Integrated Analysis of Differentially Expressed Genes in Breast Cancer Pathogenesis
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Analysis and Characterisation of the Mouse Hic2 Gene
Aus dem Institut für Entwicklungsgenetik des GSF-Forschungszentrums für Umwelt und Gesundheit, GmbH Direktor: Prof. Dr. Wolfgang Wurst Anfertigung unter der Leitung von Prof. Dr. Jochen Graw Vorgelegt über den Lehrstuhl für Molekulare Tierzucht und Biotechnologie Der Tierärztlichen Fakultät der Ludwig-Maximilians-Universität München Vorstand: Prof. Dr. Eckhard Wolf Untersuchung und Charakterisierung des Hic2-Gens der Maus Inaugural-Dissertation Zur Erlangung der tiermedizinischen Doktorwürde der Tierärztlichen Fakultät der Ludwig-Maximilians-Universität München von Aleksandra Terzic aus Sarajevo/Bosnia und Herzegowina München 2004 II Aus dem Institut für Entwicklungsgenetik des GSF-Forschungszentrums für Umwelt und Gesundheit, GmbH Direktor: Prof. Dr. Wolfgang Wurst Anfertigung unter der Leitung von Prof. Dr. Jochen Graw Vorgelegt über den Lehrstuhl für Molekulare Tierzucht und Biotechnologie Der Tierärztlichen Fakultät der Ludwig-Maximilians-Universität München Vorstand: Prof. Dr. Eckhard Wolf Analysis and characterisation of the mouse Hic2 gene Inaugural-Dissertation Zur Erlangung der tiermedizinischen Doktorwürde der Tierärztlichen Fakultät der Ludwig-Maximilians-Universität München von Aleksandra Terzic aus Sarajevo/Bosnia und Herzegowina München 2004 III Gedruckt mit Genehmigung der Tierärztlichen Fakultät der Ludwig-Maximilians-Universität München Dekan: Univ.-Prof. Dr. A.Stolle Referent: Univ.-Prof. Dr. E. Wolf Korreferent: Univ.-Prof. Dr. K. Heinritzi Tag der Promotion: 13. Februar 2004 IV List of contents 1 INTRODUCTION.............................................................................................................1 -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Nucleolin and Its Role in Ribosomal Biogenesis
NUCLEOLIN: A NUCLEOLAR RNA-BINDING PROTEIN INVOLVED IN RIBOSOME BIOGENESIS Inaugural-Dissertation zur Erlangung des Doktorgrades der Mathematisch-Naturwissenschaftlichen Fakultät der Heinrich-Heine-Universität Düsseldorf vorgelegt von Julia Fremerey aus Hamburg Düsseldorf, April 2016 2 Gedruckt mit der Genehmigung der Mathematisch-Naturwissenschaftlichen Fakultät der Heinrich-Heine-Universität Düsseldorf Referent: Prof. Dr. A. Borkhardt Korreferent: Prof. Dr. H. Schwender Tag der mündlichen Prüfung: 20.07.2016 3 Die vorgelegte Arbeit wurde von Juli 2012 bis März 2016 in der Klinik für Kinder- Onkologie, -Hämatologie und Klinische Immunologie des Universitätsklinikums Düsseldorf unter Anleitung von Prof. Dr. A. Borkhardt und in Kooperation mit dem ‚Laboratory of RNA Molecular Biology‘ an der Rockefeller Universität unter Anleitung von Prof. Dr. T. Tuschl angefertigt. 4 Dedicated to my family TABLE OF CONTENTS 5 TABLE OF CONTENTS TABLE OF CONTENTS ............................................................................................... 5 LIST OF FIGURES ......................................................................................................10 LIST OF TABLES .......................................................................................................12 ABBREVIATION .........................................................................................................13 ABSTRACT ................................................................................................................19 ZUSAMMENFASSUNG -

Differences in Aggressive Behavior and DNA Copy Number Variants Between BALB/Cj and BALB/Cbyj Substrains
