be developed, harmonized and validated as well. foodborne hazards was 33 million DALYs in 2010 (95% Addresses uncertainty interval [UI] 25–46); 40% affecting children 1 Enteric Virus Group, Department of Microbiology, University of under 5 years of age. -

Packaging of Genomic RNA in Positive-Sense Single-Stranded RNA Viruses: a Complex Story
viruses Review Packaging of Genomic RNA in Positive-Sense Single-Stranded RNA Viruses: A Complex Story Mauricio Comas-Garcia 1,2 1 Research Center for Health Sciences and Biomedicine (CICSaB), Universidad Autónoma de San Luis Potosí (UASLP), Av. Sierra Leona 550 Lomas 2da Seccion, 72810 San Luis Potosi, Mexico; [email protected] 2 Department of Sciences, Universidad Autónoma de San Luis Potosí (UASLP), Av. Chapultepec 1570, Privadas del Pedregal, 78295 San Luis Potosi, Mexico Received: 14 February 2019; Accepted: 8 March 2019; Published: 13 March 2019 Abstract: The packaging of genomic RNA in positive-sense single-stranded RNA viruses is a key part of the viral infectious cycle, yet this step is not fully understood. Unlike double-stranded DNA and RNA viruses, this process is coupled with nucleocapsid assembly. The specificity of RNA packaging depends on multiple factors: (i) one or more packaging signals, (ii) RNA replication, (iii) translation, (iv) viral factories, and (v) the physical properties of the RNA. The relative contribution of each of these factors to packaging specificity is different for every virus. In vitro and in vivo data show that there are different packaging mechanisms that control selective packaging of the genomic RNA during nucleocapsid assembly. The goals of this article are to explain some of the key experiments that support the contribution of these factors to packaging selectivity and to draw a general scenario that could help us move towards a better understanding of this step of the viral infectious cycle. Keywords: (+)ssRNA viruses; RNA packaging; virion assembly; packaging signals; RNA replication 1. Introduction Nucleocapsid assembly and the RNA replication of positive-sense single-stranded RNA [(+)ssRNA] viruses occur in the cytoplasm. -

USMLE Step 1: Predict Viral Replication in Enucleated Cells
USMLE Step 1: Predict viral replication in enucleated cells DEC 8, 2016 Staff News Writer If you’re preparing for the United States Medical Licensing Examination® (USMLE®) Step 1 exam, you might want to know which questions are most often missed by test takers. Check out this example from Kaplan Medical, and view an expert video explanation of the answer. Also check out all posts in this series. This month’s stumper An investigator is studying virus life cycles. He creates a continuous culture of kidney fibroblasts that are suitable hosts for a large variety of viral agents. In one experiment, the nuclei of these cells are removed by cytosurgery, and various viral agents are added to the cultures. Following culture of the viruses with the enucleated cells, the yield of cytopathic units of virus is quantified. Which of the following viruses would be capable of replication in these enucleated cells? A. Adenovirus B. Cytomegalovirus C. Influenza virus D. JC virus E. Poliovirus URL: https://www.ama-assn.org/residents-students/usmle/usmle-step-1-predict-viral-replication-enucleated-cells Copyright 1995 - 2021 American Medical Association. All rights reserved. The correct answer is E. Kaplan Medical explains why Most RNA viruses—for example, poliovirus—replicate in the cytoplasm and therefore can replicate in enucleated cells. Poliovirus belongs to the family Picornaviridae. These viruses are nonenveloped and have an icosahedral nucleocapsid that contains positive-sense RNA. Why the other answers are wrong Read these explanations to understand the important rationale for why each answer is incorrect. Choice A: Adenoviruses are non-enveloped and have an icosahedral nucleocapsid that contains a double-stranded linear DNA genome.