Behav Genet (2010) 40:201–210 DOI 10.1007/s10519-009-9325-5 ORIGINAL RESEARCH Differences in Aggressive Behavior and DNA Copy Number Variants Between BALB/cJ and BALB/cByJ Substrains Lady Velez • Greta Sokoloff • Klaus A. Miczek • Abraham A. Palmer • Stephanie C. Dulawa Received: 3 February 2009 / Accepted: 3 December 2009 / Published online: 23 December 2009 Ó Springer Science+Business Media, LLC 2009 Abstract Some BALB/c substrains exhibit different Keywords Aggression Á Resident intruder Á BALB/c Á levels of aggression. We compared aggression levels CNV Á Substrain between male BALB/cJ and BALB/cByJ substrains using the resident intruder paradigm. These substrains were also assessed in other tests of emotionality and information Introduction processing including the open field, forced swim, fear conditioning, and prepulse inhibition tests. We also eval- Some substrains of BALB/c mice have previously been uated single nucleotide polymorphisms (SNPs) previously suggested to exhibit robust differences in aggressive reported between these BALB/c substrains. Finally, we behavior, although the genetic basis for this behavioral dif- compared BALB/cJ and BALB/cByJ mice for genomic ference has not been identified (Ciaranello et al. 1974). The deletions or duplications, collectively termed copy number BALB/c substrains were derived from an initial BALB/c variants (CNVs), to identify candidate genes that might stock established by 1935 (Les 1990); this BALB/c stock underlie the observed behavioral differences. BALB/cJ was then acquired by several other laboratories and were mice showed substantially higher aggression levels than maintained and bred as independent stocks including BALB/ BALB/cByJ mice; however, only minor differences in cJ, BALB/cN, and BALB/cByJ. -

Appendix 2. Significantly Differentially Regulated Genes in Term Compared with Second Trimester Amniotic Fluid Supernatant
Appendix 2. Significantly Differentially Regulated Genes in Term Compared With Second Trimester Amniotic Fluid Supernatant Fold Change in term vs second trimester Amniotic Affymetrix Duplicate Fluid Probe ID probes Symbol Entrez Gene Name 1019.9 217059_at D MUC7 mucin 7, secreted 424.5 211735_x_at D SFTPC surfactant protein C 416.2 206835_at STATH statherin 363.4 214387_x_at D SFTPC surfactant protein C 295.5 205982_x_at D SFTPC surfactant protein C 288.7 1553454_at RPTN repetin solute carrier family 34 (sodium 251.3 204124_at SLC34A2 phosphate), member 2 238.9 206786_at HTN3 histatin 3 161.5 220191_at GKN1 gastrokine 1 152.7 223678_s_at D SFTPA2 surfactant protein A2 130.9 207430_s_at D MSMB microseminoprotein, beta- 99.0 214199_at SFTPD surfactant protein D major histocompatibility complex, class II, 96.5 210982_s_at D HLA-DRA DR alpha 96.5 221133_s_at D CLDN18 claudin 18 94.4 238222_at GKN2 gastrokine 2 93.7 1557961_s_at D LOC100127983 uncharacterized LOC100127983 93.1 229584_at LRRK2 leucine-rich repeat kinase 2 HOXD cluster antisense RNA 1 (non- 88.6 242042_s_at D HOXD-AS1 protein coding) 86.0 205569_at LAMP3 lysosomal-associated membrane protein 3 85.4 232698_at BPIFB2 BPI fold containing family B, member 2 84.4 205979_at SCGB2A1 secretoglobin, family 2A, member 1 84.3 230469_at RTKN2 rhotekin 2 82.2 204130_at HSD11B2 hydroxysteroid (11-beta) dehydrogenase 2 81.9 222242_s_at KLK5 kallikrein-related peptidase 5 77.0 237281_at AKAP14 A kinase (PRKA) anchor protein 14 76.7 1553602_at MUCL1 mucin-like 1 76.3 216359_at D MUC7 mucin 7, -

GATA2 Regulates Mast Cell Identity and Responsiveness to Antigenic Stimulation by Promoting Chromatin Remodeling at Super- Enhancers
ARTICLE https://doi.org/10.1038/s41467-020-20766-0 OPEN GATA2 regulates mast cell identity and responsiveness to antigenic stimulation by promoting chromatin remodeling at super- enhancers Yapeng Li1, Junfeng Gao 1, Mohammad Kamran1, Laura Harmacek2, Thomas Danhorn 2, Sonia M. Leach1,2, ✉ Brian P. O’Connor2, James R. Hagman 1,3 & Hua Huang 1,3 1234567890():,; Mast cells are critical effectors of allergic inflammation and protection against parasitic infections. We previously demonstrated that transcription factors GATA2 and MITF are the mast cell lineage-determining factors. However, it is unclear whether these lineage- determining factors regulate chromatin accessibility at mast cell enhancer regions. In this study, we demonstrate that GATA2 promotes chromatin accessibility at the super-enhancers of mast cell identity genes and primes both typical and super-enhancers at genes that respond to antigenic stimulation. We find that the number and densities of GATA2- but not MITF-bound sites at the super-enhancers are several folds higher than that at the typical enhancers. Our studies reveal that GATA2 promotes robust gene transcription to maintain mast cell identity and respond to antigenic stimulation by binding to super-enhancer regions with dense GATA2 binding sites available at key mast cell genes. 1 Department of Immunology and Genomic Medicine, National Jewish Health, Denver, CO 80206, USA. 2 Center for Genes, Environment and Health, National Jewish Health, Denver, CO 80206, USA. 3 Department of Immunology and Microbiology, University of Colorado Anschutz Medical Campus, Aurora, ✉ CO 80045, USA. email: [email protected] NATURE COMMUNICATIONS | (2021) 12:494 | https://doi.org/10.1038/s41467-020-20766-0 | www.nature.com/naturecommunications 1 ARTICLE NATURE COMMUNICATIONS | https://doi.org/10.1038/s41467-020-20766-0 ast cells (MCs) are critical effectors in immunity that at key MC genes. -

Computational Inferences of Mutations Driving Mesenchymal Differentiation in Glioblastoma
Computational Inferences of Mutations Driving Mesenchymal Differentiation in Glioblastoma James Chen Submitted in partial fulfillment of the requirements for the Doctor of Philosophy Degree in the Graduate School of Arts and Sciences Columbia University 2013 ! 2013 James Chen All rights reserved ABSTRACT Computational Inferences of Mutations Driving Mesenchymal Differentiation in Glioblastoma James Chen This dissertation reviews the development and implementation of integrative, systems biology methods designed to parse driver mutations from high- throughput array data derived from human patients. The analysis of vast amounts of genomic and genetic data in the context of complex human genetic diseases such as Glioblastoma is a daunting task. Mutations exist by the hundreds, if not thousands, and only an unknown handful will contribute to the disease in a significant way. The goal of this project was to develop novel computational methods to identify candidate mutations from these data that drive the molecular differentiation of glioblastoma into the mesenchymal subtype, the most aggressive, poorest-prognosis tumors associated with glioblastoma. TABLE OF CONTENTS CHAPTER 1… Introduction and Background 1 Glioblastoma and the Mesenchymal Subtype 3 Systems Biology and Master Regulators 9 Thesis Project: Genetics and Genomics 20 CHAPTER 2… TCGA Data Processing 23 CHAPTER 3… DIGGIn Part 1 – Selecting f-CNVs 33 Mutual Information 40 Application and Analysis 45 CHAPTER 4… DIGGIn Part 2 – Selecting drivers 52 CHAPTER 5… KLHL9 Manuscript 63 Methods 90 CHAPTER 5a… Revisions work-in-progress 105 CHAPTER 6… Discussion 109 APPENDICES… 132 APPEND01 – TCGA classifications 133 APPEND02 – GBM f-CNV list 136 APPEND03 – MES f-CNV candidate drivers 152 APPEND04 – Scripts 149 APPEND05 – Manuscript Figures and Legends 175 APPEND06 – Manuscript Supplemental Materials 185 i ACKNOWLEDGEMENTS I would like to thank the Califano Lab and my mentor, Andrea Califano, for their intellectual and motivational support during my stay in their lab. -

Quantitative Proteomic Characterization and Comparison of T Helper 17 and Induced Regulatory T Cells
METHODS AND RESOURCES Quantitative proteomic characterization and comparison of T helper 17 and induced regulatory T cells Imran Mohammad1,2, Kari Nousiainen3, Santosh D. Bhosale1,2, Inna Starskaia1,2, Robert Moulder1, Anne Rokka1, Fang Cheng4, Ponnuswamy Mohanasundaram4, John E. Eriksson4, David R. Goodlett5, Harri LaÈhdesmaÈki3, Zhi Chen1* 1 Turku Centre for Biotechnology, University of Turku and Åbo Akademi University, Turku, Finland, 2 Turku Doctoral Programme of Molecular Medicine, University of Turku, Turku, Finland, 3 Department of Computer a1111111111 Science, Aalto University, Espoo, Finland, 4 Cell Biology, Biosciences, Faculty of Science and Engineering, a1111111111 Åbo Akademi University, Turku, Finland, 5 Department of Pharmaceutical Sciences, University of Maryland a1111111111 School of Pharmacy, Baltimore, Maryland, United States of America a1111111111 a1111111111 * [email protected] Abstract OPEN ACCESS The transcriptional network and protein regulators that govern T helper 17 (Th17) cell differ- Citation: Mohammad I, Nousiainen K, Bhosale SD, entiation have been studied extensively using advanced genomic approaches. For a better Starskaia I, Moulder R, Rokka A, et al. (2018) understanding of these biological processes, we have moved a step forward, from gene- to Quantitative proteomic characterization and protein-level characterization of Th17 cells. Mass spectrometry±based label-free quantita- comparison of T helper 17 and induced regulatory T cells. PLoS Biol 16(5): e2004194. https://doi.org/ tive (LFQ) proteomics analysis were made of in vitro differentiated murine Th17 and induced 10.1371/journal.pbio.2004194 regulatory T (iTreg) cells. More than 4,000 proteins, covering almost all subcellular compart- Academic Editor: Paula Oliver, University of ments, were detected. Quantitative comparison of the protein expression profiles resulted in Pennsylvania Perelman School of Medicine, United the identification of proteins specifically expressed in the Th17 and iTreg cells. -

Target Gene Gene Description Validation Diana Miranda
Supplemental Table S1. Mmu-miR-183-5p in silico predicted targets. TARGET GENE GENE DESCRIPTION VALIDATION DIANA MIRANDA MIRBRIDGE PICTAR PITA RNA22 TARGETSCAN TOTAL_HIT AP3M1 adaptor-related protein complex 3, mu 1 subunit V V V V V V V 7 BTG1 B-cell translocation gene 1, anti-proliferative V V V V V V V 7 CLCN3 chloride channel, voltage-sensitive 3 V V V V V V V 7 CTDSPL CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) small phosphatase-like V V V V V V V 7 DUSP10 dual specificity phosphatase 10 V V V V V V V 7 MAP3K4 mitogen-activated protein kinase kinase kinase 4 V V V V V V V 7 PDCD4 programmed cell death 4 (neoplastic transformation inhibitor) V V V V V V V 7 PPP2R5C protein phosphatase 2, regulatory subunit B', gamma V V V V V V V 7 PTPN4 protein tyrosine phosphatase, non-receptor type 4 (megakaryocyte) V V V V V V V 7 EZR ezrin V V V V V V 6 FOXO1 forkhead box O1 V V V V V V 6 ANKRD13C ankyrin repeat domain 13C V V V V V V 6 ARHGAP6 Rho GTPase activating protein 6 V V V V V V 6 BACH2 BTB and CNC homology 1, basic leucine zipper transcription factor 2 V V V V V V 6 BNIP3L BCL2/adenovirus E1B 19kDa interacting protein 3-like V V V V V V 6 BRMS1L breast cancer metastasis-suppressor 1-like V V V V V V 6 CDK5R1 cyclin-dependent kinase 5, regulatory subunit 1 (p35) V V V V V V 6 CTDSP1 CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) small phosphatase 1 V V V V V V 6 DCX doublecortin V V V V V V 6 ENAH enabled homolog (Drosophila) V V V V V V 6 EPHA4 EPH receptor A4 V V V V V V 6 FOXP1 forkhead box P1 V -

Sleep Alterations in Mouse Genetic Models of Human Disease
University of Kentucky UKnowledge Theses and Dissertations--Biology Biology 2016 SLEEP ALTERATIONS IN MOUSE GENETIC MODELS OF HUMAN DISEASE Mansi Sethi University of Kentucky, [email protected] Author ORCID Identifier: http://orcid.org/0000-0002-6636-171X Digital Object Identifier: https://doi.org/10.13023/ETD.2016.522 Right click to open a feedback form in a new tab to let us know how this document benefits ou.y Recommended Citation Sethi, Mansi, "SLEEP ALTERATIONS IN MOUSE GENETIC MODELS OF HUMAN DISEASE" (2016). Theses and Dissertations--Biology. 38. https://uknowledge.uky.edu/biology_etds/38 This Doctoral Dissertation is brought to you for free and open access by the Biology at UKnowledge. It has been accepted for inclusion in Theses and Dissertations--Biology by an authorized administrator of UKnowledge. For more information, please contact [email protected]. STUDENT AGREEMENT: I represent that my thesis or dissertation and abstract are my original work. Proper attribution has been given to all outside sources. I understand that I am solely responsible for obtaining any needed copyright permissions. I have obtained needed written permission statement(s) from the owner(s) of each third-party copyrighted matter to be included in my work, allowing electronic distribution (if such use is not permitted by the fair use doctrine) which will be submitted to UKnowledge as Additional File. I hereby grant to The University of Kentucky and its agents the irrevocable, non-exclusive, and royalty-free license to archive and make accessible my work in whole or in part in all forms of media, now or hereafter known. -
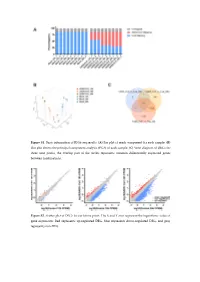
Figure S1. Basic Information of RNA-Seq Results. (A) Bar Plot of Reads Component for Each Sample
Figure S1. Basic information of RNA-seq results. (A) Bar plot of reads component for each sample. (B) Dot plot shows the principal component analysis (PCA) of each sample. (C) Venn diagram of DEGs for three time points, the overlap part of the circles represents common differentially expressed genes between combinations. Figure S2. Scatter plot of DEGs for each time point. The X and Y axes represent the logarithmic value of gene expression. Red represents up-regulated DEG, blue represents down-regulated DEG, and gray represents non-DEG. Table S1. Primers used for quantitative real-time PCR analysis of DEGs. Gene Primer Sequence Forward 5’-CTACGAGTGGATGGTCAAGAGC-3’ FOXO1 Reverse 5’-CCAGTTCCTTCATTCTGCACACG-3’ Forward 5’-GACGTCCGGCATCAGAGAAA-3’ IRS2 Reverse 5’-TCCACGGCTAATCGTCACAG-3’ Forward 5’-CACAACCAGGACCTCACACC-3’ IRS1 Reverse 5’-CTTGGCACGATAGAGAGCGT-3’ Forward 5’-AGGATACCACTCCCAACAGACCT-3’ IL6 Reverse 5’-CAAGTGCATCATCGTTGTTCATAC-3’ Forward 5’-TCACGTTGTACGCAGCTACC-3’ CCL5 Reverse 5’-CAGTCCTCTTACAGCCTTTGG-3’ Forward 5’-CTGTGCAGCCGCAGTGCCTACC-3’ BMP7 Reverse 5’-ATCCCTCCCCACCCCACCATCT-3’ Forward 5’-CTCTCCCCCTCGACTTCTGA-3’ BCL2 Reverse 5’-AGTCACGCGGAACACTTGAT-3’ Forward 5’-CTGTCGAACACAGTGGTACCTG-3’ FGF7 Reverse 5’-CCAACTGCCACTGTCCTGATTTC-3’ Forward 5’-GGGAGCCAAAAGGGTCATCA-3’ GAPDH Reverse 5’-CGTGGACTGTGGTCATGAGT-3’ Supplementary material: Differentially expressed genes log2(SADS-CoV_12h/ Qvalue (SADS-CoV _12h/ Gene Symbol Control_12h) Control_12h) PTGER4 -1.03693 6.79E-04 TMEM72 -3.08132 3.66E-04 IFIT2 -1.02918 2.11E-07 FRAT2 -1.09282 4.66E-05 -

Inbred Mouse Strains Expression in Primary Immunocytes Across
Downloaded from http://www.jimmunol.org/ by guest on September 26, 2021 Daphne is online at: average * The Journal of Immunology published online 29 September 2014 from submission to initial decision 4 weeks from acceptance to publication Sara Mostafavi, Adriana Ortiz-Lopez, Molly A. Bogue, Kimie Hattori, Cristina Pop, Daphne Koller, Diane Mathis, Christophe Benoist, The Immunological Genome Consortium, David A. Blair, Michael L. Dustin, Susan A. Shinton, Richard R. Hardy, Tal Shay, Aviv Regev, Nadia Cohen, Patrick Brennan, Michael Brenner, Francis Kim, Tata Nageswara Rao, Amy Wagers, Tracy Heng, Jeffrey Ericson, Katherine Rothamel, Adriana Ortiz-Lopez, Diane Mathis, Christophe Benoist, Taras Kreslavsky, Anne Fletcher, Kutlu Elpek, Angelique Bellemare-Pelletier, Deepali Malhotra, Shannon Turley, Jennifer Miller, Brian Brown, Miriam Merad, Emmanuel L. Gautier, Claudia Jakubzick, Gwendalyn J. Randolph, Paul Monach, Adam J. Best, Jamie Knell, Ananda Goldrath, Vladimir Jojic, J Immunol http://www.jimmunol.org/content/early/2014/09/28/jimmun ol.1401280 Koller, David Laidlaw, Jim Collins, Roi Gazit, Derrick J. Rossi, Nidhi Malhotra, Katelyn Sylvia, Joonsoo Kang, Natalie A. Bezman, Joseph C. Sun, Gundula Min-Oo, Charlie C. Kim and Lewis L. Lanier Variation and Genetic Control of Gene Expression in Primary Immunocytes across Inbred Mouse Strains Submit online. Every submission reviewed by practicing scientists ? is published twice each month by http://jimmunol.org/subscription http://www.jimmunol.org/content/suppl/2014/09/28/jimmunol.140128 0.DCSupplemental Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2014 by The American Association of Immunologists, Inc.